








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 362-373.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0563
李天源( ), 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏(
), 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏( )
)
收稿日期:2025-05-31
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
穆霄鹏,男,博士,副教授,研究方向 :园艺植物种质利用与创新;E-mail: 15110671026@163.com作者简介:李天源,女,硕士研究生,研究方向 :园艺植物种质利用与创新;E-mail: 17735426816@163.com
基金资助:
LI Tian-yuan( ), QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng(
), QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng( )
)
Received:2025-05-31
Published:2026-03-26
Online:2026-04-23
摘要:
目的 研究欧李SPL基因家族成员的结构功能和表达分析,为后续深入研究欧李SPL基因家族的功能奠定基础。 方法 基于欧李全基因组和转录组数据,系统鉴定欧李SPL基因家族成员,并对ChSPL基因家族进行生物信息学分析以及表达模式分析。 结果 共鉴定出17个欧李SPL基因家族成员(ChSPL01‒ChSPL17),ChSPLs不均等地分布在7条染色体上,各亚族内的基因结构及保守结构域均表现出高度相似性,ChSPL14‒ChSPL16定位在细胞质,其余的家族成员则定位在细胞核。顺式作用元件分析表明,ChSPLs的光响应元件数量最多,其次为厌氧诱导响应元件(ARE)、脱落酸响应元件(ABRE)和茉莉酸甲酯响应元件(CGTCA-motif)。基于转录组和RT-qPCR分析表明,ChSPL01、ChSPL02、ChSPL06、ChSPL09和ChSPL11等多数ChSPLs在果实发育早期呈现较高的表达量,且随着果实成熟表达量逐渐下降,而ChSPL12、ChSPL14和ChSPL16随果实发育表达量逐渐上调,其中ChSPL12在果实成熟期表达量最高。ChSPLs还存在明显的组织特异性表达,ChSPL02、ChSPL05、ChSPL06、ChSPL10和ChSPL17等在茎和叶中表达量相对较高;ChSPL01、ChSPL04、ChSPL09在叶中表达量相对较高;而ChSPL14、ChSPL15和ChSPL16在花和果实中表达量相对较高。 结论 共鉴定出17个欧李ChSPL基因,推测ChSPL基因家族在果实发育中发挥重要作用,其中ChSPL12、ChSPL14和ChSPL16可能为促进果实成熟的关键候选基因,为未来研究ChSPL基因功能提供参考。
李天源, 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏. 欧李SPL基因家族的鉴定及在果实发育过程中的表达分析[J]. 生物技术通报, 2026, 42(3): 362-373.
LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development[J]. Biotechnology Bulletin, 2026, 42(3): 362-373.
基因名称 Gene name | 上游引物序列 Forward primer sequence (5′‒3′) | 下游引物序列 Reverse primer sequence (5′‒3′) |
|---|---|---|
| ChSPL01 | AAGAGGAGGAGGAGGAGGAG | TGCTTGACAAGATGGCCTTG |
| ChSPL02 | GCTGTGCTCTCTCTCTTCTGTCAA | TTGAGTCAGTGACTCCTGTGACG |
| ChSPL03 | CCAAGGAAAACGTAGTTGTCGTAGA | TCAAATTCAGGAAGCTGATGAAATC |
| ChSPL04 | TGGTGCTGATTTTGTTGAGGC | CACTTCTCTGCTTGACAACATGGT |
| ChSPL05 | GCCACTGCTCAGTCTAAAGC | TTCAACTTGGCAGTGTGGTG |
| ChSPL06 | CAAGAGAGGCAGAGGAGGAG | ACTCCTTTGCGTTCATCAGC |
| ChSPL07 | ATTCTGGAGCTGGGACTCAC | CGTTGGAGCTGTGAGGAAAG |
| ChSPL08 | GCAAACAGGCAAATGCTTCC | CTGAACTCCTGCTGAATGGC |
| ChSPL09 | GGGTGATAATAGTGGAAGAATTCGG | TGGCCACATACATCCTTCAACTG |
| ChSPL10 | CTTCTGCCTGCCAACAACTT | CTGAGCTGGCCAGATTGAAC |
| ChSPL11 | CATAACCAGCATCTCCAACTGC | AAGACCGGTATCAGCCTCAGC |
| ChSPL12 | CCAAGCAGATGCAGAGGTTC | GAGAGTGAGCCGTGAGGTAA |
| ChSPL13 | GGCCTGTGGAAGCCAAAAAG | TCTGCTTTCTGCAGGTGAGG |
| ChSPL14 | ATGCCAAGGATTACCACCGA | TATTATGGCCAGCCAAACGC |
| ChSPL15 | ATTGTGATGGTGGAGCAGGA | CAGGCACTGAACTGAGGGTA |
| ChSPL16 | GCGTTTGGCTGGCCATAATA | TGGCTAGGCTCCTCAGTAGA |
| ChSPL17 | CCTTCAACAACCTCGTCCCT | TAGATGGAGTGGAGCTGCTG |
| ChActin | TTCAAAGACCAGCTCATCTGTGG | CAATGCCAGGGAACATAGTGGA |
表1 RT-qPCR引物序列
Table 1 Sequences of primers used for RT-qPCR
基因名称 Gene name | 上游引物序列 Forward primer sequence (5′‒3′) | 下游引物序列 Reverse primer sequence (5′‒3′) |
|---|---|---|
| ChSPL01 | AAGAGGAGGAGGAGGAGGAG | TGCTTGACAAGATGGCCTTG |
| ChSPL02 | GCTGTGCTCTCTCTCTTCTGTCAA | TTGAGTCAGTGACTCCTGTGACG |
| ChSPL03 | CCAAGGAAAACGTAGTTGTCGTAGA | TCAAATTCAGGAAGCTGATGAAATC |
| ChSPL04 | TGGTGCTGATTTTGTTGAGGC | CACTTCTCTGCTTGACAACATGGT |
| ChSPL05 | GCCACTGCTCAGTCTAAAGC | TTCAACTTGGCAGTGTGGTG |
| ChSPL06 | CAAGAGAGGCAGAGGAGGAG | ACTCCTTTGCGTTCATCAGC |
| ChSPL07 | ATTCTGGAGCTGGGACTCAC | CGTTGGAGCTGTGAGGAAAG |
| ChSPL08 | GCAAACAGGCAAATGCTTCC | CTGAACTCCTGCTGAATGGC |
| ChSPL09 | GGGTGATAATAGTGGAAGAATTCGG | TGGCCACATACATCCTTCAACTG |
| ChSPL10 | CTTCTGCCTGCCAACAACTT | CTGAGCTGGCCAGATTGAAC |
| ChSPL11 | CATAACCAGCATCTCCAACTGC | AAGACCGGTATCAGCCTCAGC |
| ChSPL12 | CCAAGCAGATGCAGAGGTTC | GAGAGTGAGCCGTGAGGTAA |
| ChSPL13 | GGCCTGTGGAAGCCAAAAAG | TCTGCTTTCTGCAGGTGAGG |
| ChSPL14 | ATGCCAAGGATTACCACCGA | TATTATGGCCAGCCAAACGC |
| ChSPL15 | ATTGTGATGGTGGAGCAGGA | CAGGCACTGAACTGAGGGTA |
| ChSPL16 | GCGTTTGGCTGGCCATAATA | TGGCTAGGCTCCTCAGTAGA |
| ChSPL17 | CCTTCAACAACCTCGTCCCT | TAGATGGAGTGGAGCTGCTG |
| ChActin | TTCAAAGACCAGCTCATCTGTGG | CAATGCCAGGGAACATAGTGGA |
基因 ID Gene ID | 基因 Gene | 染色体位置 Chr. No. | 编码氨基酸长度 Length (aa) | 分子量Molecular weight (kD) | 等电点 pI | 不稳定系数 Instability index | 脂肪族氨基酸指数 Aliphatic index | 亲水性 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|---|---|
| ouLi_011577-RA | SPL01 | Chr3 | 489 | 38.03 | 5.26 | 28.89 | 30.88 | 0.681 | Nucleus |
| ouLi_006828-RA | SPL02 | Chr2 | 1 152 | 95.94 | 5.01 | 58.50 | 23.96 | 0.800 | Nucleus |
| ouLi_019229-RA | SPL03 | Chr6 | 1 143 | 95.19 | 5.01 | 58.64 | 23.80 | 0.797 | Nucleus |
| ouLi_010488-RA | SPL04 | Chr3 | 489 | 38.23 | 5.24 | 19.82 | 26.99 | 0.648 | Nucleus |
| ouLi_002187-RA | SPL05 | Chr1 | 1 446 | 119.72 | 4.99 | 44.37 | 28.56 | 0.805 | Nucleus |
| ouLi_017116-RA | SPL06 | Chr5 | 1 230 | 100.61 | 5.01 | 45.43 | 24.07 | 0.786 | Nucleus |
| ouLi_024093-RA | SPL07 | Chr8 | 1 656 | 137.64 | 4.99 | 43.99 | 28.02 | 0.724 | Nucleus |
| ouLi_002823-RA | SPL08 | Chr1 | 1 497 | 124.27 | 5.01 | 42.47 | 29.33 | 0.743 | Nucleus |
| ouLi_013789-RA | SPL09 | Chr4 | 570 | 44.99 | 5.22 | 23.54 | 30.88 | 0.724 | Nucleus |
| ouLi_002175-RB | SPL10 | Chr1 | 951 | 78.33 | 5.08 | 38.32 | 28.08 | 0.802 | Nucleus |
| ouLi_024712-RA | SPL11 | Chr8 | 1 227 | 102.04 | 5.02 | 52.67 | 29.01 | 0.858 | Nucleus |
| ouLi_026147-RA | SPL12 | Chr8 | 3 213 | 265.03 | 4.84 | 42.23 | 27.23 | 0.725 | Cytoplasm/Nucleus |
| ouLi_012144-RA | SPL13 | Chr4 | 1 176 | 97.79 | 5.02 | 47.01 | 26.53 | 0.799 | Nucleus |
| ouLi_000956-RA | SPL14 | Chr1 | 3 105 | 255.36 | 4.86 | 41.47 | 28.28 | 0.716 | Cytoplasm |
| ouLi_001341-RA/RB | SPL15 | Chr1 | 2 451 | 202.05 | 4.91 | 39.97 | 27.46 | 0.696 | Cytoplasm |
| ouLi_000956-RD | SPL16 | Chr1 | 3 033 | 248.56 | 4.87 | 40.26 | 28.39 | 0.699 | Cytoplasm |
| ouLi_017119-RA | SPL17 | Chr5 | 1 137 | 97.28 | 4.94 | 62.14 | 21.37 | 1.115 | Nucleus |
表2 ChSPL家族成员的基本信息
Table 2 Basic information of ChSPL family members
基因 ID Gene ID | 基因 Gene | 染色体位置 Chr. No. | 编码氨基酸长度 Length (aa) | 分子量Molecular weight (kD) | 等电点 pI | 不稳定系数 Instability index | 脂肪族氨基酸指数 Aliphatic index | 亲水性 Grand average of hydropathicity | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|---|---|
| ouLi_011577-RA | SPL01 | Chr3 | 489 | 38.03 | 5.26 | 28.89 | 30.88 | 0.681 | Nucleus |
| ouLi_006828-RA | SPL02 | Chr2 | 1 152 | 95.94 | 5.01 | 58.50 | 23.96 | 0.800 | Nucleus |
| ouLi_019229-RA | SPL03 | Chr6 | 1 143 | 95.19 | 5.01 | 58.64 | 23.80 | 0.797 | Nucleus |
| ouLi_010488-RA | SPL04 | Chr3 | 489 | 38.23 | 5.24 | 19.82 | 26.99 | 0.648 | Nucleus |
| ouLi_002187-RA | SPL05 | Chr1 | 1 446 | 119.72 | 4.99 | 44.37 | 28.56 | 0.805 | Nucleus |
| ouLi_017116-RA | SPL06 | Chr5 | 1 230 | 100.61 | 5.01 | 45.43 | 24.07 | 0.786 | Nucleus |
| ouLi_024093-RA | SPL07 | Chr8 | 1 656 | 137.64 | 4.99 | 43.99 | 28.02 | 0.724 | Nucleus |
| ouLi_002823-RA | SPL08 | Chr1 | 1 497 | 124.27 | 5.01 | 42.47 | 29.33 | 0.743 | Nucleus |
| ouLi_013789-RA | SPL09 | Chr4 | 570 | 44.99 | 5.22 | 23.54 | 30.88 | 0.724 | Nucleus |
| ouLi_002175-RB | SPL10 | Chr1 | 951 | 78.33 | 5.08 | 38.32 | 28.08 | 0.802 | Nucleus |
| ouLi_024712-RA | SPL11 | Chr8 | 1 227 | 102.04 | 5.02 | 52.67 | 29.01 | 0.858 | Nucleus |
| ouLi_026147-RA | SPL12 | Chr8 | 3 213 | 265.03 | 4.84 | 42.23 | 27.23 | 0.725 | Cytoplasm/Nucleus |
| ouLi_012144-RA | SPL13 | Chr4 | 1 176 | 97.79 | 5.02 | 47.01 | 26.53 | 0.799 | Nucleus |
| ouLi_000956-RA | SPL14 | Chr1 | 3 105 | 255.36 | 4.86 | 41.47 | 28.28 | 0.716 | Cytoplasm |
| ouLi_001341-RA/RB | SPL15 | Chr1 | 2 451 | 202.05 | 4.91 | 39.97 | 27.46 | 0.696 | Cytoplasm |
| ouLi_000956-RD | SPL16 | Chr1 | 3 033 | 248.56 | 4.87 | 40.26 | 28.39 | 0.699 | Cytoplasm |
| ouLi_017119-RA | SPL17 | Chr5 | 1 137 | 97.28 | 4.94 | 62.14 | 21.37 | 1.115 | Nucleus |
蛋白 Protein | α-螺旋 Alpha helix (%) | 延伸链 Extended strand (%) | β-折叠 Beta turn (%) | 无规则卷曲 Random coil (%) |
|---|---|---|---|---|
| SPL01 | 31.48 | 5.56 | 0 | 62.96 |
| SPL02 | 9.40 | 4.96 | 0 | 85.64 |
| SPL03 | 11.05 | 6.05 | 0 | 82.89 |
| SPL04 | 31.48 | 5.56 | 0 | 62.96 |
| SPL05 | 9.36 | 4.57 | 0 | 86.07 |
| SPL06 | 11.98 | 6.36 | 0 | 81.66 |
| SPL07 | 12.52 | 3.63 | 0 | 83.85 |
| SPL08 | 9.04 | 4.02 | 0 | 86.95 |
| SPL09 | 28.57 | 7.41 | 0 | 64.02 |
| SPL10 | 10.76 | 6.96 | 0 | 82.28 |
| SPL11 | 10.78 | 6.13 | 0 | 83.09 |
| SPL12 | 28.04 | 5.23 | 0 | 66.73 |
| SPL13 | 9.72 | 6.14 | 0 | 84.14 |
| SPL14 | 27.37 | 7.93 | 0 | 64.70 |
| SPL15 | 28.80 | 8.46 | 0 | 62.75 |
| SPL16 | 25.64 | 4.95 | 0 | 69.41 |
| SPL17 | 11.64 | 6.88 | 0 | 81.48 |
表3 ChSPL蛋白的二级结构
Table 3 Secondary structure of ChSPL proteins
蛋白 Protein | α-螺旋 Alpha helix (%) | 延伸链 Extended strand (%) | β-折叠 Beta turn (%) | 无规则卷曲 Random coil (%) |
|---|---|---|---|---|
| SPL01 | 31.48 | 5.56 | 0 | 62.96 |
| SPL02 | 9.40 | 4.96 | 0 | 85.64 |
| SPL03 | 11.05 | 6.05 | 0 | 82.89 |
| SPL04 | 31.48 | 5.56 | 0 | 62.96 |
| SPL05 | 9.36 | 4.57 | 0 | 86.07 |
| SPL06 | 11.98 | 6.36 | 0 | 81.66 |
| SPL07 | 12.52 | 3.63 | 0 | 83.85 |
| SPL08 | 9.04 | 4.02 | 0 | 86.95 |
| SPL09 | 28.57 | 7.41 | 0 | 64.02 |
| SPL10 | 10.76 | 6.96 | 0 | 82.28 |
| SPL11 | 10.78 | 6.13 | 0 | 83.09 |
| SPL12 | 28.04 | 5.23 | 0 | 66.73 |
| SPL13 | 9.72 | 6.14 | 0 | 84.14 |
| SPL14 | 27.37 | 7.93 | 0 | 64.70 |
| SPL15 | 28.80 | 8.46 | 0 | 62.75 |
| SPL16 | 25.64 | 4.95 | 0 | 69.41 |
| SPL17 | 11.64 | 6.88 | 0 | 81.48 |
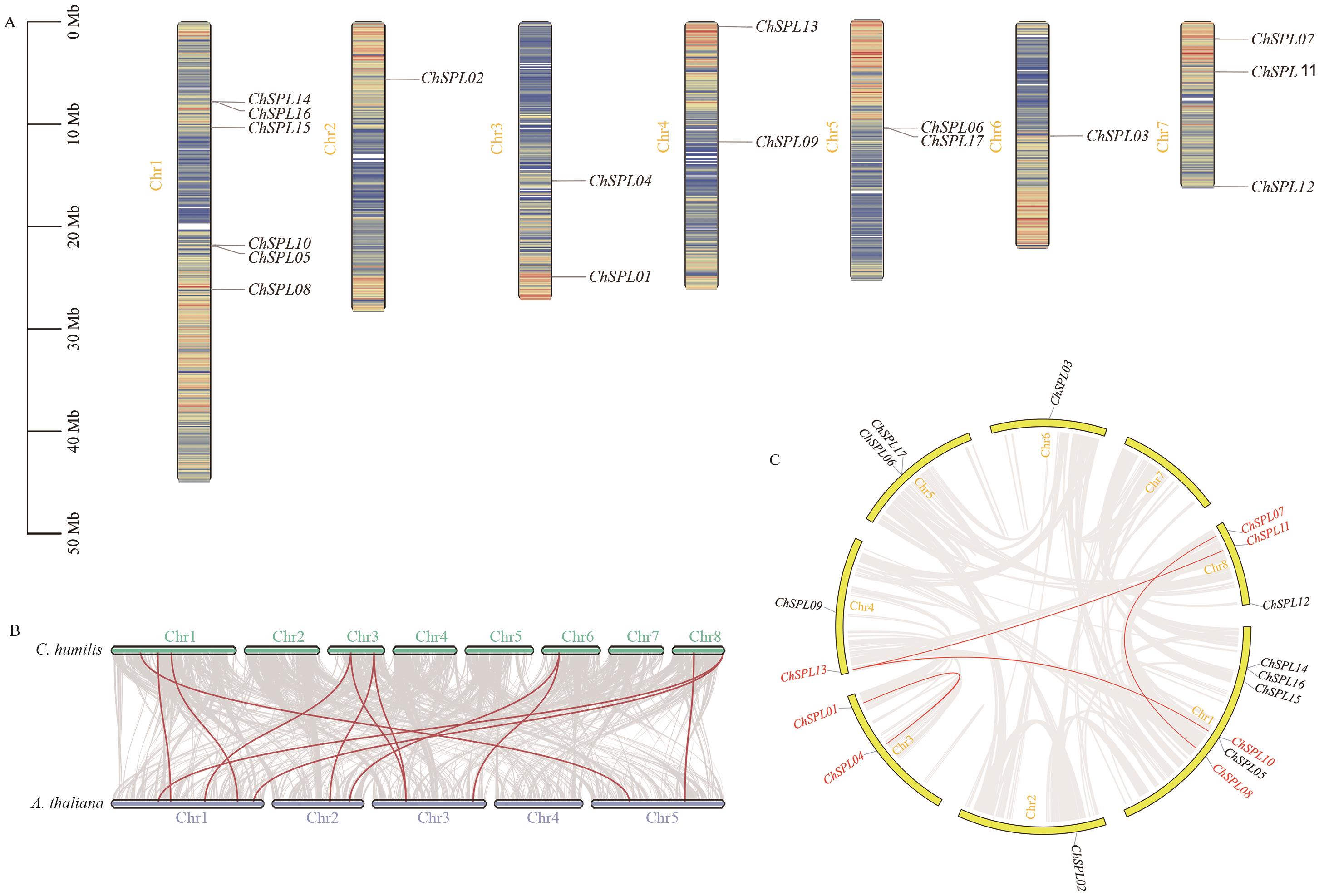
图4 ChSPL基因的染色体定位(A)与物种间共线性分析(B)、种内共线性分析(C)
Fig. 4 Chromosome location (A) and interspecies collinearity analysis (B), intraspecific collinearity analysis (C) of ChSPL gene
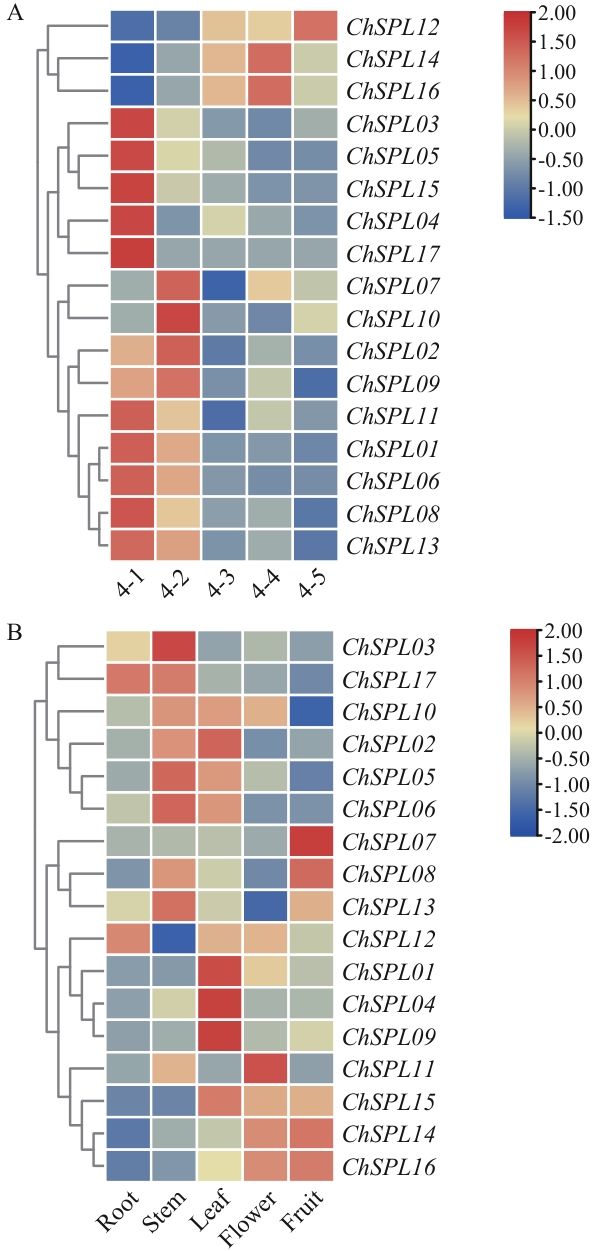
图7 ChSPL在果实不同发育时期(A)和不同组织中(B)的表达量4-1:幼果期;4-2:硬核期;4-3:转色期;4-4:着色期;4-5:成熟期
Fig. 7 Expressions of ChSPL in different stages of fruit (A) and different tissues (B)4-1: Young fruit stage. 4-2: Hard core stage. 4-3: Turning stage. 4-4: Coloring stage. 4-5: Mature stage
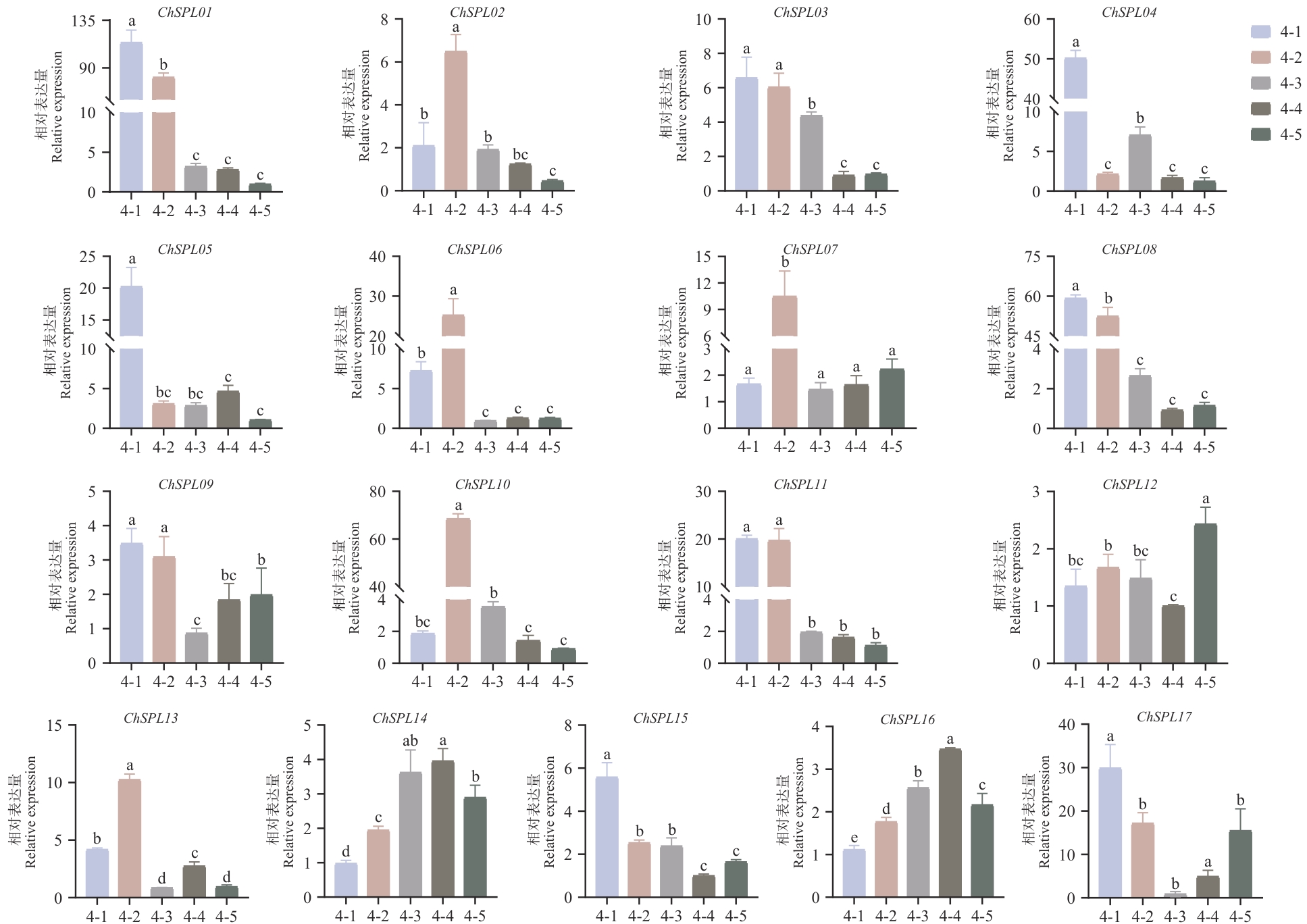
图8 ChSPL在果实中不同发育时期的表达量变化4-1:幼果期(花后6周);4-2:硬核期(花后8周);4-3:着色期(花后16周);4-4:转色期(花后17周);4-5:成熟期(花后18周),不同字母表示不同发育时期表达量差异显著(P<0.05)
Fig. 8 Changes in the expressions of ChSPL over time under different development stages4-1: Young fruit stage (6 weeks after flowering). 4-2: Stone hardening stage (8 weeks after flowering). 4-3: Coloration stage (16 weeks after flowering). 4-4: Color turning stage (17 weeks after flowering). 4-5: Maturity stage (18 weeks after flowering). Different letters indicate significant differences in expressions at different developmental stages (P<0.05)
| [1] | He AL, Zhou H, Ma C, et al. Genome-wide identification and expression analysis of the SPL gene family and its response to abiotic stress in barley (Hordeum vulgare L.) [J]. BMC Genomics, 2024, 25(1): 846. |
| [2] | Zhu DM, Geng XM, Zeng FY, et al. Identification and expression analysis of the SPL gene family during flower bud differentiation in Rhododendron molle [J]. Genes Genomics, 2025, 47(2): 171-182. |
| [3] | Li Y, Deng YF, Qin DB, et al. Study of the SPL gene family and miR156-SPL module in Populus tomentosa: Potential roles in juvenile-to-adult phase transition and reproductive phase [J]. Int J Biol Macromol, 2025, 296: 139547. |
| [4] | 赵一曼, 李景飞, 唐卓然, 等. 芥菜BjuSPL10基因表达及调控抽薹开花的功能解析 [J]. 园艺学报, 2025, 52(5): 1341-1350. |
| Zhao YM, Li JF, Tang ZR, et al. Functional identification and expression analysis of BjuSPL10 in regulating Brassica juncea bolting and flowering [J]. Acta Hortic Sin, 2025, 52(5): 1341-1350. | |
| [5] | 葛奇, 席会鹏. 植物SPL转录因子研究进展 [J]. 安徽农业科学, 2023, 51(23): 25-29, 53. |
| Ge Q, Xi HP. Research progress of SPL transcription factors in plants [J]. J Anhui Agric Sci, 2023, 51(23): 25-29, 53. | |
| [6] | Mouri IJ, Islam MS. A comprehensive in silico genome-wide identification and characterization of SQUAMOSA promoter binding protein (SBP) gene family in Musa acuminata [J]. J Genet Eng Biotechnol, 2025, 23(1): 100461. |
| [7] | Cheng HF, Li ZL, Sun BJ, et al. Genome-wide identification and expression analysis of the SPL gene family in Solanum melongena (eggplant) [J]. Agronomy, 2025, 15(2): 471. |
| [8] | Wang LY, Yu P, Lyu JY, et al. BZR1 physically interacts with SPL9 to regulate the vegetative phase change and cell elongation in Arabidopsis [J]. Int J Mol Sci, 2021, 22(19): 10415. |
| [9] | Zhang Y, Schwarz S, Saedler H, et al. SPL8, a local regulator in a subset of gibberellin-mediated developmental processes in Arabidopsis [J]. Plant Mol Biol, 2007, 63(3): 429-439. |
| [10] | Jin-Ying Gou FFF. Negative regulation of anthocyanin biosynthesis in Arabidopsis by a miR156-targeted SPL transcription factor [J]. Plant Cell, 2011, 23(4): 1512-1522. |
| [11] | Cui L, Zheng FY, Wang JF, et al. miR156a-targeted SBP-Box transcription factor SlSPL13 regulates inflorescence morphogenesis by directly activating SFT in tomato [J]. Plant Biotechnol J, 2020, 18(8): 1670-1682. |
| [12] | Wei W, Yang YY, Wu CJ, et al. MaSPL16 positively regulates fruit ripening in bananas via the direct transcriptional induction of MaNAC029 [J]. Hortic Adv, 2023, 1(1): 10. |
| [13] | Han YC, Gao HY, Chen HJ, et al. The involvement of papaya CpSBP1 in modulating fruit softening and carotenoid accumulation by repressing CpPME1/2 and CpPDS4 [J]. Sci Hortic, 2019, 256: 108582. |
| [14] | Li XY, Hou YM, Xie X, et al. A blueberry MIR156a-SPL12 module coordinates the accumulation of chlorophylls and anthocyanins during fruit ripening [J]. J Exp Bot, 2020, 71(19): 5976-5989. |
| [15] | Panda S, Chappell-Maor L, Alejandro de Haro L, et al. Molecular mechanisms driving the unusual pigmentation shift during eggplant fruit development [J]. Plant Commun, 2025, 6(5): 101321. |
| [16] | Liu HN, Shu Q, Kui LW, et al. The PyPIF5-PymiR156a-PySPL9-PyMYB114/MYB10 module regulates light-induced anthocyanin biosynthesis in red pear [J]. Mol Hortic, 2021, 1(1): 14. |
| [17] | 冯荦荦. 欧李多糖的制备、结构表征及对RAW 264.7细胞的免疫调节活性研究 [D]. 西安: 陕西师范大学, 2022. |
| Feng LL. Preparation, structural characterization and immunomodulatory activity of Prunus humilis polysaccharide on RAW 264.7 cells [D]. Xi’an: Shaanxi Normal University, 2022. | |
| [18] | 张莉, 郭炳成, 邵郅胜, 等. 欧李果实中花青素代谢物组分特征及抗氧化能力分析 [J/OL]. 分子植物育种, 2024. . |
| Zhang L, Guo BC, Shao ZS, et al. Analysis of the anthocyanin metabolite fractions and antioxidant capacity in Cerasus humilis fruit [J/OL]. Mol Plant Breed, 2024. . | |
| [19] | 杨钰, 闫松涛, 付鸿博, 等. 套袋遮光对欧李果实糖、酸和类黄酮含量的影响 [J]. 西北植物学报, 2024, 44(8): 1201-1207. |
| Yang Y, Yan ST, Fu HB, et al. Effects of bagging and shading on the content of sugar, acid and flavonoids in fruit of Cerasus humilis [J]. Acta Bot Boreali Occidentalia Sin, 2024, 44(8): 1201-1207. | |
| [20] | Bai YB, Liang MR, Cheng ZM, et al. Cloning, characterization, and ectopic expression in Arabidopsis to determine the function of FvSPL1 gene from woodland strawberry (Fragaria vesca) [J]. J Berry Res, 2022, 12(2): 267-278. |
| [21] | Liu XJ, Sun WX, Liu HB, et al. PavSPLs are key regulators of growth, development, and stress response in sweet cherry [J]. Plant Sci, 2025, 350: 112279. |
| [22] | 汪先菊, 张绍宇, 徐泽, 等. 桃SPL基因家族成员鉴定及其基因表达与果实发育的相关性分析 [J]. 农业生物技术学报, 2020, 28(9): 1521-1534. |
| Wang XJ, Zhang SY, Xu Z, et al. Identification of SPL gene family members and correlation analysis between gene expression and fruit development in peach (Prunus persica) [J]. J Agric Biotechnol, 2020, 28(9): 1521-1534. | |
| [23] | Li J, Hou HM, Li XQ, et al. Genome-wide identification and analysis of the SBP-box family genes in apple (Malus × domestica Borkh.) [J]. Plant Physiol Biochem, 2013, 70: 100-114. |
| [24] | Dong HX, Yan SL, Jing YX, et al. MIR156-targeted SPL9 is phosphorylated by SnRK2s and interacts with ABI5 to enhance ABA responses in Arabidopsis [J]. Front Plant Sci, 2021, 12: 708573. |
| [25] | Qin MM, Zhang Y, Yang YM, et al. Seed-specific overexpression of SPL12 and IPA1 improves seed dormancy and grain size in rice [J]. Front Plant Sci, 2020, 11: 532771. |
| [26] | Li HX, Wang SW, Zhai LL, et al. The miR156/SPL12 module orchestrates fruit colour change through directly regulating ethylene production pathway in blueberry [J]. Plant Biotechnol J, 2024, 22(2): 386-400. |
| [27] | Sun YT, Wang YY, Xiao YQ, et al. Genome-wide identification of the SQUAMOSA promoter-binding-binding protein-like (SPL) transcription factor family in sweet cherry fruit [J]. Int J Mol Sci, 2023, 24(3): 2880. |
| [28] | Lai TF, Wang XH, Ye BS, et al. Molecular and functional characterization of the SBP-box transcription factor SPL-CNR in tomato fruit ripening and cell death [J]. J Exp Bot, 2020, 71(10): 2995-3011. |
| [29] | Ling QH, Sadali NM, Soufi Z, et al. The chloroplast-associated protein degradation pathway controls chromoplast development and fruit ripening in tomato [J]. Nat Plants, 2021, 7(5): 655-666. |
| [30] | 汪先菊. 桃SPL基因家族的鉴定及PrupeSPL16, 17, 19在果实成熟软化过程中作用的研究 [D]. 杨凌: 西北农林科技大学, 2020. |
| Wang XJ. Identification of SPL gene family and analysis of the function of PrupeSPL16, 17, 19 in the process of fruit ripening and softening in peach [D]. Yangling: Northwest A & F University, 2020. |
| [1] | 尹跃, 秦小雅, 米佳, 安巍, 何军, 张锋锋. 枸杞FBN基因家族鉴定及与类胡萝卜素代谢的相关性分析[J]. 生物技术通报, 2026, 42(3): 338-348. |
| [2] | 罗威, 宫奥, 仲阳, 胡迪, 周洪源, 张泓欣, 艾菊, 罗有卫, 高冬丽. 黄瓜SEPALLATA2基因敲除对果实及疣状结构发育的多效性影响[J]. 生物技术通报, 2026, 42(3): 283-293. |
| [3] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [4] | 胡秋玲, 陈灵, 黄嘉怡, 赵梓乔, 潘璐怡, 刘慧丽, 刘太波. 多胺调控果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 203-212. |
| [5] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [6] | 张高翔, 吴玉碧, 郭亚静, 纪薇, 杨忠义. 葡萄WD40基因家族鉴定及表达量分析[J]. 生物技术通报, 2026, 42(3): 324-337. |
| [7] | 李成泉, 史庆华, 杨晓玉. 园艺作物果实发育的miRNA调控网络:从分子机制到种质创新[J]. 生物技术通报, 2026, 42(3): 19-36. |
| [8] | 李迎辉, 王杨博涵, 周浩博, 卢心如, 张珂欣, 于洋, 李传友, 孙传龙. 番茄VPE基因家族鉴定和抗逆功能分析[J]. 生物技术通报, 2026, 42(3): 263-274. |
| [9] | 陈静欢, 房国楠, 朱文豪, 叶广继, 苏旺, 贺苗苗, 杨生龙, 周云. 马铃薯种质资源淀粉表征及相关基因表达分析[J]. 生物技术通报, 2026, 42(1): 170-183. |
| [10] | 龙林茜, 曾银萍, 王茜, 邓玉萍, 葛敏茜, 陈彦灼, 李鑫娟, 杨军, 邹建. 向日葵GH3基因家族鉴定及其在花发育中的功能分析[J]. 生物技术通报, 2026, 42(1): 125-138. |
| [11] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [12] | 任睿斌, 司二静, 万广有, 汪军成, 姚立蓉, 张宏, 马小乐, 李葆春, 王化俊, 孟亚雄. 大麦条纹病菌GH17基因家族的鉴定及表达分析[J]. 生物技术通报, 2025, 41(8): 146-154. |
| [13] | 康琴, 汪霞, 谌明洋, 徐静天, 陈诗兰, 廖平杨, 许文志, 吴卫, 徐东北. 薄荷UV-B受体基因McUVR8的克隆与表达分析[J]. 生物技术通报, 2025, 41(8): 255-266. |
| [14] | 曾丹, 黄园, 王健, 张艳, 刘庆霞, 谷荣辉, 孙庆文, 陈宏宇. 铁皮石斛bZIP转录因子家族全基因组鉴定及表达分析[J]. 生物技术通报, 2025, 41(8): 197-210. |
| [15] | 韩燚, 侯昌林, 唐露, 孙璐, 谢晓东, 梁晨, 陈小强. 大麦HvERECTA基因的克隆及功能分析[J]. 生物技术通报, 2025, 41(7): 106-116. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||