








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 158-168.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0378
刘保财1,2,3( ), 胡学博2, 张武君1,3, 赵云青1,3, 黄颖桢1,3, 陈菁瑛1,3(
), 胡学博2, 张武君1,3, 赵云青1,3, 黄颖桢1,3, 陈菁瑛1,3( )
)
收稿日期:2025-04-11
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
陈菁瑛,女,研究员,研究方向 :药用植物资源利用与规范栽培;E-mail: cjy6601@163.com作者简介:刘保财,男,博士,助理研究员,研究方向 :药用植物繁殖、育种与栽培;E-mail: 626813844@qq.com
基金资助:
LIU Bao-cai1,2,3( ), HU Xue-bo2, ZHANG Wu-jun1,3, ZHAO Yun-qing1,3, HUANG Ying-zhen1,3, CHEN Jing-ying1,3(
), HU Xue-bo2, ZHANG Wu-jun1,3, ZHAO Yun-qing1,3, HUANG Ying-zhen1,3, CHEN Jing-ying1,3( )
)
Received:2025-04-11
Published:2026-02-26
Online:2026-03-17
摘要:
目的 石仙桃(Pholidota chinensis)系兰科(Orchidaceae)石仙桃属(Pholidota)珍稀濒危的多年附生草本植物,为了研究该物种及近源种功能基因的表达量,迫切需要筛选其稳定的内参基因用于实时荧光定量PCR(RT-qPCR)分析。 方法 采用RT-qPCR检测11个管家基因在石仙桃不同组织(根、根状茎、假鳞茎、叶片、芽、花序)和非生物胁迫(20 mmol/L MeJA和50 mmol/L NaCl喷洒叶片、不同光照强度)的表达量,用geNorm、NormFinder、BestKeeper、ΔCt和RefFinder几何平均值综合分析11个内参基因表达的稳定性。 结果 选取的11个基因在石仙桃的不同组织中具有单一的扩增条带和峰图,扩增效率及表达丰度基本达到了内参基因的要求,表达稳定性表明,Actin/f58p0可作为不同组织基因表达的最佳内参基因,而TUA3/f11p0可作为非生物胁迫的最佳内参基因。分别以Actin/f58p0和TUA3/f11p0作为内参基因对石仙桃中天麻素合成关键基因GT1、GT2、GT3-01、GT3-02、GT4、ADH-01、ADH-02、ADH-03表达分析,在不同组织中,二者表达趋势基本一致,但HCT基因表达则更适宜以Actin/f58p0作为内参基因;非生物胁迫中,以TUA3/f11p0作为内参基因更佳,尤其是在NaCl和MeJA处理中,但在光照胁迫下,Actin/f58p0和TUA3/f11p0皆可作为内参,MeJA较NaCl对石仙桃基因表达量影响时间长。 结论 Actin/f58p0和TUA3/f11p0分别可作为石仙桃不同组织、活性成分和非生物胁迫的内参基因,为石仙桃及近缘物种功能基因的发掘、表达量分析等研究奠定基础。
刘保财, 胡学博, 张武君, 赵云青, 黄颖桢, 陈菁瑛. 石仙桃内参基因筛选与应用[J]. 生物技术通报, 2026, 42(2): 158-168.
LIU Bao-cai, HU Xue-bo, ZHANG Wu-jun, ZHAO Yun-qing, HUANG Ying-zhen, CHEN Jing-ying. Screening of Internal Reference Gene for Pholidota chinensis and Their Applications[J]. Biotechnology Bulletin, 2026, 42(2): 158-168.
| 基因 Gene | 引物名称 Primer name | 引物序列 Primer sequence (5′‒3′) | 产物长度 Product length (bp) |
|---|---|---|---|
| UBQ | UBQ/f2p0 | F: GTGACTGGTGATCCTTGGCA R: ACCAACTTTCAACAGGCCAGA | 121 |
| TUA | TUA/f11p0 | F: CTGGGAAGGAAGATGCTGCA R: TCTAACCCGGTCTAGGCACA | 89 |
| TUA | TUA/f3p01 | F: GGCAAGTACATGGCATGCTG R: TCGTCTTGATTGTGGCGACA | 85 |
| TUA | TUA/f3p02 | F: GAATCAACTACCAACCTCCTACAG R: CTTCCATTCCTTCTCCGACATAC | 183 |
| EF-1α | EF-1α/f74p0 | F: GAAGGATCCAACTGGTGCCA R: TCGAGCACCATTGGAACGAA | 84 |
| EF-1α | EF-1α/f11p0 | F: GGTTCTCGACTGTCACACGT R: CCTTACCAGAGCGCCTATCG | 80 |
| Actin | Actin/f10p0 | F: GGATTTGCCGGTGATGATGC R: ATGCCAACCATGACACCAGT | 83 |
| Actin | Actin/f58p0 | F: CTCTTCTCTCCGCTGCCAAA R: CTTGGGAATACAGCCCTGGG | 142 |
| 18S rRNA | 18S/f2p0 | F: CGCTCCTACCGATTGAATGG R: TTGTTACGACTTCTCCTTCCTCTA | 125 |
| GAPDH | GAPDH/f24p0 | F: GCTCTGTTGAGGAGGGCATT R: TGATCTGAACCGTGCCAGTC | 129 |
| GAPDH | GAPDH/f94p0 | F: AGTATGACACTGTGCACGGG R: GGGATCTCCTCAGGGTTCCT | 121 |
表1 候选内参基因RT-qPCR引物序列
Table 1 Prime sequences of RT-qPCR primers for candidate internal reference genes
| 基因 Gene | 引物名称 Primer name | 引物序列 Primer sequence (5′‒3′) | 产物长度 Product length (bp) |
|---|---|---|---|
| UBQ | UBQ/f2p0 | F: GTGACTGGTGATCCTTGGCA R: ACCAACTTTCAACAGGCCAGA | 121 |
| TUA | TUA/f11p0 | F: CTGGGAAGGAAGATGCTGCA R: TCTAACCCGGTCTAGGCACA | 89 |
| TUA | TUA/f3p01 | F: GGCAAGTACATGGCATGCTG R: TCGTCTTGATTGTGGCGACA | 85 |
| TUA | TUA/f3p02 | F: GAATCAACTACCAACCTCCTACAG R: CTTCCATTCCTTCTCCGACATAC | 183 |
| EF-1α | EF-1α/f74p0 | F: GAAGGATCCAACTGGTGCCA R: TCGAGCACCATTGGAACGAA | 84 |
| EF-1α | EF-1α/f11p0 | F: GGTTCTCGACTGTCACACGT R: CCTTACCAGAGCGCCTATCG | 80 |
| Actin | Actin/f10p0 | F: GGATTTGCCGGTGATGATGC R: ATGCCAACCATGACACCAGT | 83 |
| Actin | Actin/f58p0 | F: CTCTTCTCTCCGCTGCCAAA R: CTTGGGAATACAGCCCTGGG | 142 |
| 18S rRNA | 18S/f2p0 | F: CGCTCCTACCGATTGAATGG R: TTGTTACGACTTCTCCTTCCTCTA | 125 |
| GAPDH | GAPDH/f24p0 | F: GCTCTGTTGAGGAGGGCATT R: TGATCTGAACCGTGCCAGTC | 129 |
| GAPDH | GAPDH/f94p0 | F: AGTATGACACTGTGCACGGG R: GGGATCTCCTCAGGGTTCCT | 121 |
| 基因 Gene | 引物名称 Primer name | 引物序列 Primer sequence (5′‒3′) |
|---|---|---|
| GT1 | 16563/f4p0/2237-F | TGACCTCGCAACAGATTATGAAC |
| 16563/f4p0/2237-R | TTCCAGTGCCTCAACAATAATTCT | |
| GT2 | 261/f5p0/5355-F | TCTGTAACAATCCAATGACCAAGG |
| 261/f5p0/5355-R | TCCAACTCTTCGCCTAGTATTCT | |
| GT3-01 | 17418/f3p0/2174-F | GTTAATAAGTGGAAGAGCGTAGCA |
| 17418/f3p0/2174-R | CACAACCTCCGCATCATCTC | |
| GT3-02 | 28360/f2p0/1592-F | GAAGGAGTTGATGGAAGGAGAG |
| 28360/f2p0/1592-R | TCACTTAGTAGCCGTTGAATCC | |
| GT4-01 | 20627/f2p0/1989-F | GTCAATGCTGGAGAATGCTTAATG |
| 20627/f2p0/1989-R | AAGAATGCCGCTGTTGAAGAT | |
| ADH-01 | 30246/f18p0/1465-F | AGTTGCCCATCTTCCCTCTAAT |
| 30246/f18p0/1465-R | GCATCGCTTCGTTCACATAGT | |
| ADH-02 | 32130/f3p0/1407-F | ATTGAGACAGTGAGACCAGAAGA |
| 32130/f3p0/1407-R | TTAGAACTAAGCGACGCCATTG | |
| ADH-03 | 30741/f57p0/1339-F | GGAAGCCTCATTGGTGGAATAAG |
| 30741/f57p0/1339-R | CGACATCAACAACGAACCGATAT | |
| HCT | 26704/f5p0/1679-F | GGGCACCGCAATTACTAATACTT |
| 26704/f5p0/1679-R | CACATCCACCAACAACCTCTC |
表2 天麻素合成通路中9个基因RT-qPCR引物序列
Table 2 Primer sequences of RT-qPCR for 9 key candidate genes of gastrodin biosynthesis pathway in P. chinensis
| 基因 Gene | 引物名称 Primer name | 引物序列 Primer sequence (5′‒3′) |
|---|---|---|
| GT1 | 16563/f4p0/2237-F | TGACCTCGCAACAGATTATGAAC |
| 16563/f4p0/2237-R | TTCCAGTGCCTCAACAATAATTCT | |
| GT2 | 261/f5p0/5355-F | TCTGTAACAATCCAATGACCAAGG |
| 261/f5p0/5355-R | TCCAACTCTTCGCCTAGTATTCT | |
| GT3-01 | 17418/f3p0/2174-F | GTTAATAAGTGGAAGAGCGTAGCA |
| 17418/f3p0/2174-R | CACAACCTCCGCATCATCTC | |
| GT3-02 | 28360/f2p0/1592-F | GAAGGAGTTGATGGAAGGAGAG |
| 28360/f2p0/1592-R | TCACTTAGTAGCCGTTGAATCC | |
| GT4-01 | 20627/f2p0/1989-F | GTCAATGCTGGAGAATGCTTAATG |
| 20627/f2p0/1989-R | AAGAATGCCGCTGTTGAAGAT | |
| ADH-01 | 30246/f18p0/1465-F | AGTTGCCCATCTTCCCTCTAAT |
| 30246/f18p0/1465-R | GCATCGCTTCGTTCACATAGT | |
| ADH-02 | 32130/f3p0/1407-F | ATTGAGACAGTGAGACCAGAAGA |
| 32130/f3p0/1407-R | TTAGAACTAAGCGACGCCATTG | |
| ADH-03 | 30741/f57p0/1339-F | GGAAGCCTCATTGGTGGAATAAG |
| 30741/f57p0/1339-R | CGACATCAACAACGAACCGATAT | |
| HCT | 26704/f5p0/1679-F | GGGCACCGCAATTACTAATACTT |
| 26704/f5p0/1679-R | CACATCCACCAACAACCTCTC |
| 候选基因 Candidate gene | 标准曲线斜率 Standard curve slope (k) | 扩增效率 Amplification efficiency (E)(%) | 相关系数 Correlation coefficient (R2) |
|---|---|---|---|
| UBQ/f2p0 | -3.435 8 | 95.46 | 0.998 1 |
| TUA/f11p0 | -2.523 7 | 109.03 | 0.987 9 |
| TUA/f3p01 | -3.188 3 | 105.90 | 0.993 7 |
| TUA/f3p02 | -3.492 5 | 93.34 | 0.997 2 |
| EF-1α/f74p0 | -3.509 9 | 92.71 | 0.993 8 |
| EF-1α/f11p0 | -3.239 2 | 103.57 | 0.994 3 |
| Actin/f10p0 | -3.148 9 | 100.78 | 0.991 8 |
| Actin/f58p0 | -3.312 8 | 100.38 | 0.995 6 |
| 18S/f2p0 | -3.646 5 | 88.03 | 0.992 9 |
| GAPDH/f24p0 | -3.374 4 | 97.86 | 0.997 9 |
| GAPDH/f94p0 | -3.330 2 | 99.66 | 0.985 6 |
表3 11个候选内参基因RT-qPCR的扩增参数
Table 3 Amplification parameters for RT-qPCR of 11 candidate reference genes
| 候选基因 Candidate gene | 标准曲线斜率 Standard curve slope (k) | 扩增效率 Amplification efficiency (E)(%) | 相关系数 Correlation coefficient (R2) |
|---|---|---|---|
| UBQ/f2p0 | -3.435 8 | 95.46 | 0.998 1 |
| TUA/f11p0 | -2.523 7 | 109.03 | 0.987 9 |
| TUA/f3p01 | -3.188 3 | 105.90 | 0.993 7 |
| TUA/f3p02 | -3.492 5 | 93.34 | 0.997 2 |
| EF-1α/f74p0 | -3.509 9 | 92.71 | 0.993 8 |
| EF-1α/f11p0 | -3.239 2 | 103.57 | 0.994 3 |
| Actin/f10p0 | -3.148 9 | 100.78 | 0.991 8 |
| Actin/f58p0 | -3.312 8 | 100.38 | 0.995 6 |
| 18S/f2p0 | -3.646 5 | 88.03 | 0.992 9 |
| GAPDH/f24p0 | -3.374 4 | 97.86 | 0.997 9 |
| GAPDH/f94p0 | -3.330 2 | 99.66 | 0.985 6 |
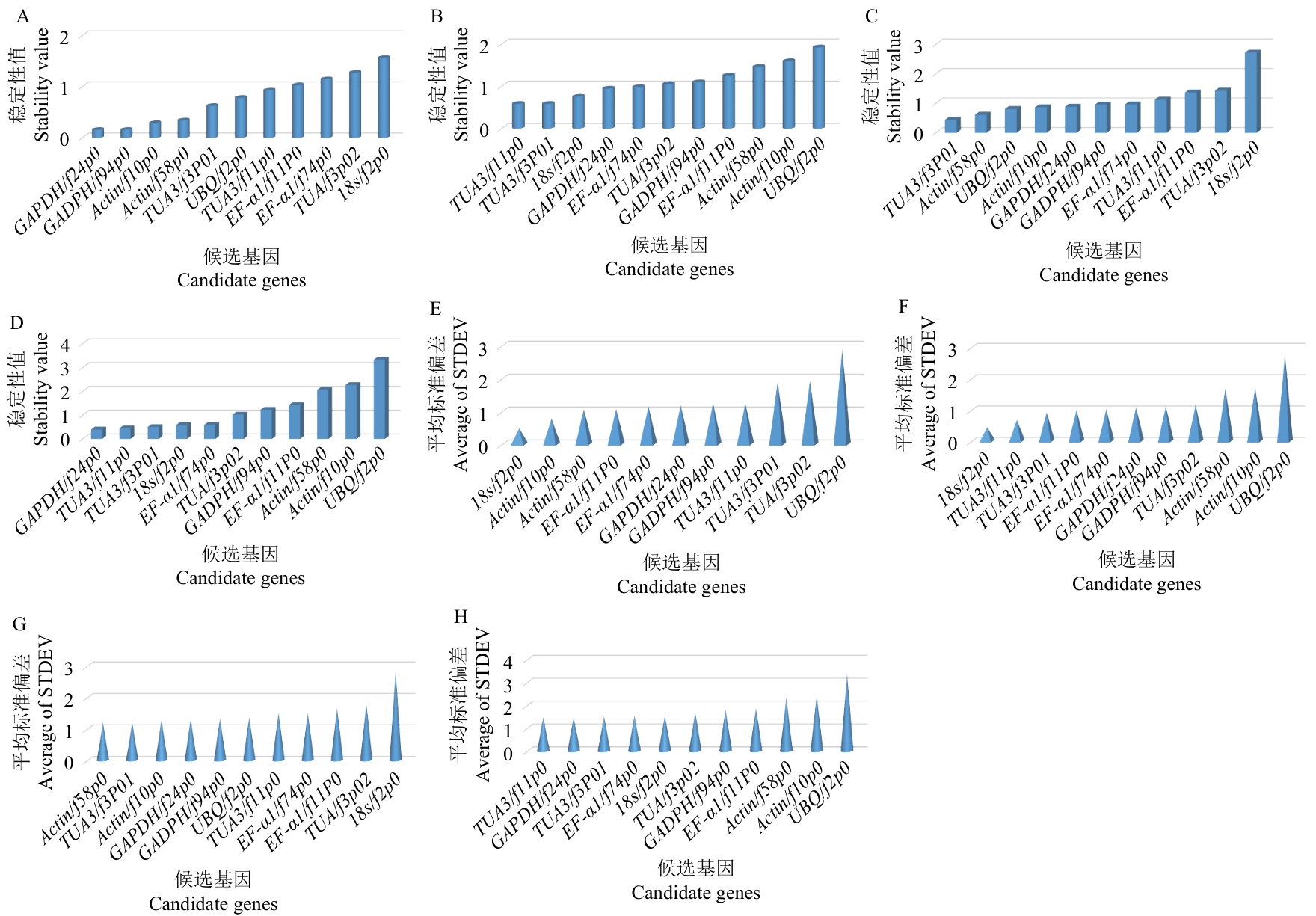
图5 石仙桃中11个候选参考基因的表达稳定性A、B:geNorm对不同组织(A)和非生物胁迫下(B)内参候选基因表达稳定性分析;C、D:NormFinder对不同组织(C)和非生物胁迫下(D)内参候选基因表达稳定性分析;E、F:BestKeeper对不同组织(E)和非生物胁迫下(F)内参候选基因表达稳定性分析;G、H:ΔCt对不同组织(G)和非生物胁迫下(H)内参候选基因表达稳定性分析
Fig. 5 Expression stabilities of 11 candidate reference genes in P. chinensisA, B: Stability analysis of geNorm on the expressions of internal candidate reference genes in different tissues (A) and under abiotic stress (B). C, D: Stability analysis of NormFinder on the expressions of internal candidate reference genes in different tissues (C) and under abiotic stress (D). E, F: Stability analysis of BestKeeper on the expressions of internal candidate reference genes in different tissues (E) and under abiotic stress (F). G, H: Stability analysis of ΔCt on the expressions of internal candidate reference genes in different tissues (G) and under abiotic stress (H)
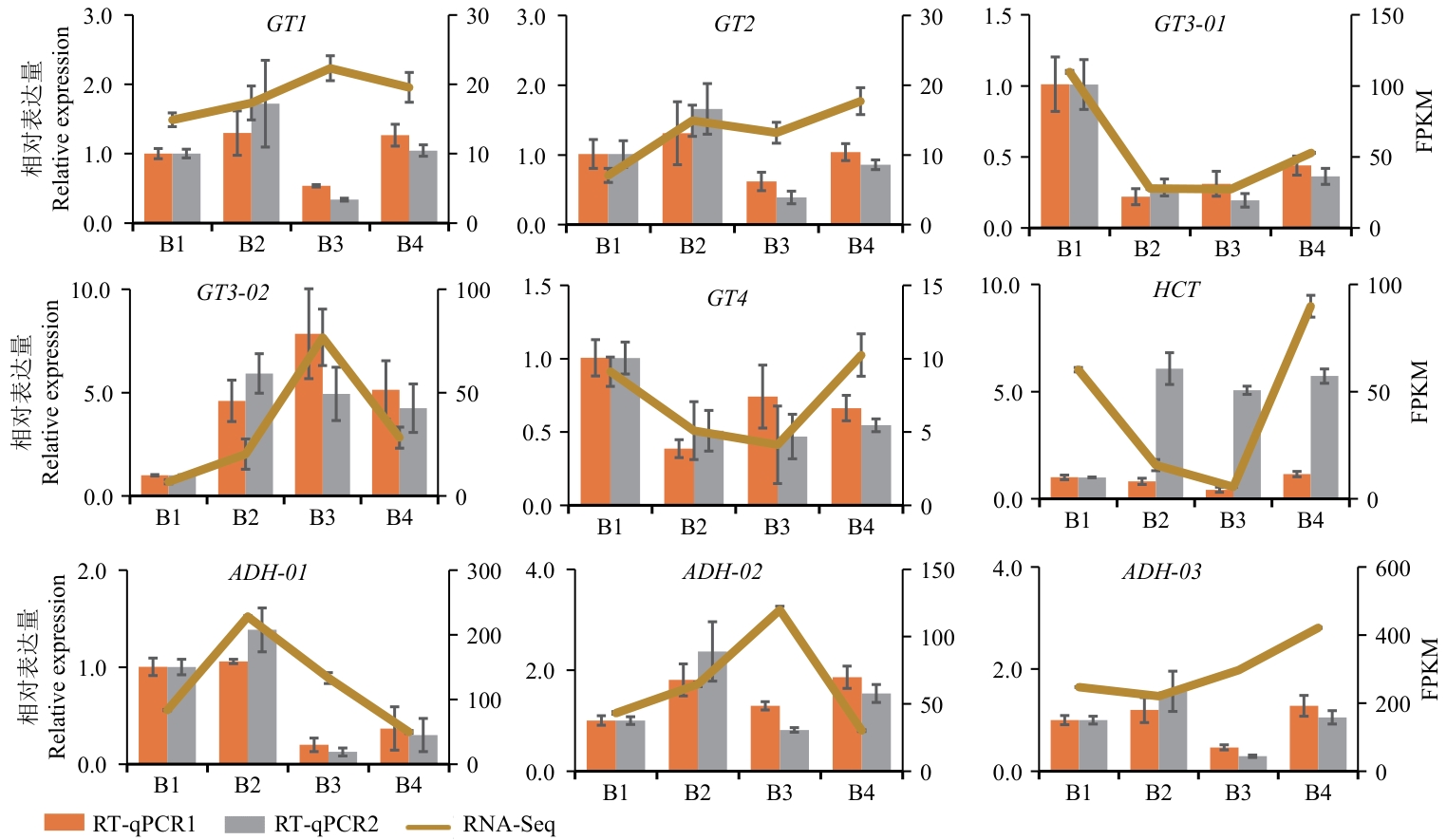
图6 石仙桃不同组织中ADH、HCT、GT基因家族的9个基因的RT-qPCR相对表达量和Illumina测序表达量RT-qPCR1:以Actin/f58p0为内参基因;RT-qPCR2:以TUA3/f11p为内参基因;RNA-Seq:用Illumina测序获得的表达量。B1:根;B2:根状茎;B3:假鳞茎;B4:叶
Fig. 6 RT-qPCR relative expressions and Illumina sequencing expressions of nine genes within ADH, HCT and GT gene families in different tissues of P. chinensisRT-qPCR1 with Actin/f58p0 as the internal reference gene; RT-qPCR2 with TUA3/f11p as the internal reference gene; RNA-Seq: sequencing with Illumina expression obtained. B1: Roots; B2: rhizomes; B3: pseudobulbs; B4: leaves
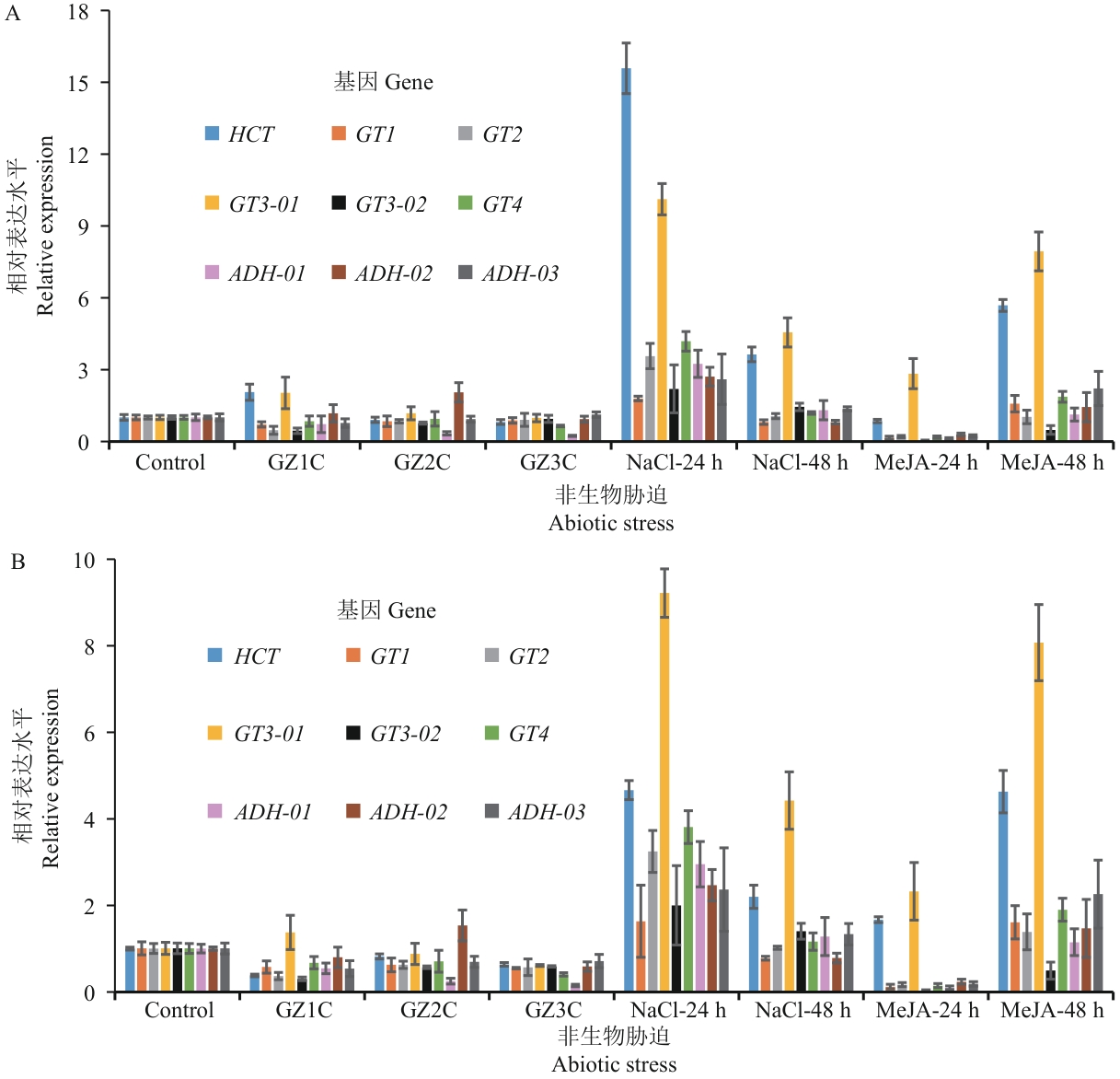
图7 石仙桃非生物胁迫中ADH、HCT、GT基因家族的9个基因的RT-qPCR相对表达量A:以Actin/f58p0为内参基因;B:以TUA3/f11p为内参基因;Control:对照;GZ1C:1层遮阳网;GZ2C:2层遮阳网;GZ3C:3层遮阳网;NaCl-24 h:氯化钠处理后24 h;NaCl-48 h:氯化钠处理后48 h;MeJA-24 h:茉莉酸甲酯处理后24 h;MeJA-48 h:茉莉酸甲酯处理后48 h
Fig. 7 RT-qPCR relative expressions of nine genes on ADH, HCT, and GT gene families, in abiotic stress of P. chinensisA: Actin/f58p0 as the internal reference gene; B: TUA3/f11p as the internal reference gene. Control: Control; GZ1C: one layer of sunshade; GZ2C: two layers of sunshade; GZ3C: three layers of sunshade; NaCl-24 h: 24 h after NaCl treatment; NaCl-48 h: 48 h after NaCl treatment; MeJA-24 h: 24 h after methyl jasmonate treatment; MeJA-48 h: 48 h after methyl jasmonate treatment
| [1] | Zhang LY, Zhang RM, Hu XL, et al. Reference gene selection for qRT-PCR analysis in the shoots and roots of Camellia sinensis var. sinensis under nutritional stresses [J]. Sci Hortic, 2023, 320: 112237. |
| [2] | Yang C, Lin YS, Qiu ZW, et al. Reference gene selection for qRT-PCR normalization of gene expression analysis in Melaleuca bracteata F. Muell. under abiotic stresses and hormonal stimuli [J]. Sci Hortic, 2023, 319: 112184. |
| [3] | Moshchenskaya YL, Galibina NA, Korzhenevskiy MA, et al. High-quality RNA extraction and evaluation of reference genes for qPCR assay of Pinus sylvestris L. trunk tissues [J]. Russ J Dev Biol, 2023, 54(1): 24-36. |
| [4] | Paul S, Singh S, Chakrabarti A, et al. Selection and evaluation of appropriate reference genes for RT-qPCR based expression analysis in Candida tropicalis following azole treatment [J]. Sci Rep, 2020, 10(1): 1972. |
| [5] | Tang J, Li EJ, Liu JX, et al. Selection of reliable reference genes for gene expression normalization in Sagittaria trifolia [J]. Genes, 2023, 14(7): 1321. |
| [6] | Chen XQ, Wood JJ. Flora of China [M]. Beijing: Sciences Press, 2009, 335-339. |
| [7] | 吴晓婷, 刘凯良. 闽东传统畲药草本植物资源调查与分析 [J]. 亚太传统医药, 2025, 21(5): 193-199. |
| Wu XT, Liu KL. Investigation and analysis of native plant resources of traditional she medicinal herbs in eastern Fujian [J]. Asia Pac Tradit Med, 2025, 21(5): 193-199. | |
| [8] | Li DL, Zheng XL, Duan L, et al. Ethnobotanical survey of herbal tea plants from the traditional markets in Chaoshan, China [J]. J Ethnopharmacol, 2017, 205: 195-206. |
| [9] | 刘保财, 黄颖桢, 赵云青, 等. 野生石仙桃立地环境与生长特性研究 [J]. 福建农业学报, 2017, 32(8): 864-869. |
| Liu BC, Huang YZ, Zhao YQ, et al. Natural habitat and growth characteristics of wild Pholidota chinensis Lindl [J]. Fujian J Agric Sci, 2017, 32(8): 864-869. | |
| [10] | Liu L, Zou MJ, Zeng KW, et al. Chemical constituents and their antioxidant, anti-inflammatory and anti-acetylcholinesterase activities from Pholidota cantonensis [J]. Plant Foods Hum Nutr, 2021, 76(1): 105-110. |
| [11] | 吴秀彩, 梁爽, 粟正英. 石仙桃属植物化学成分和药理活性的研究概况 [J]. 中国民族民间医药, 2020, 29(4): 60-62. |
| Wu XC, Liang S, Su ZY. Advances in chemical constituents and pharmacological activity of Pholidota chinensis Lindl [J]. Chin J Ethnomed Ethnopharmacy, 2020, 29(4): 60-62. | |
| [12] | Luo DH, Wang ZJ, Li ZM, et al. Structure of an entangled heteropolysaccharide from Pholidota chinensis Lindl and its antioxidant and anti-cancer properties [J]. Int J Biol Macromol, 2018, 112: 921-928. |
| [13] | Liu BC, Chen JY, Zhang WJ, et al. The gastrodin biosynthetic pathway in Pholidota chinensis Lindl. revealed by transcriptome and metabolome profiling [J]. Front Plant Sci, 2022, 13: 1024239. |
| [14] | 刘保财, 张武君, 赵云青, 等. 石仙桃中4种转录因子鉴定及其组织表达的研究 [J]. 生物技术通报, 2024, 40(5): 141-152. |
| Liu BC, Zhang WJ, Zhao YQ, et al. Identification and tissue expression of four transcription factors in Pholidota chinensis [J]. Biotechnol Bull, 2024, 40(5): 141-152. | |
| [15] | Ruijter JM, Ramakers C, Hoogaars WMH, et al. Amplification efficiency: linking baseline and bias in the analysis of quantitative PCR data [J]. Nucleic Acids Res, 2009, 37(6): e45. |
| [16] | Larsen B, Fuller VL, Pollier J, et al. Identification of iridoid glucoside transporters in Catharanthus roseus [J]. Plant Cell Physiol, 2017, 58(9): 1507-1518. |
| [17] | 徐圆圆, 赵国春, 郝颖颖, 等. 无患子RT-qPCR内参基因的筛选与验证 [J]. 生物技术通报, 2022, 38(10): 80-89. |
| Xu YY, Zhao GC, Hao YY, et al. Reference genes selection and validation for RT-qPCR in Sapindus mukorossi [J]. Biotechnol Bull, 2022, 38(10): 80-89. | |
| [18] | Xie FL, Xiao P, Chen DL, et al. miRDeepFinder: a miRNA analysis tool for deep sequencing of plant small RNAs [J]. Plant Mol Biol, 2012, 80: 75-84. |
| [19] | Andersen CL, Jensen JL, Ørntoft TF. Normalization of real-time quantitative reverse transcription-PCR data: a model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets [J]. Cancer Res, 2004, 64(15): 5245-5250. |
| [20] | Vandesompele J, De Preter K, Pattyn F, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes [J]. Genome Biol, 2002, 3(7): RESEARCH0034. |
| [21] | Pfaffl MW, Tichopad A, Prgomet C, et al. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pair-wise correlations [J]. Biotechnol Lett, 2004, 26(6): 509-515. |
| [22] | Silver N, Best S, Jiang J, et al. Selection of housekeeping genes for gene expression studies in human reticulocytes using real-time PCR [J]. BMC Mol Biol, 2006, 7: 33. |
| [23] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method [J]. Methods, 2001, 25(4): 402-408. |
| [24] | Pei YF, Leng L, Sun W, et al. Whole-genome sequencing in medicinal plants: current progress and prospect [J]. Sci China Life Sci, 2024, 67(2): 258-273. |
| [25] | Chaturvedi T, Gupta AK, Lal RK, et al. March of molecular breeding techniques in the genetic enhancement of herbal medicinal plants: present and future prospects [J]. Nucleus, 2022, 65(3): 413-436. |
| [26] | Clark JW. Genome evolution in plants and the origins of innovation [J]. New Phytol, 2023, 240(6): 2204-2209. |
| [27] | Liu S, Zhang Q, Kollie L, et al. Molecular networks of secondary metabolism accumulation in plants: Current understanding and future challenges [J]. Ind Crops Prod, 2023, 201: 116901. |
| [28] | Lin Y, Liu GF, Rao Y, et al. Identification and validation of reference genes for qRT-PCR analyses under different experimental conditions in Allium wallichii [J]. J Plant Physiol, 2023, 281: 153925. |
| [29] | Wang QZ, Guo CQ, Yang SP, et al. Screening and verification of reference genes for analysis of gene expression in garlic (Allium sativum L.) under cold and drought stress [J]. Plants, 2023, 12(4): 763. |
| [30] | Zhang BB, Zhu HJ, Zhang LH, et al. Evaluation of quality characteristics of 15 Gastrodia elata germplasm resources based on HS-SPME-GC-MS, HPLC and electronic nose techniques [J]. Food Biosci, 2025, 71: 107158. |
| [31] | Cao H, Li H, Lu L, et al. Screening and validation of internal reference genes for quantitative real-time PCR analysis of leaf color mutants in Dendrobium officinale [J]. Genes, 2023, 14(5): 1112. |
| [32] | 陈龙, 张淼, 秦慧真, 等. 石仙桃及其混伪品的DNA条形码鉴定 [J/OL]. 分子植物育种, 2023. . |
| Chen L, Zhang M, Qin HZ, et al. DNA barcoding identification of Pholidota chinensis lindl. and its fakes [J/OL]. Mol Plant Breed, 2023. . | |
| [33] | Mehralian M, Bidabadi SS, Azad M, et al. Melatonin-mediated alleviation of drought stress by modulation of physio-biochemical and metabolic status in Dracocephalum kotschyi Boiss. (Lamiaceae) [J]. Ind Crops Prod, 2023, 204: 117321. |
| [34] | Nazir F, Jahan B, Iqbal N, et al. Methyl jasmonate influences ethylene formation, defense systems, nutrient homeostasis and carbohydrate metabolism to alleviate arsenic-induced stress in rice (Oryza sativa) [J]. Plant Physiol Biochem, 2023, 202: 107990. |
| [1] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [2] | 张驰昊, 刘晋囡, 董家乐, 晁跃辉. 紫花苜蓿酵母表达文库的构建及候选耐盐基因的筛选与鉴定[J]. 生物技术通报, 2026, 42(2): 239-249. |
| [3] | 付涵, 孙书豪, 张思晴, 艾妞, 于杨, 于连伟, 王琼琼, 韩晓玉, 施艳, 韩卫丽, 杨雪. 本氏烟BOI家族基因的全基因组鉴定与表达分析[J]. 生物技术通报, 2026, 42(2): 207-217. |
| [4] | 李勒松, 张金金, 贺佳蝶, 梁艳丽, 杨生超, 栗孟飞, 赵艳. 当归4-香豆酸∶辅酶A连接酶基因家族的鉴定与功能表征[J]. 生物技术通报, 2026, 42(2): 293-305. |
| [5] | 倪莹, 李雷, 汪进萱, 马波, 孟昕, 冷平生, 吴静, 胡增辉. 紫丁香So4CL的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 139-149. |
| [6] | 陈静欢, 房国楠, 朱文豪, 叶广继, 苏旺, 贺苗苗, 杨生龙, 周云. 马铃薯种质资源淀粉表征及相关基因表达分析[J]. 生物技术通报, 2026, 42(1): 170-183. |
| [7] | 陈强, 于璎霏, 张颖, 张冲. 茉莉酸甲酯对薄皮甜瓜‘绿宝石’采后冷害的调控[J]. 生物技术通报, 2025, 41(9): 105-114. |
| [8] | 于文杰, 范斯然, 高文丽, 邢宇, 秦岭. 板栗核黄素合成通路关键基因鉴定及功能验证[J]. 生物技术通报, 2025, 41(9): 232-241. |
| [9] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [10] | 李玉珍, 李梦丹, 张蔚, 彭婷. 基于月季扩展蛋白基因家族鉴定的野蔷薇RmEXPB2基因功能研究[J]. 生物技术通报, 2025, 41(9): 182-194. |
| [11] | 胡鑫, 罗正英, 刘新龙, 吴才文, 吴转娣. 果蔗中甘蔗线条花叶病毒脱除技术[J]. 生物技术通报, 2025, 41(9): 62-70. |
| [12] | 任睿斌, 司二静, 万广有, 汪军成, 姚立蓉, 张宏, 马小乐, 李葆春, 王化俊, 孟亚雄. 大麦条纹病菌GH17基因家族的鉴定及表达分析[J]. 生物技术通报, 2025, 41(8): 146-154. |
| [13] | 程雪, 付颖, 柴晓娇, 王红艳, 邓欣. 谷子LHC基因家族鉴定及非生物胁迫表达分析[J]. 生物技术通报, 2025, 41(8): 102-114. |
| [14] | 曾丹, 黄园, 王健, 张艳, 刘庆霞, 谷荣辉, 孙庆文, 陈宏宇. 铁皮石斛bZIP转录因子家族全基因组鉴定及表达分析[J]. 生物技术通报, 2025, 41(8): 197-210. |
| [15] | 韩燚, 侯昌林, 唐露, 孙璐, 谢晓东, 梁晨, 陈小强. 大麦HvERECTA基因的克隆及功能分析[J]. 生物技术通报, 2025, 41(7): 106-116. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||