








生物技术通报 ›› 2026, Vol. 42 ›› Issue (2): 239-249.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0844
收稿日期:2025-08-04
出版日期:2026-02-26
发布日期:2026-03-17
通讯作者:
晁跃辉,男,博士,教授,研究方向 :草类植物遗传育种;E-mail: chaoyuehui@bjfu.edu.cn作者简介:张驰昊,男,硕士研究生,研究方向 :草类植物遗传育种;E-mail: zzzch0127@126.com
基金资助:
ZHANG Chi-hao1( ), LIU Jin-nan1, DONG Jia-le2, CHAO Yue-hui1(
), LIU Jin-nan1, DONG Jia-le2, CHAO Yue-hui1( )
)
Received:2025-08-04
Published:2026-02-26
Online:2026-03-17
摘要:
目的 构建紫花苜蓿(Medicago sativa)盐诱导基因表达文库,筛选候选耐盐基因并进行验证。 方法 使用1% NaCl处理紫花苜蓿幼苗,收集植物材料并提取总RNA,反转录后构建基因表达酵母文库。将文库质粒与空白质粒(pYES2-NTB)转入酵母菌株,确定用于文库筛选的NaCl浓度及候选耐盐基因。挑选196个单克隆菌落进行耐盐性分析,并进行PCR鉴定和测序分析。选择其中4个候选基因进行转基因功能分析。 结果 获得高质量RNA,并成功建立紫花苜蓿盐诱导基因表达文库,文库库容为7.68×107 CFU/mL,总克隆数为1.536×108 CFU。文库菌液在SG/-Ura+1.5 mol/L NaCl条件下仍有单克隆酵母菌落存活,而对照组则在1.3 mol/L NaCl培养基上已无法存活。测序结果经序列比对后,鉴定出75个不同基因。筛选并获得4个候选耐盐基因Msa1025270、Msa0320180、Msa0819320、Msa0859120,通过NCBI网站查询得这4个基因的功能与脱落酸受体、ABA反应、脱落酸和环境胁迫诱导、脱水早期反应等蛋白有关。转基因功能鉴定结果显示,4个候选基因表达能够显著提高植物对NaCl的抵抗能力。 结论 文库菌液具有良好的耐盐抗性,候选耐盐基因转基因植株验证结果表明,Msa1025270、Msa0320180、Msa0819320、Msa0859120 4个基因具有耐盐相关功能。
张驰昊, 刘晋囡, 董家乐, 晁跃辉. 紫花苜蓿酵母表达文库的构建及候选耐盐基因的筛选与鉴定[J]. 生物技术通报, 2026, 42(2): 239-249.
ZHANG Chi-hao, LIU Jin-nan, DONG Jia-le, CHAO Yue-hui. Construction of Alfalfa Yeast Expression Library and Screening and Identification of Salt-tolerance Genes[J]. Biotechnology Bulletin, 2026, 42(2): 239-249.
| 引物名称 Primer name | 序列 Sequence (5′‒3′) | 用途 Usage |
|---|---|---|
| CDS III/3' PCR Primer | GACGATAAGGTACCTAAGGATCCACCGGG | 逆转录 Reverse transcription |
| SMART IV Oligonucleotide | GTGCTGGATATCTGCAGAATTCCATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTVN | |
| P1-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAGGG | ds cDNA扩增 ds cDNA amplification |
| P2-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAGGG | |
| P3-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAAGGG | |
| P4-R | ACTCGAGCGGCCGCCACTGTGCTGGATATCTGCAGAATTCCA | |
| pYES2-NTB-F | CAGCTGTAATACGACTCACTATAGG | 文库菌落PCR鉴定 PCR identification of library colonies |
| pYES2-NTB-R | AGGGTTAGGGATAGGCTTACCTTCG | |
| NtActin-F | AGGTGGAGACATGGGTGGTG | 烟草内参基因表达分析 Analysis of tobacco internal reference gene expression |
| NtActin-R | TCATTAGGCACACAGATCTCTG | |
| MsG5270-qRT-F | CGGCTTGCTAACTACCGTTCT | 基因表达分析 Gene expression analysis |
| MsG5270-qRT-R | TTGATTTGATCTCGTTGAGGC | |
| MsG0180-qRT-F | GCTGATGAAATCGTCCCA | |
| MsG0180-qRT-R | CAACCCTGTTCCTCCCAC | |
| MsG9320-qRT-F | CTCTTGGCTATGGTTCTTC | |
| MsG9320-qRT-R | TTGGCATCGTTTACTTCA | |
| MsG9120-qRT-F | TCCAATGTTCGTCCCACT | |
| MsG9120-qRT-R | CTCTTTCCCATCTTCGTG | |
| MsG5270-F | TCACCATTTACGAACGATACATGTTACCAAACCCAACAAC | 克隆及表达载体构建 Construction of cloning and expression vectors |
| MsG5270-R | ATGGTGATGCATTCCCGGGTTTATCGTTGATTTGATCTCG | |
| MsG9320-F | TCACCATTTACGAACGATACATGGATTCGAGAAAAGCAAT | |
| MsG9320-R | ATGGTGATGCATTCCCGGGTTCAATTCTGAGTGTTGTCCT | |
| MsG9120-F | TCACCATTTACGAACGATACATGGAAGTTCTTTCTGCTAC | |
| MsG9120-R | ATGGTGATGCATTCCCGGGTCTACCTGGGCTGATGAATAG | |
| MsG0180-F | TCACCATTTACGAACGATACATGGGTGTTTTTACTTTCAA | |
| MsG0180-R | ATGGTGATGCATTCCCGGGTCTAGTAATTAGGGTTTGCTA | |
| 3302Y-F | TGACGCACAATCCCACTATCCTT | 植物表达载体PCR鉴定 PCR identification of plant expression vector |
| NOS181-R | CGTATTAAATGTATAATTGCGGGAC |
表1 引物及序列
Table 1 Primers and sequences
| 引物名称 Primer name | 序列 Sequence (5′‒3′) | 用途 Usage |
|---|---|---|
| CDS III/3' PCR Primer | GACGATAAGGTACCTAAGGATCCACCGGG | 逆转录 Reverse transcription |
| SMART IV Oligonucleotide | GTGCTGGATATCTGCAGAATTCCATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTVN | |
| P1-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAGGG | ds cDNA扩增 ds cDNA amplification |
| P2-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAGGG | |
| P3-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAAGGG | |
| P4-R | ACTCGAGCGGCCGCCACTGTGCTGGATATCTGCAGAATTCCA | |
| pYES2-NTB-F | CAGCTGTAATACGACTCACTATAGG | 文库菌落PCR鉴定 PCR identification of library colonies |
| pYES2-NTB-R | AGGGTTAGGGATAGGCTTACCTTCG | |
| NtActin-F | AGGTGGAGACATGGGTGGTG | 烟草内参基因表达分析 Analysis of tobacco internal reference gene expression |
| NtActin-R | TCATTAGGCACACAGATCTCTG | |
| MsG5270-qRT-F | CGGCTTGCTAACTACCGTTCT | 基因表达分析 Gene expression analysis |
| MsG5270-qRT-R | TTGATTTGATCTCGTTGAGGC | |
| MsG0180-qRT-F | GCTGATGAAATCGTCCCA | |
| MsG0180-qRT-R | CAACCCTGTTCCTCCCAC | |
| MsG9320-qRT-F | CTCTTGGCTATGGTTCTTC | |
| MsG9320-qRT-R | TTGGCATCGTTTACTTCA | |
| MsG9120-qRT-F | TCCAATGTTCGTCCCACT | |
| MsG9120-qRT-R | CTCTTTCCCATCTTCGTG | |
| MsG5270-F | TCACCATTTACGAACGATACATGTTACCAAACCCAACAAC | 克隆及表达载体构建 Construction of cloning and expression vectors |
| MsG5270-R | ATGGTGATGCATTCCCGGGTTTATCGTTGATTTGATCTCG | |
| MsG9320-F | TCACCATTTACGAACGATACATGGATTCGAGAAAAGCAAT | |
| MsG9320-R | ATGGTGATGCATTCCCGGGTTCAATTCTGAGTGTTGTCCT | |
| MsG9120-F | TCACCATTTACGAACGATACATGGAAGTTCTTTCTGCTAC | |
| MsG9120-R | ATGGTGATGCATTCCCGGGTCTACCTGGGCTGATGAATAG | |
| MsG0180-F | TCACCATTTACGAACGATACATGGGTGTTTTTACTTTCAA | |
| MsG0180-R | ATGGTGATGCATTCCCGGGTCTAGTAATTAGGGTTTGCTA | |
| 3302Y-F | TGACGCACAATCCCACTATCCTT | 植物表达载体PCR鉴定 PCR identification of plant expression vector |
| NOS181-R | CGTATTAAATGTATAATTGCGGGAC |
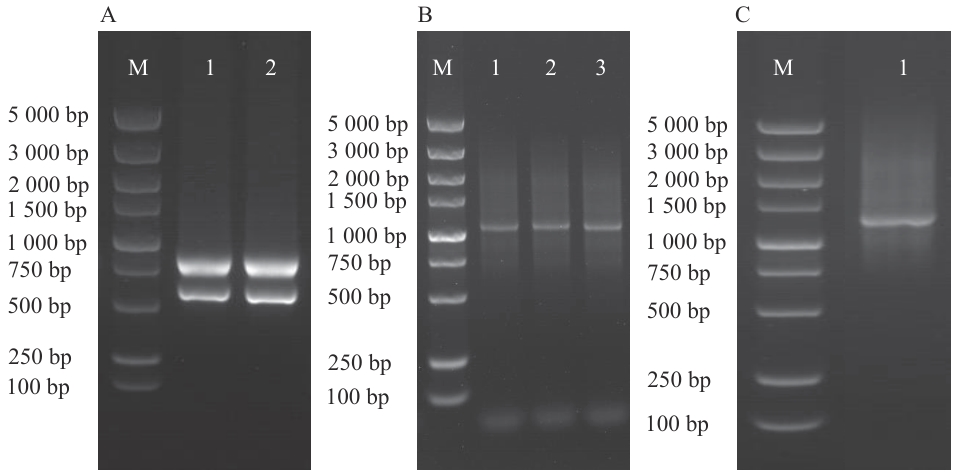
图1 RNA提取及合成ds cDNA的鉴定A:样品RNA提取(M:Marker;1-2:紫花苜蓿样品);B:合成ds cDNA扩增结果(M:Marker;1:P1-F/P4-R扩增ds cDNA;2:P2-F/P4-R扩增ds cDNA;3:P3-F/P4-R扩增ds cDNA);C:纯化与去除小片段(M:Marker,1:3种ds cDNA混合纯化后结果)
Fig. 1 RNA extraction and identification of synthesized ds cDNAA: Sample RNA extraction (M: Marker, 1‒2: alfalfa (Medicago sativa) samples). B: Amplified results of synthesized ds cDNA (M: Marker; 1: P1-F/P4-R amplified ds cDNA; 2: P2-F/P4-R amplified ds cDNA; 3: P3-F/P4-R amplified ds cDNA). C: Purified and removed small fragments (M: Marker; 1: results after the three ds cDNAs were mixed and purified)
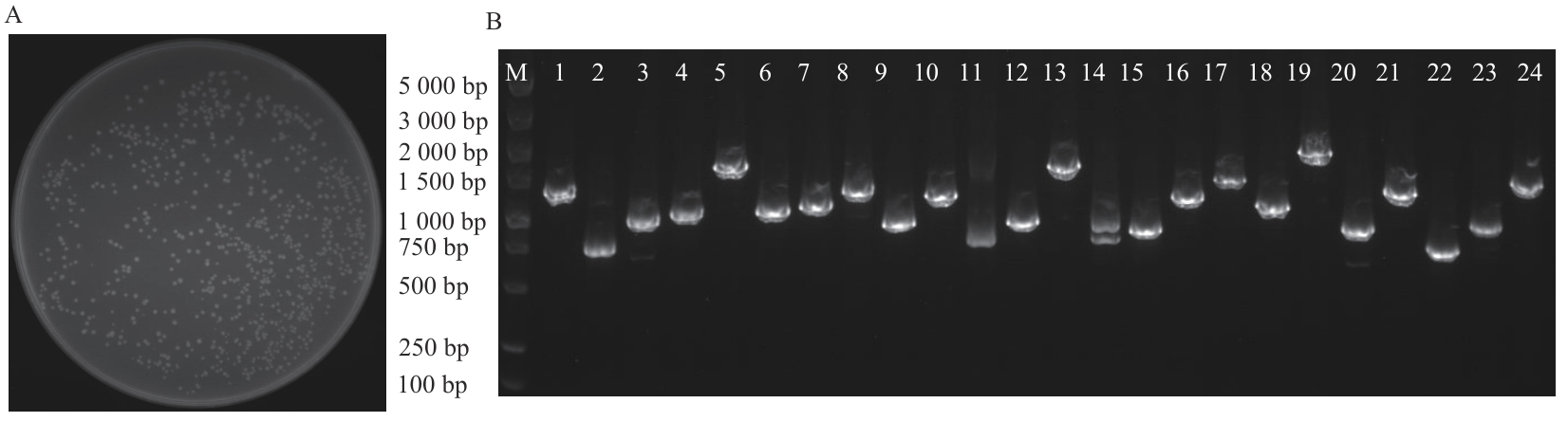
图2 大肠杆菌菌落计数及文库鉴定A:文库库容克隆计数;B:文库质量PCR鉴定(M:DNA marker;1‒24:24个不同单克隆菌落)
Fig. 2 Count and library identification of Escherichia coli colonyA: Library capacity and clone count. B: PCR identification of library quality (M: DNA marker; 1‒24: 24 different cloned colonies)
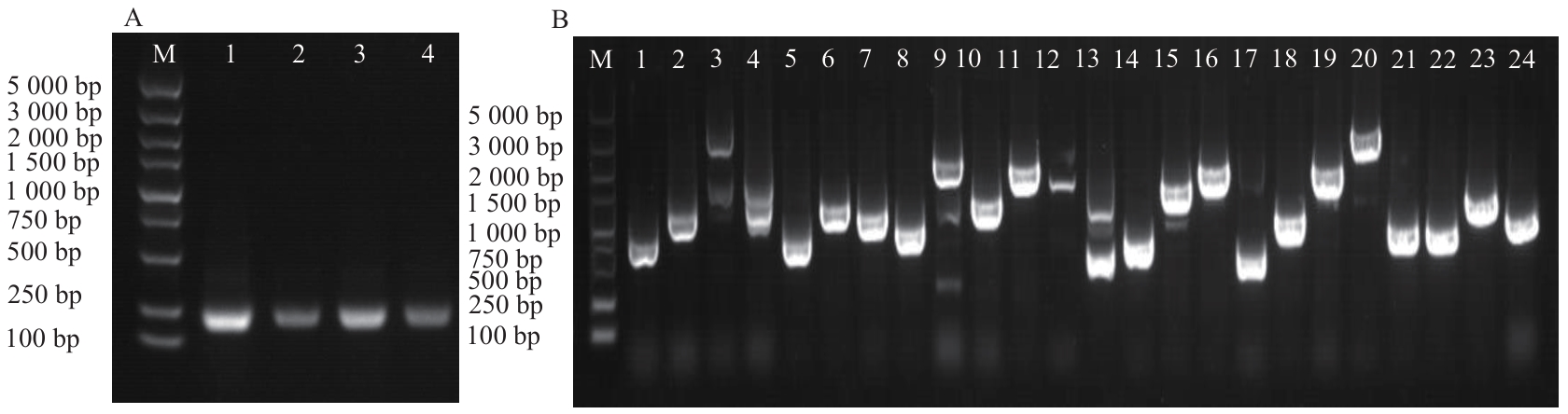
图3 空载(A)、文库菌液(B)酵母转化PCR验证图A中,M:Marker,1-4:空载酵母转化;图B中,M:DNA marker,1-24:酵母文库菌落
Fig. 3 PCR identification of yeast transformed with empty vector (A) and library cultures (B)In Fig. A, M: marker, 1-4: yeast transformation empty vector. In Fig. B, M: DNA marker, 1-24: yeast library colonies
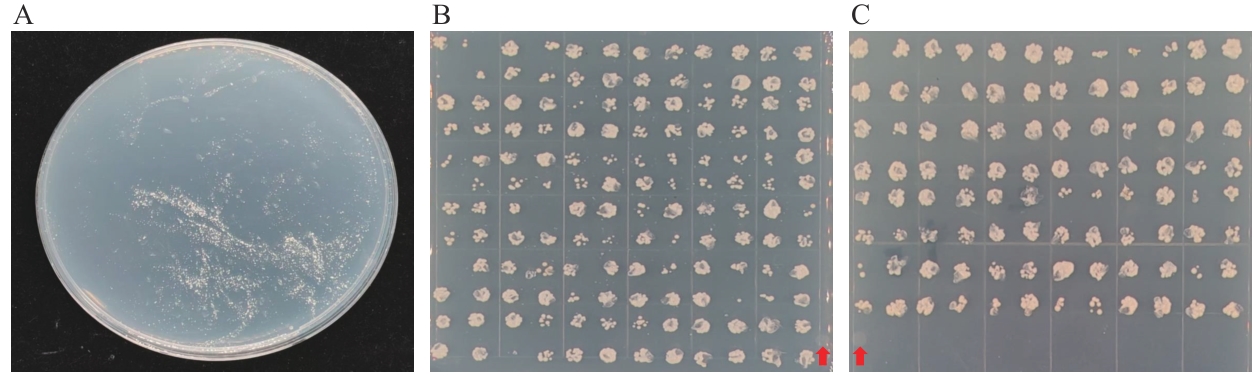

图5 阳性克隆菌板及回转验证结果A:文库菌液涂布于1.5 mol/L NaCl固体培养基结果图;B:单克隆菌落回转验证结果(144个);C:单克隆菌落回转验证结果(96个);:对照组
Fig. 5 Positive clone plate and results of rotation verificationA: The result of applying the library bacterial solution on 1.5 mol/L NaCl solid culture medium. B: Verification results of monoclonal colony rotation (144). C: Verification results of monoclonal colony rotation (96). : Control group
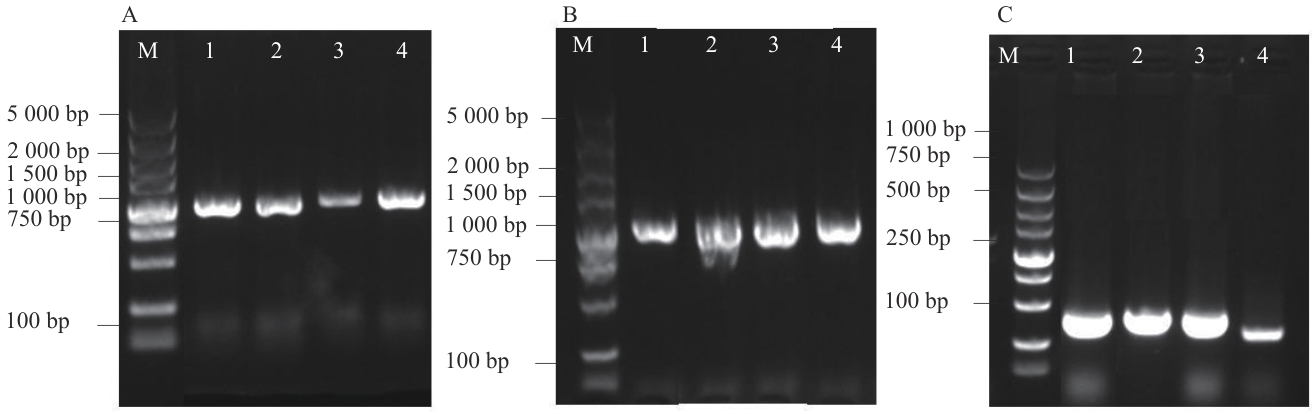
图8 检测载体菌落PCRA:载体构建菌落PCR;B:农杆菌转化菌落PCR;C:cDNA PCR鉴定
Fig. 8 Detection vector colony PCRA: Vector construction colony PCR (1: pBⅠ-MsG5270; 2: pBⅠ-MsG0180; 3: pBⅠ-MsG9320; 4: pBⅠ-MsG9120). B: Agrobacterium transformed colony PCR (1: pBⅠ-MsG5270; 2: pBⅠ- MsG0180; 3: pBⅠ-MsG9320; 4: pBⅠ-MsG9120). C: cDNA PCR identification (1: pBⅠ-MsG5270; 2: pBⅠ-MsG0180; 3: pBⅠ-MsG9320; 4: pBⅠ-MsG9120)
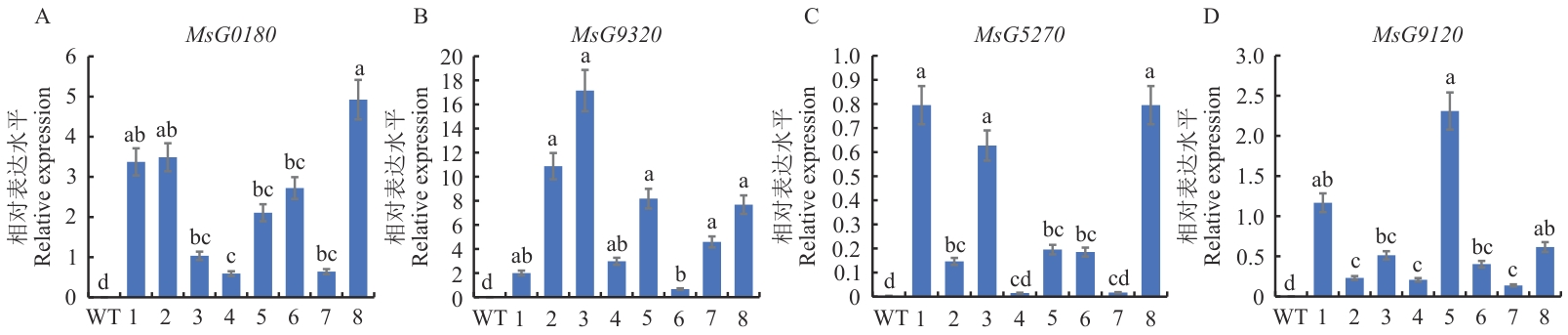
图9 MsG0180、MsG9320、MsG5270、MsG9120表达分析不同字母表示P<0.05水平差异显著,数值为平均值±标准误(n=3). WT:野生型;1‒8:转基因植株
Fig. 9 Expression analysis of MsG0180, MsG9320, MsG5270, and MsG9120Different letters indicate significant differences at P<0.05 level. The data are in means ±SE (n=3), WT: Wild type; 1‒8: transgenic plants
| [1] | Gong ZZ. Plant abiotic stress: New insights into the factors that activate and modulate plant responses [J]. J Integr Plant Biol, 2021, 63(3): 429-430. |
| [2] | Joshi S, Nath J, Singh AK, et al. Ion transporters and their regulatory signal transduction mechanisms for salinity tolerance in plants [J]. Physiol Plant, 2022, 174(3): e13702. |
| [3] | Liang XY, Li JF, Yang YQ, et al. Designing salt stress-resilient crops: Current progress and future challenges [J]. J Integr Plant Biol, 2024, 66(3): 303-329. |
| [4] | Ali Aazami M, Rasouli F, Ebrahimzadeh A. Oxidative damage, antioxidant mechanism and gene expression in tomato responding to salinity stress under in vitro conditions and application of iron and zinc oxide nanoparticles on callus induction and plant regeneration [J]. BMC Plant Biol, 2021, 21(1): 597. |
| [5] | van Zelm E, Zhang YX, Testerink C. Salt tolerance mechanisms of plants [J]. Annu Rev Plant Biol, 2020, 71: 403-433. |
| [6] | Soltabayeva A, Ongaltay A, Omondi JO, et al. Morphological, physiological and molecular markers for salt-stressed plants [J]. Plants, 2021, 10(2): 243. |
| [7] | Prăvălie R, Patriche C, Borrelli P, et al. Arable lands under the pressure of multiple land degradation processes. A global perspective [J]. Environ Res, 2021, 194: 110697. |
| [8] | Popova LG, Khramov DE, Nedelyaeva OI, et al. Yeast heterologous expression systems for the study of plant membrane proteins [J]. Int J Mol Sci, 2023, 24(13): 10768. |
| [9] | Du LY, Huang XL, Ding L, et al. TaERF87 and TaAKS1 synergistically regulate TaP5CS1/TaP5CR1-mediated proline biosynthesis to enhance drought tolerance in wheat [J]. New Phytol, 2023, 237(1): 232-250. |
| [10] | Kumari S, Singh P, Singla-Pareek SL, et al. Heterologous expression of a salinity and developmentally regulated rice cyclophilin gene (OsCyp2) in E. coli and S. cerevisiae confers tolerance towards multiple abiotic stresses [J]. Mol Biotechnol, 2009, 42(2): 195-204. |
| [11] | Kumar R, Mustafiz A, Sahoo KK, et al. Functional screening of cDNA library from a salt tolerant rice genotype Pokkali identifies mannose-1-phosphate guanyl transferase gene (OsMPG1) as a key member of salinity stress response [J]. Plant Mol Biol, 2012, 79(6): 555-568. |
| [12] | Lee JH, Van Montagu M, Verbruggen N. A highly conserved kinase is an essential component for stress tolerance in yeast and plant cells [J]. Proc Natl Acad Sci USA, 1999, 96(10): 5873-5877. |
| [13] | Abulfaraj AA, Alshareef SA. Concordant gene expression and alternative splicing regulation under abiotic stresses in Arabidopsis [J]. Genes, 2024, 15(6): 675. |
| [14] | Yamanaka T, Miyama M, Tada Y. Transcriptome profiling of the mangrove plant Bruguiera gymnorhiza and identification of salt tolerance genes by Agrobacterium functional screening [J]. Biosci Biotechnol Biochem, 2009, 73(2): 304-310. |
| [15] | Song A, Lu J, Jiang J, et al. Isolation and characterisation of Chrysanthemum crassum SOS1, encoding a putative plasma membrane Na+/H+ antiporter [J]. Plant Biol, 2012, 14(5): 706-713. |
| [16] | Kappachery S, Yu JW, Baniekal-Hiremath G, et al. Rapid identification of potential drought tolerance genes from Solanum tuberosum by using a yeast functional screening method [J]. Comptes Rendus Biol, 2013, 336(11/12): 530-545. |
| [17] | Eswaran N, Parameswaran S, Sathram B, et al. Yeast functional screen to identify genetic determinants capable of conferring abiotic stress tolerance in Jatropha curcas . [J]. BMC Biotechnol, 2010, 10: 23. |
| [18] | Obata T, Kitamoto HK, NakamUra A, et al. Rice shaker potassium channel OsKAT1 confers tolerance to salinity stress on yeast and rice cells [J]. Plant Physiol, 2007, 144(4): 1978-1985. |
| [19] | Burezq H. Reproductive ecology of forage alfalfa (Medicago sativa L.): recent advances [M]//Plant Reproductive Ecology-Recent Advances. London: IntechOpen, 2022. |
| [20] | 王馨磊, 朱思琪, 田晓娜, 等. 紫花苜蓿RALF基因家族的全基因组鉴定及盐胁迫下表达分析 [J]. 江苏农业学报, 2025, 41(7): 1260-1269. |
| Wang XL, Zhu SQ, Tian XN, et al. Genome-wide identification of the RALF gene family in Medicago sativa and expression analysis under salt stress [J]. Jiangsu J Agric Sci, 2025, 41(7): 1260-1269. | |
| [21] | 杭嘉慧. 紫花苜蓿褪黑素合成基因MsSNAT1和MsASMT1调控耐盐性机制研究 [D]. 银川: 宁夏大学, 2025. |
| Hang JH. Regulation mechanism of melatonin synthesis genes MsSNAT1 and MsASMT1 in salt tolerance of alfalfa [D]. Yinchuan: Ningxia University, 2025. | |
| [22] | 沙吾列·沙比汗, 赛里克·都曼, 张丽萍, 等. 20份苜蓿种质材料苗期耐盐性综合评价 [J]. 现代农业科技, 2016(8): 247-250, 262. |
| Shawulie·S, Sailike·D, Zhang LP, et al. A comprehensive evaluation on salt tolerance of 20 alfalfa germplasm materials in seedling stage [J]. XianDai NongYe KeJi, 2016(8): 247-250, 262. | |
| [23] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method [J]. Methods, 2001, 25(4): 402-408. |
| [24] | Li YK, Jiang FL, Niu LF, et al. Synergistic regulation at physiological, transcriptional and metabolic levels in tomato plants subjected to a combination of salt and heat stress [J]. Plant J, 2024, 117(6): 1656-1675. |
| [25] | Afzal M, El Sayed Hindawi S, Alghamdi SS, et al. Potential breeding strategies for improving salt tolerance in crop plants [J]. J Plant Growth Regul, 2023, 42(6): 3365-3387. |
| [26] | Ashraf M, Munns R. Evolution of approaches to increase the salt tolerance of crops [J]. Crit Rev Plant Sci, 2022, 41(2): 128-160. |
| [27] | Rasheed A, Raza A, Jie HD, et al. Molecular tools and their applications in developing salt-tolerant soybean (Glycine max L.) cultivars [J]. Bioengineering, 2022, 9(10): 495. |
| [28] | Fan XR, Jiang HZ, Meng LJ, et al. Gene mapping, cloning and association analysis for salt tolerance in rice [J]. Int J Mol Sci, 2021, 22(21): 11674. |
| [29] | Kausar R, Komatsu S. Proteomic approaches to uncover salt stress response mechanisms in crops [J]. Int J Mol Sci, 2022, 24(1): 518. |
| [30] | Atta K, Mondal S, Gorai S, et al. Impacts of salinity stress on crop plants: improving salt tolerance through genetic and molecular dissection [J]. Front Plant Sci, 2023, 14: 1241736. |
| [31] | Fidler J, Graska J, Gietler M, et al. PYR/PYL/RCAR receptors play a vital role in the abscisic-acid-dependent responses of plants to external or internal stimuli [J]. Cells, 2022, 11(8): 1352. |
| [32] | Fan WQ, Zhao MY, Li SX, et al. Contrasting transcriptional responses of PYR1/PYL/RCAR ABA receptors to ABA or dehydration stress between maize seedling leaves and roots [J]. BMC Plant Biol, 2016, 16: 99. |
| [33] | Zhao Y, Chan ZL, Gao JH, et al. ABA receptor PYL9 promotes drought resistance and leaf senescence [J]. Proc Natl Acad Sci USA, 2016, 113(7): 1949-1954. |
| [34] | Krishnaswamy SS, Srivastava S, Mohammadi M, et al. Transcriptional profiling of pea ABR17 mediated changes in gene expression in Arabidopsis thaliana . [J]. BMC Plant Biol, 2008, 8: 91. |
| [35] | Luo M, Liu JH, Mohapatra S, et al. Characterization of a gene family encoding abscisic acid- and environmental stress-inducible proteins of alfalfa [J]. J Biol Chem, 1992, 267(22): 15367-15374. |
| [36] | Wang XS, Huo ZC, Ma L, et al. The salt and ABA inducible transcription factor gene, SlAITR3, negatively regulates tomato salt tolerance [J]. Plant Physiol Biochem, 2025, 222: 109735. |
| [37] | 刘颖慧, 王天宇, 黎裕. 拟南芥脱水诱导早期应答基因研究进展 [J]. 中国农业大学学报, 2009, 14(3): 7-11. |
| Liu YH, Wang TY, Li Y. Review of early responsive genes expressed during dehydration in Arabidopsis sp [J]. J China Agric Univ, 2009, 14(3): 7-11. | |
| [38] | Kariola T, Brader G, Helenius E, et al. EARLY RESPONSIVE TO DEHYDRATION 15, a negative regulator of abscisic acid responses in Arabidopsis [J]. Plant Physiol, 2006, 142(4): 1559-1573. |
| [1] | 刘青媛, 吴洪启, 陈秀娥, 陈剑, 姜远泽, 何燕子, 喻奇伟, 刘仁祥. 转录因子NtMYB96a调控烟草耐旱性的功能研究[J]. 生物技术通报, 2026, 42(4): 239-250. |
| [2] | 王亚乐, 李雪君, 孙计平, 孙焕. 烟草豫浓香201对低温胁迫的生理响应及转录组分析[J]. 生物技术通报, 2026, 42(2): 228-238. |
| [3] | 刘梦茹, 夏琳, 刘瑞霞, 闫新可, 杨军, 张菁华, 武明珠. 烟草多酚氧化酶NtPPO1-2基因克隆及功能分析[J]. 生物技术通报, 2026, 42(2): 218-227. |
| [4] | 张驰昊, 刘晋囡, 晁跃辉. 蒺藜苜蓿bZIP转录因子MtbZIP29的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 241-250. |
| [5] | 翟莹, 计俊杰, 陈炯辛, 于海伟, 李珊珊, 赵艳, 马天意. 异源过表达大豆GmNF-YB24提高转基因烟草抗旱性[J]. 生物技术通报, 2025, 41(8): 137-145. |
| [6] | 黄旭升, 周雅莉, 柴旭东, 闻婧, 王计平, 贾小云, 李润植. 紫苏质体型PfLPAT1B基因的克隆及其在油脂合成中的功能分析[J]. 生物技术通报, 2025, 41(7): 226-236. |
| [7] | 董栩焜, 车永梅, 王明硕, 罗政刚, 管恩森, 赵方贵, 叶青, 刘新. 纳米硅和蜡样芽胞杆菌SS1共处理对烟草生长的影响[J]. 生物技术通报, 2025, 41(7): 292-298. |
| [8] | 王伟伟, 赵振杰, 王志, 邹景伟, 罗政辉, 张玉杰, 钮力亚, 于亮, 杨学举. 盐胁迫下与小麦生理响应相关的耐盐基因研究进展[J]. 生物技术通报, 2025, 41(5): 14-22. |
| [9] | 卢勇杰, 夏海乾, 李永铃, 张文建, 余婧, 赵会纳, 王兵, 许本波, 雷波. 烟草AP2/ERF转录因子NtESR2的克隆及功能分析[J]. 生物技术通报, 2025, 41(4): 266-277. |
| [10] | 季盈彤, 高嘉宁, 钱孟英, 潘宁, 张筠, 崔红, 闫筱筱. NtMYC2调控烟草表皮毛发育的功能研究[J]. 生物技术通报, 2025, 41(4): 278-288. |
| [11] | 赵春夺, 李玉娥, 刘友杰, 王新航, 赵伟, 黄永成, 李虎林, 姬文秀. 轮作与连作对烟草根际土壤养分、酶活性及微生物群落结构的影响[J]. 生物技术通报, 2025, 41(4): 312-322. |
| [12] | 马耀武, 张麒宇, 杨淼, 蒋诚, 张振宇, 张伊琳, 李梦莎, 许嘉阳, 张斌, 崔光周, 姜瑛. 烟草根际促生菌的筛选鉴定及促生性能研究[J]. 生物技术通报, 2025, 41(3): 271-281. |
| [13] | 韩梦荞, 吴疆, 李丽华, 王昭懿, 邓习, 韦凤杰, 任民, 孙洋洋, 李富欣. 烟草染色体制备体系的建立与优化[J]. 生物技术通报, 2025, 41(3): 44-50. |
| [14] | 冯小虎, 张文梅, 徐婧, 熊书斌, 王利兵, 宋文静. 一组靶向烟草青枯病的噬菌体鸡尾酒筛选与防控效果[J]. 生物技术通报, 2025, 41(11): 311-318. |
| [15] | 王碧成, 景海青, 万坤, 张莹莹, 丁家豪, 李润植, 薛金爱, 张海平. 大豆BCAT基因家族鉴定及GmBCAT3在干旱胁迫中的功能分析[J]. 生物技术通报, 2025, 41(10): 196-209. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||