








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 349-361.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0437
徐泽( ), 周陈平, 邝瑞彬, 吴夏明, 杨敏, 刘传和, 贺涵, 魏岳荣(
), 周陈平, 邝瑞彬, 吴夏明, 杨敏, 刘传和, 贺涵, 魏岳荣( )
)
收稿日期:2025-04-24
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
魏岳荣,男,博士,研究员,研究方向 :番木瓜种质资源收集评价、细胞工程和分子育种;E-mail: weid18@163.com作者简介:徐泽,女,博士,助理研究员,研究方向 :果实贮藏保鲜;E-mail: xuzeizi@163.com
基金资助:
XU Ze( ), ZHOU Chen-ping, KUANG Rui-bin, WU Xia-ming, YANG Min, LIU Chuan-he, HE Han, WEI Yue-rong(
), ZHOU Chen-ping, KUANG Rui-bin, WU Xia-ming, YANG Min, LIU Chuan-he, HE Han, WEI Yue-rong( )
)
Received:2025-04-24
Published:2026-03-26
Online:2026-04-23
摘要:
目的 多聚半乳糖醛酸酶(PG)是调控果实软化的关键因子,鉴定番木瓜PG基因家族成员并进行功能分析,为揭示番木瓜采后软化的分子机制提供理论基础。 方法 基于‘紫晖’番木瓜基因组数据,全面鉴定CpPG家族成员,分析其蛋白理化性质、系统进化关系、保守基序、基因结构、基因复制事件、启动子顺式作用元件和表达水平;进一步利用果实瞬时过表达试验,探究CpPG1、CpPG2、CpPG28在番木瓜果实采后软化中的功能。 结果 在‘紫晖’番木瓜基因组中鉴定出38个PG家族成员,命名为CpPG1-CpPG38,这些成员不均匀分布在8条染色体上。系统进化分析表明CpPG成员聚类为亚组A-F,各亚组内基因成员的外显子/内含子结构相对保守,不同组别CpPG成员在保守结构域和蛋白基序具有显著差异。共线性分析结果表明,番木瓜PG家族存在基因复制事件和基因串联重复。CpPG成员的启动子区域包含多种光响应、植物生长发育与逆境胁迫、植物激素应答相关作用元件,参与多种生长发育进程。通过分析38个CpPG基因的表达模式,筛选出CpPG1、CpPG2和CpPG28 3个核心成员,这些基因在成熟番木瓜果实中高表达并响应乙烯诱导。瞬时过表达试验进一步证实,CpPG1、CpPG2和CpPG28均能显著降低果实硬度,揭示其在番木瓜果实软化中的关键作用。 结论 番木瓜基因组中鉴定出38个CpPG基因,挖掘3个调控番木瓜果实软化的关键家族成员CpPG1、CpPG2和CpPG28。
徐泽, 周陈平, 邝瑞彬, 吴夏明, 杨敏, 刘传和, 贺涵, 魏岳荣. 番木瓜PG基因家族鉴定及其在果实软化中的功能[J]. 生物技术通报, 2026, 42(3): 349-361.
XU Ze, ZHOU Chen-ping, KUANG Rui-bin, WU Xia-ming, YANG Min, LIU Chuan-he, HE Han, WEI Yue-rong. Identification of PG Gene Family and Their Roles in Papaya Fruit Softening[J]. Biotechnology Bulletin, 2026, 42(3): 349-361.
引物名称 Gene name | 引物序列 Primer sequence (5′‒3′) |
|---|---|
| CpPG1-F | AATGGTGGTGCGTATAGATGGAAC |
| CpPG1-R | GGGAGTTGAGAAACCCGATTGAC |
| CpPG2-F | CACCCTTCATTTCTCAGCTCCATT |
| CpPG2-R | TTTCTCCCAAGCCTCCTCAAATAC |
| CpPG28-F | ACCATTGACGGACAAGGCTCTAT |
| CpPG28-R | TCAGACAAGGTGAACTCGACAAGA |
| OE-CpPG1-F | CCAAATCGACTCTAGTCTAGAATGACGACAATCCGCTCTCACAAC |
| OE-CpPG1-R | GCTCACCATCTGCAGAAGCTTCAAGCAACTATTGGGCTGAACCG |
| OE-CpPG2-F | CCAAATCGACTCTAGTCTAGAATGGCTGTTCTATATTATGATAATTATAATCGTACCC |
| OE-CpPG2-R | GCTCACCATCTGCAGAAGCTTCCAGATTATCATCTTAAACATGACGAGCTT |
| OE-CpPG28-F | CCAAATCGACTCTAGTCTAGAATGAAGATGCCAGTAGCATTTTTCTTGC |
| OE-CpPG28-R | GCTCACCATCTGCAGAAGCTTTACCGGGACAGCCACAGTTTCA |
| EIF4A-F | AGGCAGGCAAGAGAAGAT |
| EIF4A-R | TTCATACCGAGTAGCGATTC |
表1 引物序列信息
Table 1 Information of primers
引物名称 Gene name | 引物序列 Primer sequence (5′‒3′) |
|---|---|
| CpPG1-F | AATGGTGGTGCGTATAGATGGAAC |
| CpPG1-R | GGGAGTTGAGAAACCCGATTGAC |
| CpPG2-F | CACCCTTCATTTCTCAGCTCCATT |
| CpPG2-R | TTTCTCCCAAGCCTCCTCAAATAC |
| CpPG28-F | ACCATTGACGGACAAGGCTCTAT |
| CpPG28-R | TCAGACAAGGTGAACTCGACAAGA |
| OE-CpPG1-F | CCAAATCGACTCTAGTCTAGAATGACGACAATCCGCTCTCACAAC |
| OE-CpPG1-R | GCTCACCATCTGCAGAAGCTTCAAGCAACTATTGGGCTGAACCG |
| OE-CpPG2-F | CCAAATCGACTCTAGTCTAGAATGGCTGTTCTATATTATGATAATTATAATCGTACCC |
| OE-CpPG2-R | GCTCACCATCTGCAGAAGCTTCCAGATTATCATCTTAAACATGACGAGCTT |
| OE-CpPG28-F | CCAAATCGACTCTAGTCTAGAATGAAGATGCCAGTAGCATTTTTCTTGC |
| OE-CpPG28-R | GCTCACCATCTGCAGAAGCTTTACCGGGACAGCCACAGTTTCA |
| EIF4A-F | AGGCAGGCAAGAGAAGAT |
| EIF4A-R | TTCATACCGAGTAGCGATTC |
| 基因名称 | 基因号 | 氨基酸数 | 分子量 | 等电点 | 不稳定指数 | 脂肪系数 | 亲疏水性 | 信号肽 |
|---|---|---|---|---|---|---|---|---|
| Gene name | Gene ID | Number of amino acids (aa) | Molecular weight (kD) | pI | Instability index | Aliphatic index | Hydropathicity index | Signal peptide |
| CpPG1 | Cp_zihui11610 | 359 | 38.0 | 9.24 | 31.42 | 85.46 | -0.060 | + |
| CpPG2 | Cp_zihui13876 | 437 | 48.2 | 5.69 | 34.70 | 77.12 | -0.509 | - |
| CpPG3 | Cp_zihui12587 | 428 | 47.2 | 8.75 | 28.24 | 79.88 | -0.373 | + |
| CpPG4 | Cp_zihui01876 | 390 | 42.2 | 8.63 | 44.78 | 87.95 | -0.046 | + |
| CpPG5 | Cp_zihui19603 | 482 | 52.9 | 8.34 | 41.88 | 86.72 | -0.111 | - |
| CpPG6 | Cp_zihui08791 | 447 | 48.7 | 7.57 | 36.53 | 89.80 | -0.009 | + |
| CpPG7 | Cp_zihui00474 | 401 | 43.2 | 7.41 | 26.91 | 86.28 | -0.026 | + |
| CpPG8 | Cp_zihui00470 | 278 | 30.7 | 8.94 | 28.18 | 79.93 | -0.251 | - |
| CpPG9 | Cp_zihui00483 | 461 | 50.0 | 8.34 | 33.67 | 81.58 | -0.211 | - |
| CpPG10 | Cp_zihui02232 | 458 | 49.0 | 6.30 | 44.43 | 77.86 | -0.171 | + |
| CpPG11 | Cp_zihui06380 | 262 | 29.1 | 5.77 | 58.76 | 84.43 | -0.274 | - |
| CpPG12 | Cp_zihui07099 | 432 | 45.8 | 5.35 | 45.09 | 77.59 | -0.076 | + |
| CpPG13 | Cp_zihui08226 | 374 | 40.8 | 8.64 | 33.01 | 85.78 | -0.102 | + |
| CpPG14 | Cp_zihui08889 | 488 | 53.6 | 5.38 | 35.37 | 90.20 | -0.143 | + |
| CpPG15 | Cp_zihui09177 | 481 | 52.4 | 8.34 | 34.35 | 88.90 | -0.025 | + |
| CpPG16 | Cp_zihui10261 | 349 | 37.8 | 7.89 | 35.97 | 86.07 | -0.200 | - |
| CpPG17 | Cp_zihui10262 | 371 | 40.4 | 8.71 | 30.26 | 84.88 | -0.188 | - |
| CpPG18 | Cp_zihui10270 | 335 | 35.8 | 6.72 | 45.96 | 94.84 | -0.067 | + |
| CpPG19 | Cp_zihui10273 | 431 | 47.5 | 8.55 | 44.22 | 88.19 | -0.016 | + |
| CpPG20 | Cp_zihui10665 | 475 | 51.6 | 5.48 | 50.72 | 83.28 | -0.080 | - |
| CpPG21 | Cp_zihui10984 | 498 | 54.8 | 5.12 | 37.87 | 89.22 | -0.144 | - |
| CpPG22 | Cp_zihui11565 | 530 | 58.3 | 5.71 | 31.68 | 87.19 | 0.085 | - |
| CpPG23 | Cp_zihui11577 | 425 | 45.4 | 6.78 | 29.06 | 84.24 | -0.052 | + |
| CpPG24 | Cp_zihui11602 | 356 | 37.6 | 9.6 | 28.92 | 87.36 | 0.043 | + |
| CpPG25 | Cp_zihui11601 | 393 | 42.0 | 9.68 | 36.76 | 80.87 | -0.097 | + |
| CpPG26 | Cp_zihui11608 | 395 | 43.1 | 7.58 | 38.66 | 84.89 | -0.072 | + |
| CpPG27 | Cp_zihui11609 | 403 | 42.8 | 5.40 | 38.28 | 80.77 | -0.082 | + |
| CpPG28 | Cp_zihui11883 | 457 | 49.6 | 5.59 | 48.03 | 90.83 | -0.042 | + |
| CpPG29 | Cp_zihui12006 | 494 | 53.1 | 4.80 | 53.12 | 73.00 | -0.252 | + |
| CpPG30 | Cp_zihui13178 | 477 | 52.2 | 5.35 | 33.45 | 85.97 | -0.203 | + |
| CpPG31 | Cp_zihui13181 | 454 | 49.9 | 8.72 | 43.42 | 88.41 | -0.148 | + |
| CpPG32 | Cp_zihui13193 | 499 | 54.3 | 5.20 | 34.53 | 81.42 | -0.164 | + |
| CpPG33 | Cp_zihui14064 | 477 | 52.8 | 6.71 | 44.22 | 88.68 | -0.208 | - |
| CpPG34 | Cp_zihui14065 | 429 | 46.3 | 5.38 | 42.46 | 91.17 | -0.078 | + |
| CpPG35 | Cp_zihui16353 | 240 | 27.3 | 6.11 | 29.48 | 116.88 | 0.202 | + |
| CpPG36 | Cp_zihui16358 | 277 | 30.2 | 7.21 | 42.86 | 92.82 | -0.201 | - |
| CpPG37 | Cp_zihui16904 | 536 | 60.3 | 8.70 | 44.09 | 87.95 | -0.238 | - |
| CpPG38 | Cp_zihui19689 | 415 | 45.0 | 4.61 | 39.69 | 77.49 | -0.295 | + |
表2 番木瓜PG基因家族成员基本理化性质
Table 2 Physicochemical properties of the PG gene family membersinpapaya (Carica papaya L.)
| 基因名称 | 基因号 | 氨基酸数 | 分子量 | 等电点 | 不稳定指数 | 脂肪系数 | 亲疏水性 | 信号肽 |
|---|---|---|---|---|---|---|---|---|
| Gene name | Gene ID | Number of amino acids (aa) | Molecular weight (kD) | pI | Instability index | Aliphatic index | Hydropathicity index | Signal peptide |
| CpPG1 | Cp_zihui11610 | 359 | 38.0 | 9.24 | 31.42 | 85.46 | -0.060 | + |
| CpPG2 | Cp_zihui13876 | 437 | 48.2 | 5.69 | 34.70 | 77.12 | -0.509 | - |
| CpPG3 | Cp_zihui12587 | 428 | 47.2 | 8.75 | 28.24 | 79.88 | -0.373 | + |
| CpPG4 | Cp_zihui01876 | 390 | 42.2 | 8.63 | 44.78 | 87.95 | -0.046 | + |
| CpPG5 | Cp_zihui19603 | 482 | 52.9 | 8.34 | 41.88 | 86.72 | -0.111 | - |
| CpPG6 | Cp_zihui08791 | 447 | 48.7 | 7.57 | 36.53 | 89.80 | -0.009 | + |
| CpPG7 | Cp_zihui00474 | 401 | 43.2 | 7.41 | 26.91 | 86.28 | -0.026 | + |
| CpPG8 | Cp_zihui00470 | 278 | 30.7 | 8.94 | 28.18 | 79.93 | -0.251 | - |
| CpPG9 | Cp_zihui00483 | 461 | 50.0 | 8.34 | 33.67 | 81.58 | -0.211 | - |
| CpPG10 | Cp_zihui02232 | 458 | 49.0 | 6.30 | 44.43 | 77.86 | -0.171 | + |
| CpPG11 | Cp_zihui06380 | 262 | 29.1 | 5.77 | 58.76 | 84.43 | -0.274 | - |
| CpPG12 | Cp_zihui07099 | 432 | 45.8 | 5.35 | 45.09 | 77.59 | -0.076 | + |
| CpPG13 | Cp_zihui08226 | 374 | 40.8 | 8.64 | 33.01 | 85.78 | -0.102 | + |
| CpPG14 | Cp_zihui08889 | 488 | 53.6 | 5.38 | 35.37 | 90.20 | -0.143 | + |
| CpPG15 | Cp_zihui09177 | 481 | 52.4 | 8.34 | 34.35 | 88.90 | -0.025 | + |
| CpPG16 | Cp_zihui10261 | 349 | 37.8 | 7.89 | 35.97 | 86.07 | -0.200 | - |
| CpPG17 | Cp_zihui10262 | 371 | 40.4 | 8.71 | 30.26 | 84.88 | -0.188 | - |
| CpPG18 | Cp_zihui10270 | 335 | 35.8 | 6.72 | 45.96 | 94.84 | -0.067 | + |
| CpPG19 | Cp_zihui10273 | 431 | 47.5 | 8.55 | 44.22 | 88.19 | -0.016 | + |
| CpPG20 | Cp_zihui10665 | 475 | 51.6 | 5.48 | 50.72 | 83.28 | -0.080 | - |
| CpPG21 | Cp_zihui10984 | 498 | 54.8 | 5.12 | 37.87 | 89.22 | -0.144 | - |
| CpPG22 | Cp_zihui11565 | 530 | 58.3 | 5.71 | 31.68 | 87.19 | 0.085 | - |
| CpPG23 | Cp_zihui11577 | 425 | 45.4 | 6.78 | 29.06 | 84.24 | -0.052 | + |
| CpPG24 | Cp_zihui11602 | 356 | 37.6 | 9.6 | 28.92 | 87.36 | 0.043 | + |
| CpPG25 | Cp_zihui11601 | 393 | 42.0 | 9.68 | 36.76 | 80.87 | -0.097 | + |
| CpPG26 | Cp_zihui11608 | 395 | 43.1 | 7.58 | 38.66 | 84.89 | -0.072 | + |
| CpPG27 | Cp_zihui11609 | 403 | 42.8 | 5.40 | 38.28 | 80.77 | -0.082 | + |
| CpPG28 | Cp_zihui11883 | 457 | 49.6 | 5.59 | 48.03 | 90.83 | -0.042 | + |
| CpPG29 | Cp_zihui12006 | 494 | 53.1 | 4.80 | 53.12 | 73.00 | -0.252 | + |
| CpPG30 | Cp_zihui13178 | 477 | 52.2 | 5.35 | 33.45 | 85.97 | -0.203 | + |
| CpPG31 | Cp_zihui13181 | 454 | 49.9 | 8.72 | 43.42 | 88.41 | -0.148 | + |
| CpPG32 | Cp_zihui13193 | 499 | 54.3 | 5.20 | 34.53 | 81.42 | -0.164 | + |
| CpPG33 | Cp_zihui14064 | 477 | 52.8 | 6.71 | 44.22 | 88.68 | -0.208 | - |
| CpPG34 | Cp_zihui14065 | 429 | 46.3 | 5.38 | 42.46 | 91.17 | -0.078 | + |
| CpPG35 | Cp_zihui16353 | 240 | 27.3 | 6.11 | 29.48 | 116.88 | 0.202 | + |
| CpPG36 | Cp_zihui16358 | 277 | 30.2 | 7.21 | 42.86 | 92.82 | -0.201 | - |
| CpPG37 | Cp_zihui16904 | 536 | 60.3 | 8.70 | 44.09 | 87.95 | -0.238 | - |
| CpPG38 | Cp_zihui19689 | 415 | 45.0 | 4.61 | 39.69 | 77.49 | -0.295 | + |
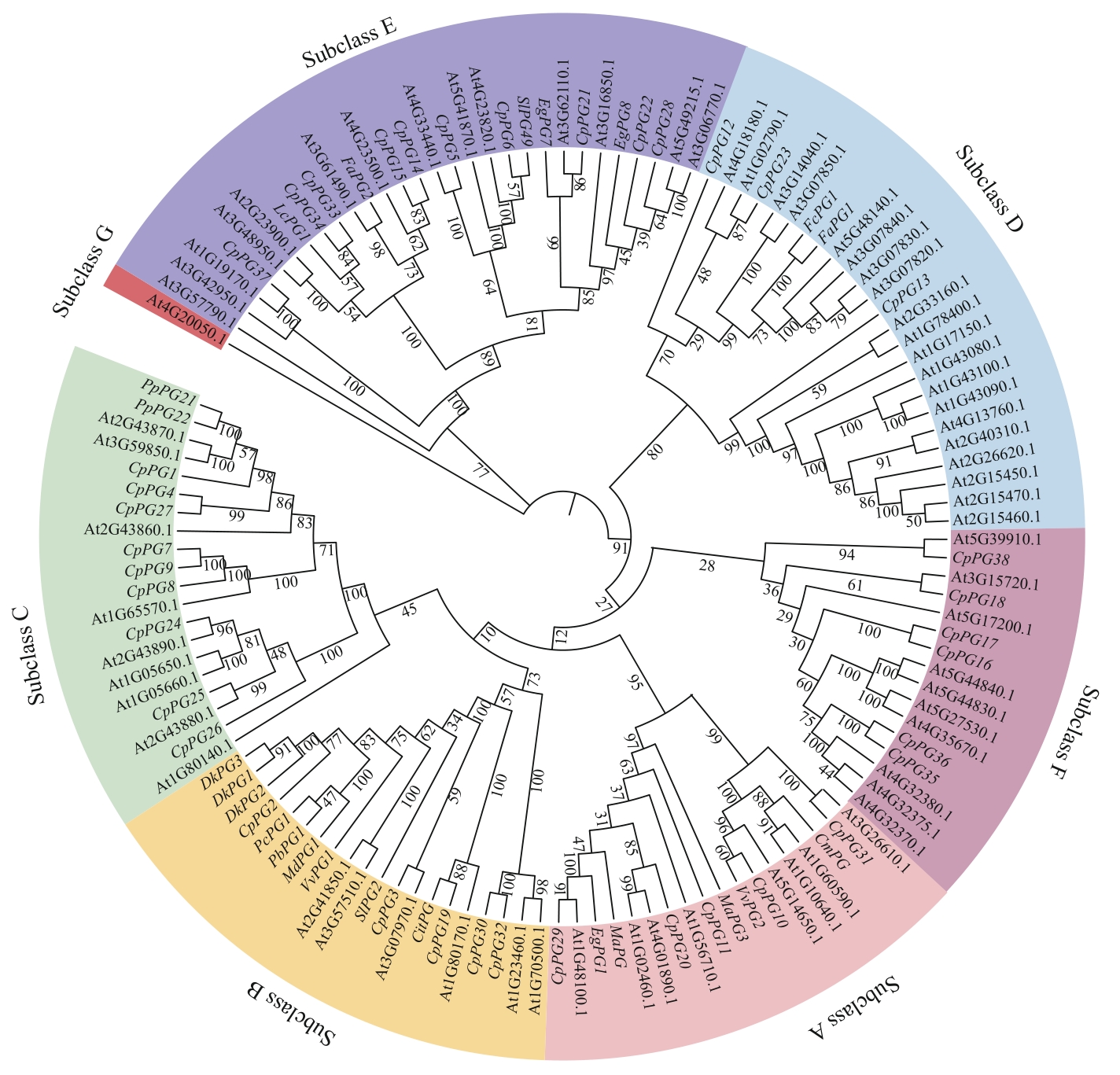
图3 PG基因家族系统发育树At:拟南芥;Lc:荔枝;Fa:栽培草莓;Sl:番茄;Eg:油棕;Fc:智利草莓;Cm:甜瓜;Vv:葡萄;Ma:香蕉;Dk:柿;Pc:西洋梨;Pb:白梨;Md:苹果;Cit:甜橙;Pp:桃
Fig. 3 Phylogenetic tree of the PG gene familyAt: Arabidopsis thaliana; Lc: Litchi chinensis; Fa: Fragaria × ananassa; Sl: Solanum lycopersicum; Eg: Elaeis guineensis; Fc: Fragaria chiloensis; Cm: Cucumis melo; Vv: Vitis vinifera; Ma: Musa acuminata; Dk: Diospyros kaki; Pc: Pyrus communis; Pb: Pyrus bretschneideri; Md: Malus domestica; Cit: Citrus sinensis; Pp: Prunus persica
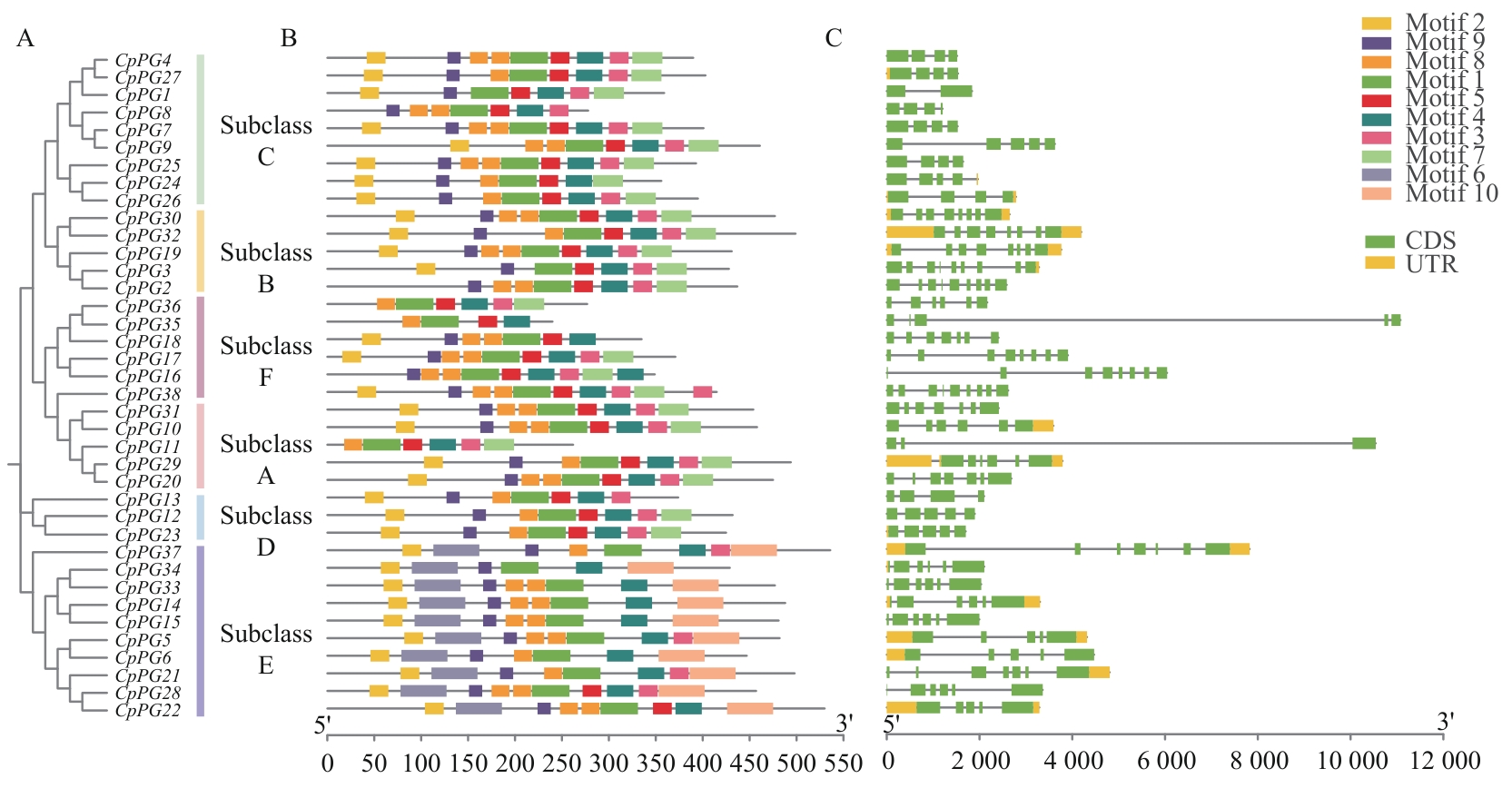
图4 番木瓜PG基因系统发育关系(A)、保守基序(B)和基因结构(C)
Fig. 4 Phylogenetic relationship (A), distribution of conserved motif (B) and gene structure (C) in the papaya PG members
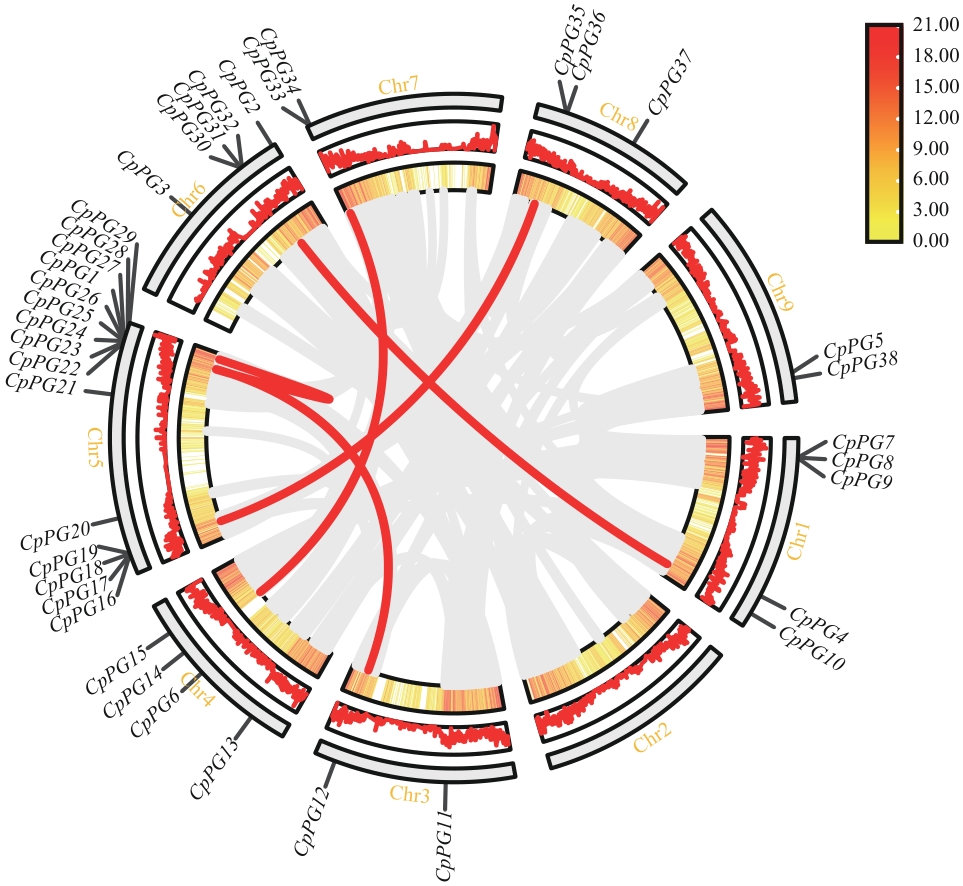
图5 番木瓜PG家族共线性分析灰线表示番木瓜基因组内共线的片段,红线表示共线的PG基因对
Fig. 5 Syntenic analysis of PG gene family in papayaThe gray lines indicate collinear blocks within the papaya genomes, while the red lines indicate collinear PG gene pairs
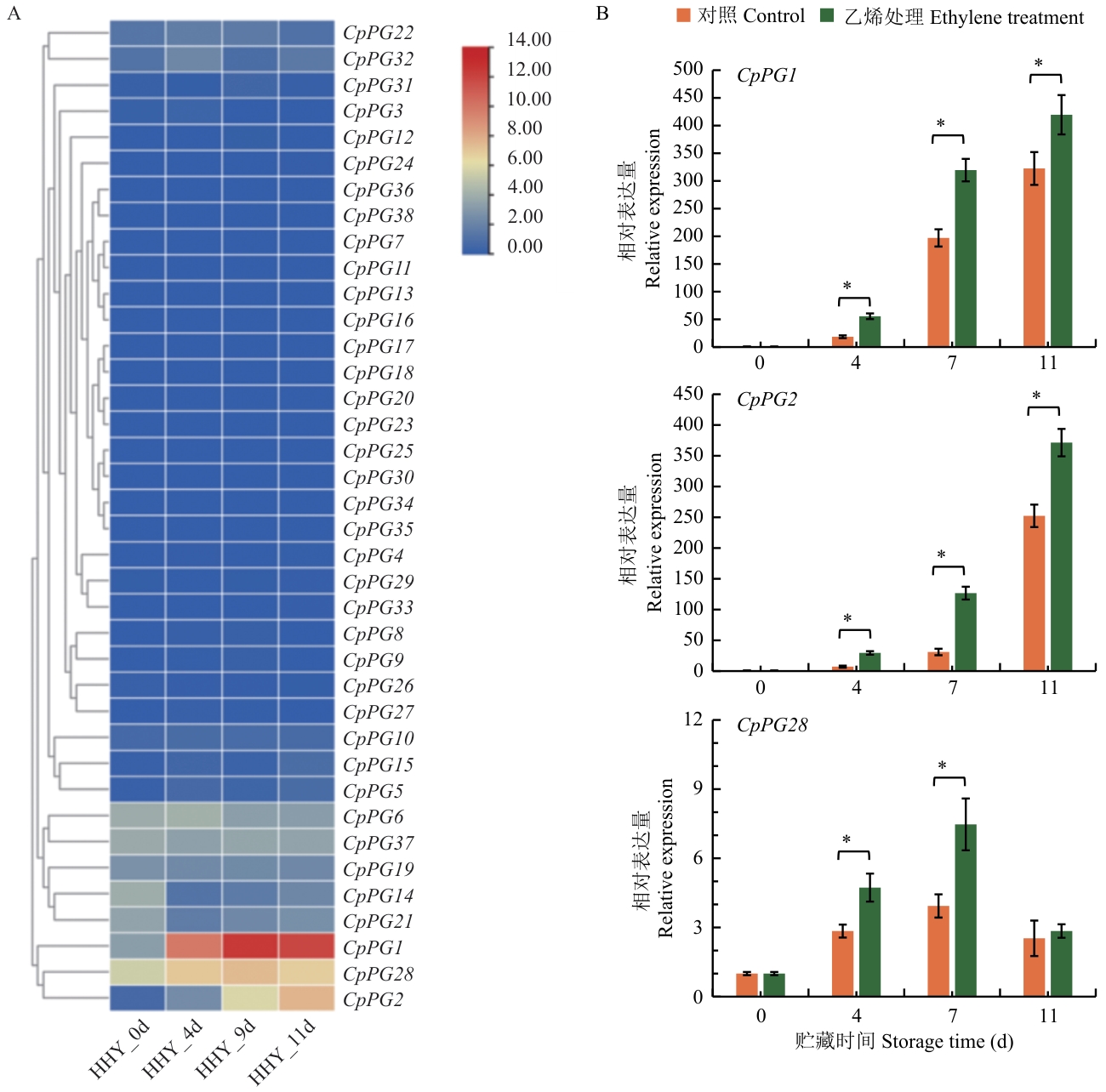
图7 番木瓜PG基因表达模式A:转录组数据分析不同CpPG在果实采后贮藏阶段的表达水平(HHY_0d:‘黄花佑’番木瓜果实采后贮藏0 d;HHY_4d:‘黄花佑’番木瓜果实采后贮藏4 d;HHY_9d:‘黄花佑’番木瓜果实采后贮藏9 d;HHY_11d:‘黄花佑’番木瓜果实采后贮藏11 d);B:乙烯处理对CpPG1、CpPG2、CpPG28表达水平的影响;* P<0.05,下同
Fig. 7 Expression pattern of papaya PG genesA: Transcriptome analysis of different CpPG expressions during storage stage (HHY_0d: ‘Huanghuayou’ fruits stored for 0 d; HHY_4d: ‘Huanghuayou’ fruits stored for 4 d; HHY_9d: ‘Huanghuayou’ fruits stored for 9 d; HHY_11d: ‘Huanghuayou’ fruits stored for 11 d). B: Impact of ethylene treatment on the expression of CpPG1, CpPG2, and CpPG28. * P<0.05. The same below
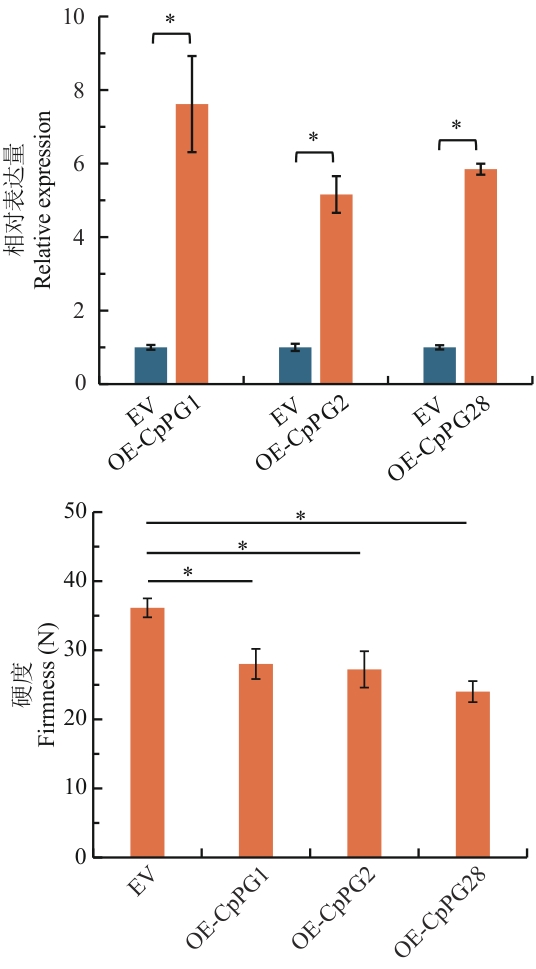
图8 分别瞬时过表达CpPG1、CpPG2和CpPG28对果实软化的影响EV:空载体;OE-CpPG1:瞬时过表达CpPG1;OE-CpPG2:瞬时过表达CpPG2;OE-CpPG28:瞬时过表达CpPG28
Fig. 8 Effects of transient overexpression of CpPG1, CpPG2 and CpPG28 on fruit softeningEV: Empty vector; OE-CpPG1: transient overexpression of CpPG1; OE-CpPG2: transient overexpression of CpPG2; OE-CpPG28: transient overexpression of CpPG28
| [1] | Wang WH, Wang YY, Chen T, et al. Current insights into posttranscriptional regulation of fleshy fruit ripening [J]. Plant Physiol, 2023, 192(3): 1785-1798. |
| [2] | Brummell DA, Bowen JK, Gapper NE. Biotechnological approaches for controlling postharvest fruit softening [J]. Curr Opin Biotechnol, 2022, 78: 102786. |
| [3] | 张芮宁, 袁舟宇, 陈萍. 番木瓜采后保鲜技术研究进展 [J]. 园艺与种苗, 2021, 41(2): 47-52. |
| Zhang RN, Yuan ZY, Chen P. Research progress on postharvest preservation technology of Carica papaya [J]. Hortic Seed, 2021, 41(2): 47-52. | |
| [4] | 葛廷, 黄雪, 谢让金. 多聚半乳糖醛酸基因在果树中的研究进展 [J]. 植物生理学报, 2019, 55(8): 1075-1088. |
| Ge T, Huang X, Xie RJ. Recent advances in polygalacturonase gene in fruit tree species [J]. Plant Physiol J, 2019, 55(8): 1075-1088. | |
| [5] | Nie HM, Shi Y, Geng XQ, et al. CRISRP/Cas9-mediated targeted mutagenesis of tomato polygalacturonase gene (SlPG) delays fruit softening [J]. Front Plant Sci, 2022, 13: 729128. |
| [6] | Niture SK. Comparative biochemical and structural characterizations of fungal polygalacturonases [J]. Biologia, 2008, 63(1): 1-19. |
| [7] | Zou QW, Tu RR, Wu JJ, et al. A polygalacturonase gene OsPG1 modulates water homeostasis in rice [J]. Crop J, 2024, 12(1): 79-91. |
| [8] | Fei YC, Mo Y, Xu JJ, et al. Genome-wide analysis of the polygalacturonase gene family in Macadamia and identification of members involved in fruit abscission [J]. Plants, 2025, 14(11): 1610. |
| [9] | Mahmood U, Li XD, Qian MC, et al. Comparative transcriptome and co-expression network analysis revealed the genes associated with senescence and polygalacturonase activity involved in pod shattering of rapeseed [J]. Biotechnol Biofuels Bioprod, 2023, 16(1): 20. |
| [10] | Liao YX, Xiang B, Xue ZZ, et al. HCC1, a polygalacturonase, regulates chlorophyll degradation via the ethylene synthesis pathway [J]. Rice, 2023, 16(1): 57. |
| [11] | Rhee SY, Osborne E, Poindexter PD, et al. Microspore separation in the quartet 3 mutants of Arabidopsis is impaired by a defect in a developmentally regulated polygalacturonase required for pollen mother cell wall degradation [J]. Plant Physiol, 2003, 133(3): 1170-1180. |
| [12] | Gu C, Wang L, Wang W, et al. Copy number variation of a gene cluster encoding endopolygalacturonase mediates flesh texture and stone adhesion in peach [J]. J Exp Bot, 2016, 67(6): 1993-2005. |
| [13] | Atkinson RG, Sutherland PW, Johnston SL, et al. Down-regulation of POLYGALACTURONASE1 alters firmness, tensile strength and water loss in apple (Malus × domestica) fruit [J]. BMC Plant Biol, 2012, 12: 129. |
| [14] | Fabi JP, Broetto SG, da Silva SLGL, et al. Analysis of Papaya cell wall-related genes during fruit ripening indicates a central role of polygalacturonases during pulp softening [J]. PLoS One, 2014, 9(8): e105685. |
| [15] | Fu CC, Chen HJ, Gao HY, et al. Histone deacetylase CpHDA3 is functionally associated with CpERF9 in suppression of CpPME1/2 and CpPG5 genes during papaya fruit ripening [J]. J Agric Food Chem, 2019, 67(32): 8919-8925. |
| [16] | Yang M, Kong XD, Zhou CP, et al. Genomic insights into the domestication and genetic basis of yield in papaya [J]. Hortic Res, 2025, 12(5): uhaf045. |
| [17] | Torki M, Mandaron P, Mache R, et al. Characterization of a ubiquitous expressed gene family encoding polygalacturonase in Arabidopsis thaliana [J]. Gene, 2000, 242(1/2): 427-436. |
| [18] | Ke XB, Wang HS, Li Y, et al. Genome-wide identification and analysis of polygalacturonase genes in Solanum lycopersicum [J]. Int J Mol Sci, 2018, 19(8): 2290. |
| [19] | Lu L, Hou QC, Wang LL, et al. Genome-wide identification and characterization of polygalacturonase gene family in maize (Zea mays L.) [J]. Int J Mol Sci, 2021, 22(19): 10722. |
| [20] | Zhao MT, Hu RX, Lin YX, et al. Genome-wide analysis of polygalacturonase gene family reveals its role in strawberry softening [J]. Plants, 2024, 13(13): 1838. |
| [21] | Huang WJ, Chen MY, Zhao TT, et al. Genome-wide identification and expression analysis of polygalacturonase gene family in kiwifruit (Actinidia chinensis) during fruit softening [J]. Plants, 2020, 9(3): 327. |
| [22] | Qian M, Zhang YK, Yan XY, et al. Identification and expression analysis of polygalacturonase family members during peach fruit softening [J]. Int J Mol Sci, 2016, 17(11): 1933. |
| [23] | Chen HF, Shao HX, Fan S, et al. Identification and phylogenetic analysis of the POLYGALACTURONASE gene family in apple [J]. Hortic Plant J, 2016, 2(5): 241-252. |
| [24] | Khan N, Fatima F, Haider MS, et al. Genome-wide identification and expression profiling of the polygalacturonase (PG) and pectin methylesterase (PME) genes in grapevine (Vitis vinifera L.) [J]. Int J Mol Sci, 2019, 20(13): 3180. |
| [25] | Wang Y, Fan ZY, Zhai YL, et al. Polygalacturonase gene family analysis identifies FcPG12 as a key player in fig (Ficus carica L.) fruit softening [J]. BMC Plant Biol, 2023, 23(1): 320. |
| [26] | Zhai ZF, Feng C, Wang YY, et al. Genome-wide identification of the xyloglucan endotransglucosylase/hydrolase (XTH) and polygalacturonase (PG) genes and characterization of their role in fruit softening of sweet cherry [J]. Int J Mol Sci, 2021, 22(22): 12331. |
| [27] | 黄少鹏, 符栩, 任正恺, 等. 基于转录组的荔枝PG基因家族鉴定及表达分析 [J]. 分子植物育种, 2022, 20(20): 6687-6695. |
| Huang SP, Fu X, Ren ZK, et al. Identification and expression analysis of litchi PG gene family based on transcriptome [J]. Mol Plant Breed, 2022, 20(20): 6687-6695. | |
| [28] | 刘若南, 刘林婷, 周秋蓉, 等. 柚多聚半乳糖醛酸酶基因家族的鉴定和分析 [J]. 热带作物学报, 2022, 43(4): 666-674. |
| Liu RN, Liu LT, Zhou QR, et al. Identification and analysis of polygalacturonase genes family in pomelo [J]. Chin J Trop Crops, 2022, 43(4): 666-674. | |
| [29] | Mahmood U, Fan YH, Wei SY, et al. Comprehensive analysis of polygalacturonase genes offers new insights into their origin and functional evolution in land plants [J]. Genomics, 2021, 113(1): 1096-1108. |
| [30] | Liu J, Sharma A, Niewiara MJ, et al. Papain-like cysteine proteases in Carica papaya: lineage-specific gene duplication and expansion [J]. BMC Genomics, 2018, 19(1): 26. |
| [31] | Sun MY, Zhang MY, Chen XN, et al. Rearrangement and domestication as drivers of Rosaceae mitogenome plasticity [J]. BMC Biol, 2022, 20(1): 181. |
| [32] | Peng G, Wu JY, Lu WJ, et al. A polygalacturonase gene clustered into clade E involved in lychee fruitlet abscission [J]. Sci Hortic, 2013, 150: 244-250. |
| [33] | Paniagua C, Ric-Varas P, García-Gago JA, et al. Elucidating the role of polygalacturonase genes in strawberry fruit softening [J]. J Exp Bot, 2020, 71(22): 7103-7117. |
| [34] | Li WQ, Xu LA, Xia R, et al. Cloning and functional identification of SlPG49 in Solanum lycopersicum [J]. Appl Sci, 2021, 11(23): 11450. |
| [35] | Ohashi T, Sari N, Misaki R, et al. Biochemical characterization of Arabidopsis clade F polygalacturonase shows a substrate preference toward oligogalacturonic acids [J]. J Biosci Bioeng, 2022, 133(1): 1-7. |
| [36] | Önder S, Tonguç M, Önder D, et al. Dynamic changes occur in the cell wall composition and related enzyme activities during flower development in Rosa damascena [J]. Front Plant Sci, 2023, 14: 1120098. |
| [37] | Qian M, Xu Z, Zhang ZH, et al. The downregulation of PpPG21 and PpPG22 influences peach fruit texture and softening [J]. Planta, 2021, 254(2): 22. |
| [38] | Deytieux-Belleau C, Vallet A, Donèche B, et al. Pectin methylesterase and polygalacturonase in the developing grape skin [J]. Plant Physiol Biochem, 2008, 46(7): 638-646. |
| [39] | Wang ZY, MacRae EA, Wright MA, et al. Polygalacturonase gene expression in kiwifruit: relationship to fruit softening and ethylene production [J]. Plant Mol Biol, 2000, 42(2): 317-328. |
| [40] | Liu YZ, Dong T, Lei Y, et al. Isolation of a polygalacturonase gene from Citrus sinensis fruit and its expression relative to fruit mastication trait, fruit development, and calcium or boron treatments [J]. Plant Mol Biol Report, 2011, 29(1): 51-59. |
| [41] | Shi YN, Li BJ, Grierson D, et al. Insights into cell wall changes during fruit softening from transgenic and naturally occurring mutants [J]. Plant Physiol, 2023, 192(3): 1671-1683. |
| [42] | Peng ZZ, Liu GS, Li HL, et al. Molecular and genetic events determining the softening of fleshy fruits: a comprehensive review [J]. Int J Mol Sci, 2022, 23(20): 12482. |
| [43] | 邵远志, 高毫杰, 贾志伟, 等. 1-MCP和乙烯利处理对番木瓜果实软化生理的影响 [J]. 中国食品学报, 2013, 13(2): 143-148. |
| Shao YZ, Gao HJ, Jia ZW, et al. The effects of 1-MCP and ethephon on softening physiology of Papaya fruit [J]. J Chin Inst Food Sci Technol, 2013, 13(2): 143-148. |
| [1] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [2] | 苏燕竹, 李达, 张爱爱, 刘永光, 张秀荣, 薛其勤. 大豆CAD基因家族的鉴定及表达分析[J]. 生物技术通报, 2026, 42(4): 101-113. |
| [3] | 刘淼, 林涛, 贾乐松, 胡丰, 李涛, 李志万, 刘美芳, 郑方燕, 崔龙. 从野生到栽培:番茄果实色泽的演化与调控机制[J]. 生物技术通报, 2026, 42(3): 187-202. |
| [4] | 颜晨琳, 李凡, 闫春婷, 程蛟文, 胡开林, 叶志彪, 宋建文. 番茄果实形态发育相关基因研究进展[J]. 生物技术通报, 2026, 42(3): 172-186. |
| [5] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [6] | 胡秋玲, 陈灵, 黄嘉怡, 赵梓乔, 潘璐怡, 刘慧丽, 刘太波. 多胺调控果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 203-212. |
| [7] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [8] | 崔之瀚, 魏庆镇, 胡娜, 包崇来, 王华森. 园艺作物IQD基因研究进展[J]. 生物技术通报, 2026, 42(3): 230-241. |
| [9] | 张高翔, 吴玉碧, 郭亚静, 纪薇, 杨忠义. 葡萄WD40基因家族鉴定及表达量分析[J]. 生物技术通报, 2026, 42(3): 324-337. |
| [10] | 彭楚, 孙娟利, 郑蓓蓓, 张若西, 韩月彭, 赵云. 果实花色苷酶促降解机制研究进展[J]. 生物技术通报, 2026, 42(3): 133-144. |
| [11] | 罗龙鑫, 李智, 李彤, 冯资权, 翟欣悦, 梁成林, 张亚丽, 吴上, 李媛媛, 姜翰. 苹果果实糖分形成的分子基础[J]. 生物技术通报, 2026, 42(3): 156-171. |
| [12] | 殷诗情, 田泰, 黄凤庭, 冯珑强, 王浩, 张静, 何文, 陈清, 王小蓉, 王燕. 果树果实硬度的调控机制研究进展[J]. 生物技术通报, 2026, 42(3): 213-229. |
| [13] | 杜丹, 郭翔, 胡鑫, 潘宇. 质体发育调控果实成熟与品质的研究进展[J]. 生物技术通报, 2026, 42(3): 48-59. |
| [14] | 李成泉, 史庆华, 杨晓玉. 园艺作物果实发育的miRNA调控网络:从分子机制到种质创新[J]. 生物技术通报, 2026, 42(3): 19-36. |
| [15] | 赵艳侠, 李倩, 孙家波, 梁红敏, 李冰冰. 草莓果实品质形成的关键调控基因及分子网络解析[J]. 生物技术通报, 2026, 42(3): 111-132. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||