








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 324-337.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0274
收稿日期:2025-03-14
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
纪薇,女,博士,教授,研究方向 :葡萄遗传种质研究与创新利用;E-mail: jiweiputao@163.com作者简介:张高翔,男,硕士研究生,研究方向 :园艺植物栽培生理与品质调控;E-mail: 1450735446@qq.com
基金资助:
ZHANG Gao-xiang( ), WU Yu-bi, GUO Ya-jing, JI Wei(
), WU Yu-bi, GUO Ya-jing, JI Wei( ), YANG Zhong-yi(
), YANG Zhong-yi( )
)
Received:2025-03-14
Published:2026-03-26
Online:2026-04-23
摘要:
目的 WD40超家族广泛存在于真核生物中,具有作为蛋白质复合物支架的基本亚基。WD40蛋白在植物发育和生理过程中发挥重要的调控作用。解析葡萄VvWD40基因家族成员的序列结构和理化性质,了解其在葡萄果实发育过程中的作用,为葡萄分子育种提供基因资源。 方法 基于葡萄全基因组信息,利用生物信息学方法对VvWD40家族进行鉴定,并对家族成员理化性质、染色体分布、基因结构、蛋白保守结构、系统进化、启动子顺式作用元件以及组织表达特性等进行分析,进一步利用转录组及RT-qPCR分析VvWD40基因在‘早黑宝’葡萄不同发育时期的表达模式。 结果 在葡萄基因组鉴定出86个WD40基因,通过系统发育分析将其分为7个组。在葡萄中共鉴定出12对共线基因对。Ka/Ks分析表明,它们在进化过程中主要经历了纯化选择。顺式作用元件分析表明,VvWD40基因家族含有激素及胁迫响应顺式作用元件,这些元件与葡萄果实发育密切相关。其中,VvWD40-62/84含有类黄酮合成元件,表明它们可能与花青素合成有关。‘早黑宝’葡萄4个不同阶段的表达谱分析显示,大多数VvWD40s在果实发育过程中表达。通过RT-qPCR验证8个基因在‘早黑宝’葡萄发育过程中表达量变化情况,VvWD40-34/49基因在葡萄果实发育初期表达量较低,成熟时期表达量最高,表明这2个基因可能在葡萄果实发育转色过程中发挥关键作用。 结论 葡萄中鉴定出86个VvWD40基因,不均匀地分布在19条染色体上。大部分VvWD40在葡萄果实发育过程中表达量变化显著,表明其在葡萄生长发育中发挥重要作用。
张高翔, 吴玉碧, 郭亚静, 纪薇, 杨忠义. 葡萄WD40基因家族鉴定及表达量分析[J]. 生物技术通报, 2026, 42(3): 324-337.
ZHANG Gao-xiang, WU Yu-bi, GUO Ya-jing, JI Wei, YANG Zhong-yi. Identification and Expression Analysis of WD40 Gene Family in Grape[J]. Biotechnology Bulletin, 2026, 42(3): 324-337.
| 基因 Gene | 正向引物 Forward primer (5′‒3′) | 反向引物 Reverse primer (5′‒3′) |
|---|---|---|
| WD40-3 | F: TTGGAAGGACACGAGAAACC | R: GGATACCTGCCACACCTTAATC |
| WD40-9 | F: GACTTCCTCAAGAGCTCCATAAA | R: GTCCAAGCTCAAGCAAGAAAC |
| WD40-27 | F: GTTCCTTCGATGAGACCGTTAG | R: GAGCCATCACGATTGAAATTGG |
| WD40-34 | F: CAGTTTAGAGAGCGGCAAGAA | R: GTCCGTACAGCCATGTCATTAG |
| WD40-42 | F: AGTGCCAGCAAGGATTCTAC | R: TCTAGGCTGCAACTCCAAAC |
| WD40-49 | F: GTGAGGACTCAAAGGTGTATGT | R: GGACTCCAGCTCACACAATTA |
| WD40-51 | F: CGTGCGATCTCTCGTGTATT | R: GCTCTTTCCCTCAGCATCATA |
| WD40-77 | F: CTGACTCGCCACAGGAATATC | R: CTTTCACCGACCAGACATAGAG |
| EF1α | F: GAACTGGGTGCTTGATAGGC | R: ACCAAAATATCCGGAGTAAAAGA |
表1 引物信息
Table 1 Primer information
| 基因 Gene | 正向引物 Forward primer (5′‒3′) | 反向引物 Reverse primer (5′‒3′) |
|---|---|---|
| WD40-3 | F: TTGGAAGGACACGAGAAACC | R: GGATACCTGCCACACCTTAATC |
| WD40-9 | F: GACTTCCTCAAGAGCTCCATAAA | R: GTCCAAGCTCAAGCAAGAAAC |
| WD40-27 | F: GTTCCTTCGATGAGACCGTTAG | R: GAGCCATCACGATTGAAATTGG |
| WD40-34 | F: CAGTTTAGAGAGCGGCAAGAA | R: GTCCGTACAGCCATGTCATTAG |
| WD40-42 | F: AGTGCCAGCAAGGATTCTAC | R: TCTAGGCTGCAACTCCAAAC |
| WD40-49 | F: GTGAGGACTCAAAGGTGTATGT | R: GGACTCCAGCTCACACAATTA |
| WD40-51 | F: CGTGCGATCTCTCGTGTATT | R: GCTCTTTCCCTCAGCATCATA |
| WD40-77 | F: CTGACTCGCCACAGGAATATC | R: CTTTCACCGACCAGACATAGAG |
| EF1α | F: GAACTGGGTGCTTGATAGGC | R: ACCAAAATATCCGGAGTAAAAGA |
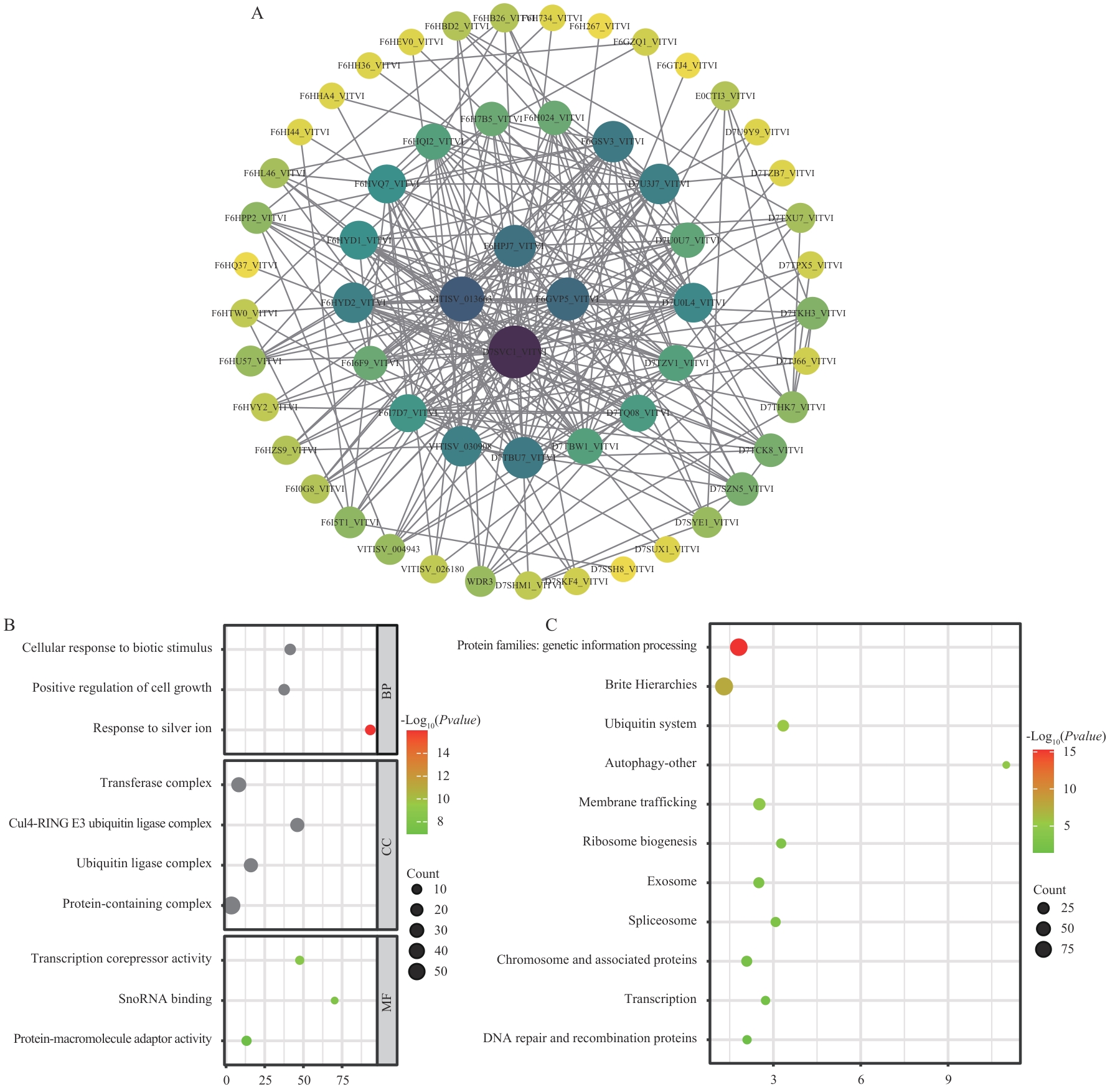
图10 VvWD40基因家族蛋白互作网络分析(A)、GO功能注释(B)和KEGG富集分析(C)
Fig. 10 Protein interaction network analysis (A), GO functional annotation (B), and KEGG enrichment analysis of the VvWD40 gene family
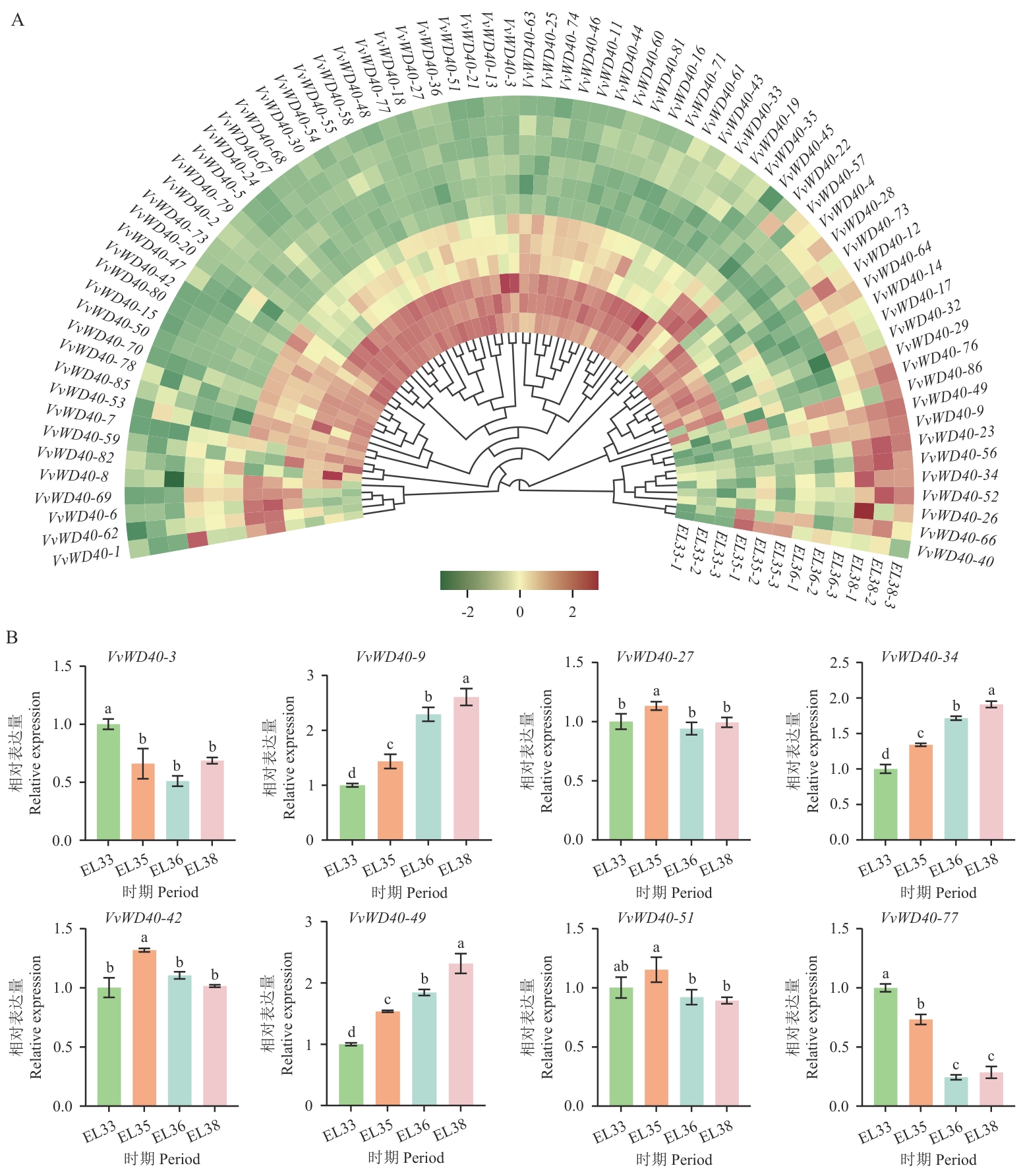
图11 VvWD40基因家族表达模式分析及RT-qPCR验证EL33:浆果仍然坚硬且呈绿色;EL35:浆果开始着色并增大;EL36:浆果具有中等可溶性固形物含量;EL38:浆果收获成熟。不同小写字母表示存在显著差异(P<0.05)
Fig. 11 Analysis of the expression pattern of the VvWD40 gene family and RT-qPCR verificationEL33: The berries are still firm and green. EL35: The berries start to color and increase in size. EL36: The berries have a medium content of soluble solids. EL38: The berries are ready for harvest and fully mature. Different lowercase letters indicate significant differences (P<0.05)
| [1] | Wang WN, Qian YH, Liu RH, et al. Effects of table grape cultivars on fruit quality and aroma components [J]. Foods, 2023, 12(18): 3371. |
| [2] | Ma MZ, Ma XY, Ma ZH, et al. Effects of foliar fertilizer additives on grape fruit quality and endogenous hormones in leaves [J]. BMC Plant Biol, 2025, 25(1): 516. |
| [3] | Jia N, Yin YG, Li MM, et al. Berry thinning affects the fruit quality composition of two table grape cultivars under linkage greenhouse conditions [J]. HortScience, 2023, 58(1): 134-140. |
| [4] | Feng RC, Zhang CH, Ma RJ, et al. Identification and characterization of WD40 superfamily genes in peach [J]. Gene, 2019, 710: 291-306. |
| [5] | Yan CY, Yang T, Wang BK, et al. Genome-wide identification of the WD40 gene family in tomato (Solanum lycopersicum L.) [J]. Genes, 2023, 14(6): 1273. |
| [6] | Ji XL, Zhang MY, Wang D, et al. Genome-wide identification of WD40 superfamily in Cerasus humilis and functional characteristics of ChTTG1 [J]. Int J Biol Macromol, 2023, 225: 376-388. |
| [7] | Wu ZR, Zhang TY, Li JN, et al. Genome-wide analysis of WD40 protein family and functional characterization of BvWD40-82 in sugar beet [J]. Front Plant Sci, 2023, 14: 1185440. |
| [8] | Fan ZY, Zhai YL, Wang Y, et al. Genome-wide analysis of anthocyanin biosynthesis regulatory WD40 gene FcTTG1 and related family in Ficus carica L [J]. Front Plant Sci, 2022, 13: 948084. |
| [9] | Wang C, Tang YF, Li Y, et al. Genome-wide identification and bioinformatics analysis of the WD40 transcription factor family and candidate gene screening for anthocyanin biosynthesis in Rhododendron simsii [J]. BMC Genomics, 2023, 24(1): 488. |
| [10] | Kim YJ, Jung KH. WD40-domain protein GORI is an integrative scaffold that is required for pollen tube growth in rice [J]. Plant Signal Behav, 2023, 18(1): 2082678. |
| [11] | Salih H, Gong WF, Mkulama M, et al. Genome-wide characterization, identification, and expression analysis of the WD40 protein family in cotton [J]. Genome, 2018, 61(7): 539-547. |
| [12] | Colanero S, Tagliani A, Perata P, et al. Alternative splicing in the anthocyanin fruit gene encoding an R2R3 MYB transcription factor affects anthocyanin biosynthesis in tomato fruits [J]. Plant Commun, 2020, 1(1): 100006. |
| [13] | Yang MG, Chen SK, Geng JH, et al. Comprehensive analysis of the Spartina alterniflora WD40 gene family reveals the regulatory role of SaTTG1 in plant development [J]. Front Plant Sci, 2024, 15: 1390461. |
| [14] | Šulc M, Kotíková Z, Paznocht L, et al. Changes in anthocyanidin levels during the maturation of color-fleshed potato (Solanum tuberosum L.) tubers [J]. Food Chem, 2017, 237: 981-988. |
| [15] | Su MY, Wang S, Liu WJ, et al. MdJa2 participates in the brassinosteroid signaling pathway to regulate the synthesis of anthocyanin and proanthocyanidin in red-fleshed apple [J]. Front Plant Sci, 2022, 13: 830349. |
| [16] | Chen C, Yang YT, Pan L, et al. Genome-wide identification of WD40 proteins in Cucurbita maxima reveals its potential functions in fruit development [J]. Genes, 2023, 14(1): 220. |
| [17] | Sun YB, Zhang XJ, Zhong MC, et al. Genome-wide identification of WD40 genes reveals a functional diversification of COP1-like genes in Rosaceae [J]. Plant Mol Biol, 2020, 104(1/2): 81-95. |
| [18] | 尹魁林, 程莎莎, 艾长丰, 等. 枣BZR基因家族的鉴定及其在果实发育中的表达分析 [J/OL]. 分子植物育种. . |
| Yin KL, Cheng SS, Ai CF, et al. Genome-wide identification of ZjBZR gene family and expression analysis in jujube fruit [J/OL]. Mol Plant Breed, 2023. . | |
| [19] | 吴松平. 花椒WD40基因家族分析及ZbTTG1功能研究 [D]. 杨凌: 西北农林科技大学, 2023. |
| Wu SP. Identification of WD40 gene family and functional analysis of ZbTTG1 in Zanthoxylum bungeanum [D]. Yangling: Northwest A & F University, 2023. | |
| [20] | 刘冰晶. 梅花WD40基因家族的鉴定及表达分析 [D]. 合肥: 安徽农业大学, 2018. |
| Liu BJ. Identification and analysis on expression of the WD40 gene family in Prunus mume [D]. Hefei: Anhui Agricultural University, 2018. | |
| [21] | 杨煜琛, 靳雅荣, 骆金婵, 等. 珍珠粟WD40基因家族鉴定及表达特征分析 [J]. 作物学报, 2024, 50(9): 2219-2236. |
| Yang YC, Jin YR, Luo JC, et al. Identification and expression analysis of the WD40 gene family in pearl millet [J]. Acta Agron Sin, 2024, 50(9): 2219-2236. | |
| [22] | 钟梦军. 生菜WD40基因家族分析和控制苋菜红色叶片的基因定位 [D]. 武汉: 华中农业大学, 2021. |
| Zhong MJ. Analysis of WD40 gene family in lettuce and fine gene mapping of controlling red leaf in amaranth [D]. Wuhan: Huazhong Agricultural University, 2021. | |
| [23] | 胡瑞. 小麦WD40基因家族鉴定及TaLWD1L、TaTTG1的功能研究 [D]. 武汉: 华中科技大学, 2020. |
| Hu R. Identification of wheat WD40 gene family and functional analysis of TaLWD1L and TaTTG1 [D]. Wuhan: Huazhong University of Science and Technology, 2020. | |
| [24] | 牛禹极, 吴锦林, 陈耀阳, 等. 木豆WD40基因家族鉴定及响应茉莉酸甲酯的表达分析 [J]. 石河子大学学报: 自然科学版, 2024, 42(1): 36-46. |
| Niu YJ, Wu JL, Chen YY, et al. Identification of WD40 gene family in pigeon pea and their expression pattern responded to Methyl Jasmonate [J]. J Shihezi Univ Nat Sci, 2024, 42(1): 36-46. | |
| [25] | 王刚. 红花WD40基因家族分析及CtWD40-1基因克隆与功能初步研究 [D]. 长春: 吉林农业大学, 2020. |
| Wang G. Bioinformatics of WD40 gene family in safflower and the cloning and preliminary characterization of CtWD40-1 [D]. Changchun: Jilin Agricultural University, 2020. | |
| [26] | 马文. 丹参WD40基因家族鉴定及SmWD40-170基因功能研究 [D]. 西安: 陕西师范大学, 2019. |
| Ma W. Identification of the WD40 gene family in Salvia miltiorrhiza and functional study of the SmWD40-170 gene [D]. Xi’an: Shaanxi Normal University, 2019. | |
| [27] | Grativol C, Thiebaut F, Sangi S, et al. A miniature inverted-repeat transposable element, AddIn-MITE located inside a WD40 gene is conserved in Andropogoneae grasses [J]. PeerJ, 2019, 7: e6080. |
| [28] | Mishra AK, Muthamilarasan M, Khan Y, et al. Genome-wide investigation and expression analyses of WD40 protein family in the model plant foxtail millet (Setaria italica L.) [J]. PLoS One, 2014, 9(1): e86852. |
| [29] | Tevatia R, Oyler GA. Evolution of DDB1-binding WD40 (DWD) in the viridiplantae [J]. PLoS One, 2018, 13(1): e0190282. |
| [30] | Xin Y, Wu YQ, Han X, et al. Overexpression of the Ginkgo biloba WD40 gene GbLWD1-like improves salt tolerance in transgenic Populus [J]. Plant Sci, 2021, 313: 111092. |
| [31] | Chen L, Cui YM, Yao YH, et al. Genome-wide identification of WD40 transcription factors and their regulation of the MYB-bHLH-WD40 (MBW) complex related to anthocyanin synthesis in Qingke (Hordeum vulgare L. var. nudum Hook. f.) [J]. BMC Genomics, 2023, 24(1): 166. |
| [32] | Luo RH, Pan WQ, Liu WQ, et al. The barley DIR gene family: an expanded gene family that is involved in stress responses [J]. Front Genet, 2022, 13: 1042772. |
| [33] | El-Sayed ASA, Mohamed NZ, Safan S, et al. Restoring the taxol biosynthetic machinery of Aspergillus terreus by Podocarpus gracilior pilger microbiome, with retrieving the ribosome biogenesis proteins of WD40 superfamily [J]. Sci Rep, 2019, 9(1): 11534. |
| [34] | Alvarez JM, Bueno N, Cañas RA, et al. Analysis of the WUSCHEL-RELATED HOMEOBOX gene family in Pinus pinaster: New insights into the gene family evolution [J]. Plant Physiol Biochem, 2018, 123: 304-318. |
| [35] | Xu MY, Fu JJ, Ni Y, et al. Genome-wide analysis of the MYB gene family in pumpkin [J]. PeerJ, 2024, 12: e17304. |
| [36] | Zhang Y, Gao WL, Li HT, et al. Genome-wide analysis of the bZIP gene family in Chinese jujube (Ziziphus jujuba Mill.) [J]. BMC Genomics, 2020, 21(1): 483. |
| [37] | Nguyen HM, Putterill J, Dare AP, et al. Two genes, ANS and UFGT2 from Vaccinium spp. are key steps for modulating anthocyanin production [J]. Front Plant Sci, 2023, 14: 1082246. |
| [38] | Zhao ZC, Hu GB, Hu FC, et al. The UDP glucose: flavonoid-3-O-glucosyltransferase (UFGT) gene regulates anthocyanin biosynthesis in litchi (Litchi chinesis Sonn.) during fruit coloration [J]. Mol Biol Rep, 2012, 39(6): 6409-6415. |
| [39] | Gao HN, Jiang H, Lian XY, et al. Identification and functional analysis of the MdLTPG gene family in apple [J]. Plant Physiol Biochem, 2021, 163: 338-347. |
| [40] | Gao YL, Zhang ZX, Cheng J, et al. Genome-wide identification of the CER1 gene family in apple and response of MdCER1-1 to drought stress [J]. Funct Integr Genomics, 2022, 23(1): 17. |
| [41] | Che LL, Lu SX, Liang GP, et al. Identification and expression analysis of the grape pentatricopeptide repeat (PPR) gene family in abiotic stress [J]. Physiol Mol Biol Plants, 2022, 28(10): 1849-1874. |
| [42] | Ge MQ, Zhong R, Sadeghnezhad E, et al. Genome-wide identification and expression analysis of magnesium transporter gene family in grape (Vitis vinifera) [J]. BMC Plant Biol, 2022, 22(1): 217. |
| [43] | Wu XX, Gong QH, Ni XP, et al. UFGT: the key enzyme associated with the petals variegation in Japanese apricot [J]. Front Plant Sci, 2017, 8: 108. |
| [44] | Hu CY, Gong YF, Jin S, et al. Molecular analysis of a UDP-glucose: flavonoid 3-O-glucosyltransferase (UFGT) gene from purple potato (Solanum tuberosum) [J]. Mol Biol Rep, 2011, 38(1): 561-567. |
| [1] | 苏燕竹, 李达, 张爱爱, 刘永光, 张秀荣, 薛其勤. 大豆CAD基因家族的鉴定及表达分析[J]. 生物技术通报, 2026, 42(4): 101-113. |
| [2] | 刘青媛, 吴洪启, 陈秀娥, 陈剑, 姜远泽, 何燕子, 喻奇伟, 刘仁祥. 转录因子NtMYB96a调控烟草耐旱性的功能研究[J]. 生物技术通报, 2026, 42(4): 239-250. |
| [3] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [4] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [5] | 胡秋玲, 陈灵, 黄嘉怡, 赵梓乔, 潘璐怡, 刘慧丽, 刘太波. 多胺调控果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 203-212. |
| [6] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [7] | 崔之瀚, 魏庆镇, 胡娜, 包崇来, 王华森. 园艺作物IQD基因研究进展[J]. 生物技术通报, 2026, 42(3): 230-241. |
| [8] | 彭楚, 孙娟利, 郑蓓蓓, 张若西, 韩月彭, 赵云. 果实花色苷酶促降解机制研究进展[J]. 生物技术通报, 2026, 42(3): 133-144. |
| [9] | 李成泉, 史庆华, 杨晓玉. 园艺作物果实发育的miRNA调控网络:从分子机制到种质创新[J]. 生物技术通报, 2026, 42(3): 19-36. |
| [10] | 刘林娅, 刘欢艳, 梁鑫钰, 宋姝熠, 何斌, 王绪英, 黄亚成. ‘红阳’猕猴桃BGAL基因家族的全基因组鉴定与表达分析[J]. 生物技术通报, 2026, 42(3): 312-323. |
| [11] | 李亚妮, 韩鸿宇, 耿梦爽, 米若兰, 王韦琪, 于文静, 孟宪文, 李传友. ChiC基因调控番茄灰霉病抗性的机制研究[J]. 生物技术通报, 2026, 42(3): 255-262. |
| [12] | 李天源, 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏. 欧李SPL基因家族的鉴定及在果实发育过程中的表达分析[J]. 生物技术通报, 2026, 42(3): 362-373. |
| [13] | 徐泽, 周陈平, 邝瑞彬, 吴夏明, 杨敏, 刘传和, 贺涵, 魏岳荣. 番木瓜PG基因家族鉴定及其在果实软化中的功能[J]. 生物技术通报, 2026, 42(3): 349-361. |
| [14] | 罗威, 宫奥, 仲阳, 胡迪, 周洪源, 张泓欣, 艾菊, 罗有卫, 高冬丽. 黄瓜SEPALLATA2基因敲除对果实及疣状结构发育的多效性影响[J]. 生物技术通报, 2026, 42(3): 283-293. |
| [15] | 尹跃, 秦小雅, 米佳, 安巍, 何军, 张锋锋. 枸杞FBN基因家族鉴定及与类胡萝卜素代谢的相关性分析[J]. 生物技术通报, 2026, 42(3): 338-348. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||