








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 263-274.doi: 10.13560/j.cnki.biotech.bull.1985.2026-0160
李迎辉1( ), 王杨博涵2, 周浩博3, 卢心如3, 张珂欣3, 于洋4, 李传友4, 孙传龙2(
), 王杨博涵2, 周浩博3, 卢心如3, 张珂欣3, 于洋4, 李传友4, 孙传龙2( )
)
收稿日期:2026-02-01
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
孙传龙,男,博士,教授,研究方向 :番茄驯化机理解析;E-mail: clsun@sdau.edu.cn作者简介:李迎辉,男,硕士,研究方向 :分子遗传育种;E-mail: liyinghui@hainanu.edu.cn
基金资助:
LI Ying-hui1( ), WANG Yang-bo-han2, ZHOU Hao-bo3, LU Xin-ru3, ZHANG Ke-xin3, YU Yang4, LI Chuan-you4, SUN Chuan-long2(
), WANG Yang-bo-han2, ZHOU Hao-bo3, LU Xin-ru3, ZHANG Ke-xin3, YU Yang4, LI Chuan-you4, SUN Chuan-long2( )
)
Received:2026-02-01
Published:2026-03-26
Online:2026-04-23
摘要:
目的 系统鉴定番茄液泡加工酶(vacuolar processing enzyme, VPE)基因家族成员,解析其进化关系、表达模式及在非生物胁迫响应中的潜在功能。 方法 利用生物信息学方法鉴定SlVPE家族成员,并对其理化性质、系统进化、基因结构、保守域、染色体分布、顺式作用元件及种间共线性进行系统分析;整合公共转录组数据与RT-qPCR技术,分析该家族的组织特异性表达模式及其在干旱、高低温、盐、碱及盐碱混合胁迫下的表达响应。 结果 在番茄中鉴定到13个含有Peptidase_C13与legumain_C结构域的SlVPE基因,分为3个亚家族。除SlVPE4和SlVPE5外,所有成员以基因簇的形式聚集于第8染色体。表达分析表明,SlVPE家族具有明显组织及时空特异性,SlVPE3、SlVPE5为组成型高表达基因,SlVPE2、SlVPE7为果实特异性高表达基因;非生物胁迫下,家族整体响应以表达抑制为主,SlVPE3为干旱、盐及盐碱混合胁迫的核心响应基因,SlVPE7受碱胁迫特异性诱导,SlVPE8主要响应高温和低温胁迫。根据野生种醋栗番茄和栽培番茄的序列变异,在SlVPE基因启动子区或基因下游开发了4个多态性分子标记。 结论 番茄SlVPE基因家族包含13个成员,其中11个成员以基因簇形式聚集于第8号染色体。SlVPE3和SlVPE5为组成型高表达基因,SlVPE2和SlVPE7在果实发育早期特异性高表达,而非生物胁迫主要诱导SlVPE3、SlVPE7和SlVPE8基因的表达。
李迎辉, 王杨博涵, 周浩博, 卢心如, 张珂欣, 于洋, 李传友, 孙传龙. 番茄VPE基因家族鉴定和抗逆功能分析[J]. 生物技术通报, 2026, 42(3): 263-274.
LI Ying-hui, WANG Yang-bo-han, ZHOU Hao-bo, LU Xin-ru, ZHANG Ke-xin, YU Yang, LI Chuan-you, SUN Chuan-long. Identification of VPE Gene Family and Their Functional Analysis under Abiotic Stress in Tomato[J]. Biotechnology Bulletin, 2026, 42(3): 263-274.
基因名称 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids | 分子量 Molecular weight (kD) | 疏水性 GRAVY | 等电点 pI |
|---|---|---|---|---|---|
| SlVPE1 | Solyc08g065790.2.1 | 460 | 51.77 | -0.49 | 6.92 |
| SlVPE2 | Solyc08g065780.1.1 | 451 | 50.09 | -0.12 | 5.76 |
| SlVPE3 | Solyc08g065610.2.1 | 470 | 52.11 | -0.36 | 7.27 |
| SlVPE4 | Solyc08g079160.2.1 | 469 | 52.00 | -0.33 | 7.59 |
| SlVPE5 | Solyc12g095910.1.1 | 463 | 51.04 | -0.20 | 5.05 |
| SlVPE6 | Solyc08g065530.1.1 | 483 | 53.88 | -0.31 | 5.83 |
| SlVPE7 | Solyc08g065570.1.1 | 428 | 47.54 | -0.28 | 7.49 |
| SlVPE8 | Solyc08g065590.1.1 | 446 | 49.35 | -0.30 | 7.26 |
| SlVPE9 | Solyc08g065690.1.1 | 450 | 49.50 | -0.20 | 7.27 |
| SlVPE10 | Solyc08g065710.1.1 | 451 | 50.68 | -0.44 | 7.78 |
| SlVPE11 | Solyc08g065720.1.1 | 482 | 53.22 | -0.34 | 5.67 |
| SlVPE12 | Solyc08g065740.1.1 | 497 | 55.21 | -0.37 | 5.49 |
| SlVPE13 | Solyc08g065750.1.1 | 480 | 52.97 | -0.27 | 6.10 |
表1 SlVPEs基本信息
Table 1 Basic information about SlVPEs
基因名称 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids | 分子量 Molecular weight (kD) | 疏水性 GRAVY | 等电点 pI |
|---|---|---|---|---|---|
| SlVPE1 | Solyc08g065790.2.1 | 460 | 51.77 | -0.49 | 6.92 |
| SlVPE2 | Solyc08g065780.1.1 | 451 | 50.09 | -0.12 | 5.76 |
| SlVPE3 | Solyc08g065610.2.1 | 470 | 52.11 | -0.36 | 7.27 |
| SlVPE4 | Solyc08g079160.2.1 | 469 | 52.00 | -0.33 | 7.59 |
| SlVPE5 | Solyc12g095910.1.1 | 463 | 51.04 | -0.20 | 5.05 |
| SlVPE6 | Solyc08g065530.1.1 | 483 | 53.88 | -0.31 | 5.83 |
| SlVPE7 | Solyc08g065570.1.1 | 428 | 47.54 | -0.28 | 7.49 |
| SlVPE8 | Solyc08g065590.1.1 | 446 | 49.35 | -0.30 | 7.26 |
| SlVPE9 | Solyc08g065690.1.1 | 450 | 49.50 | -0.20 | 7.27 |
| SlVPE10 | Solyc08g065710.1.1 | 451 | 50.68 | -0.44 | 7.78 |
| SlVPE11 | Solyc08g065720.1.1 | 482 | 53.22 | -0.34 | 5.67 |
| SlVPE12 | Solyc08g065740.1.1 | 497 | 55.21 | -0.37 | 5.49 |
| SlVPE13 | Solyc08g065750.1.1 | 480 | 52.97 | -0.27 | 6.10 |
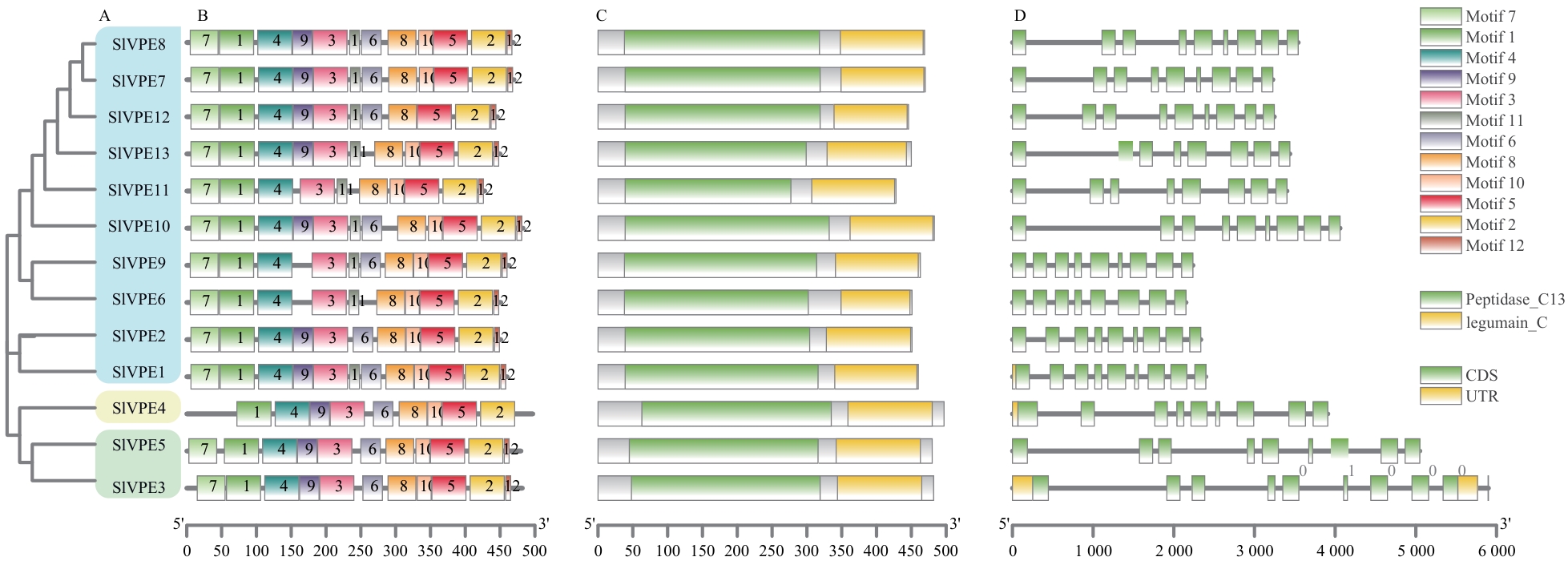
图2 SlVPEs蛋白基序、保守结构域和基因结构分析A:SlVPE蛋白系统发育树;B:SlVPE蛋白保守基序分析;C:SlVPE蛋白保守结构域分析,绿色和黄色分别代表Peptidase_C13结构域和legumain_C结构域;D:SlVPE基因结构分析;绿框代表CDS,黄框代表UTR,灰色线条代表内含子
Fig. 2 Motifs, conserved domains and gene structure analysis of SlVPE proteinA: Phylogenetic tree of SlVPE proteins. B: Analysis of conserved motifs in SlVPE proteins. C: Analysis of conserved domains of SlVPE proteins, with green and yellow boxes representing Peptidase_C13 and legumain_C domains, respectively. D: Analysis of SlVPE gene structures. CDS, UTRs, and introns are indicated by green boxes, yellow boxes, and gray lines, respectively
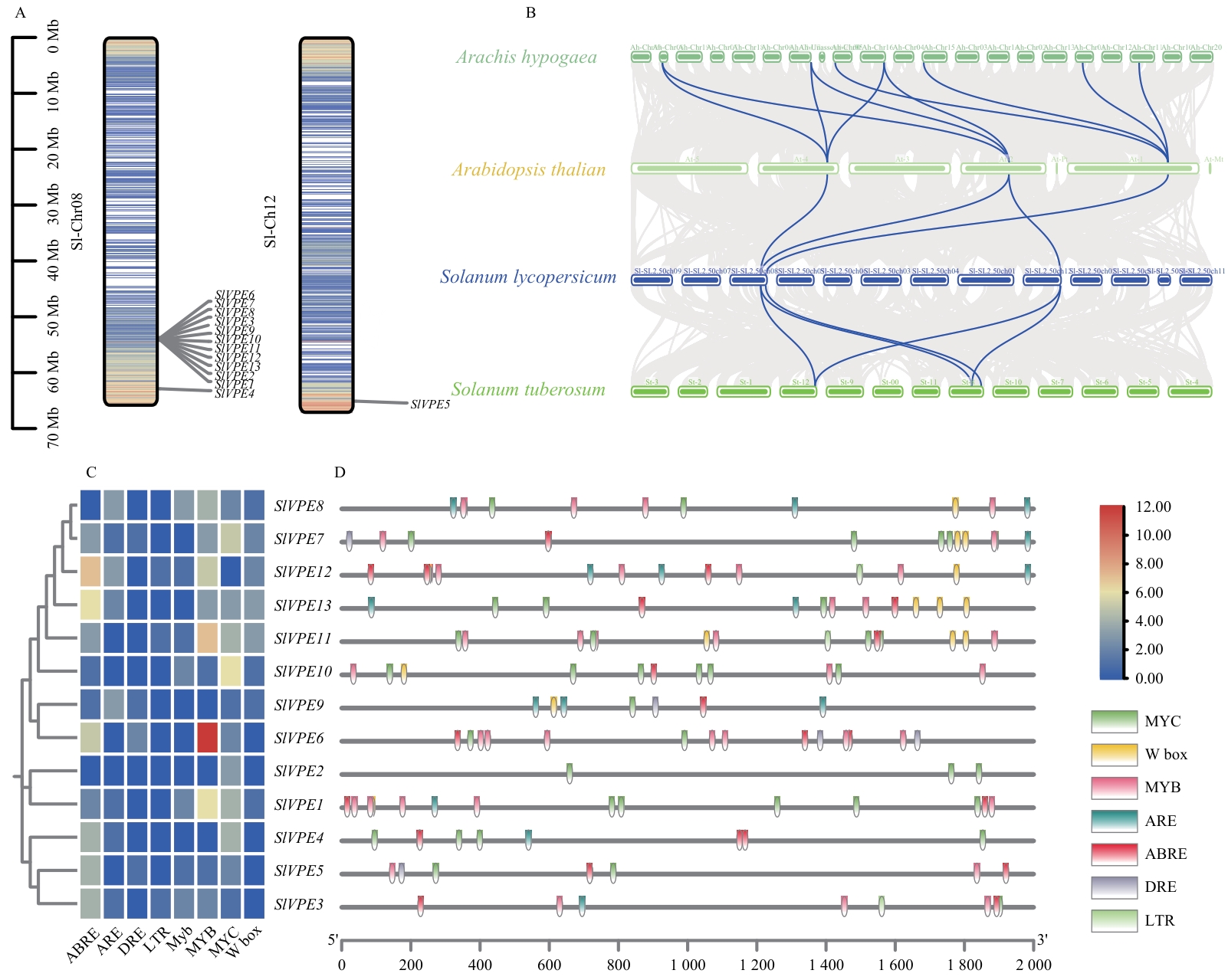
图3 SlVPE基因染色体定位、种间共线性和顺式作用元件分析A:SlVPE基因在染色体中的位置;B:SlVPE基因在番茄、拟南芥、马铃薯和花生基因组之间的共线性关系;灰色和蓝色分别代表基因组所有基因和VPE基因的共线性;C:SlVPEs启动子的顺式作用元件热图;D:SlVPEs启动子顺式作用元件的种类、位置和数量的可视化
Fig. 3 Analysis of chromosomal localization, interspecies collinearity, and cis-acting elements of SlVPE genesA: Chromosomal locations of the SlVPE genes. B: Collinear relationships of SlVPE genes among tomato, Arabidopsis, potato, and peanut genomes. Gray and blue lines indicate the collinearity of all genes and VPE genes, respectively. C: Heatmap of cis-acting elements in SlVPEs promoters. D: Visualization of the types, positions, and numbers of cis-acting elements in SlVPEs promoters
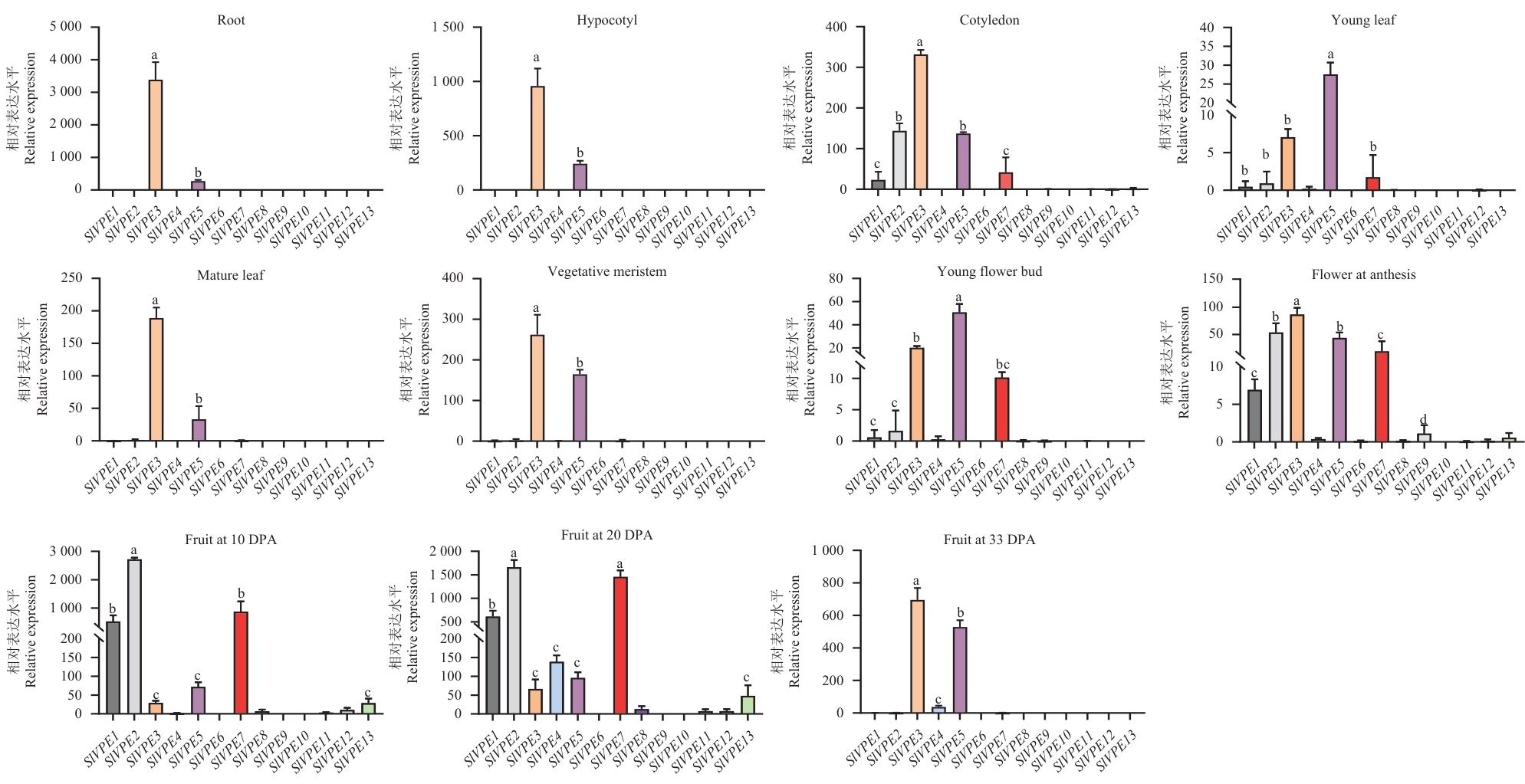
图4 SlVPE基因在番茄不同组织中的表达谱分析数据以4个生物学重复(n=4)的均值±标准误(SE)表示。不同小写字母表示差异显著(P<0.05)
Fig. 4 Expression profile analysis of SlVPE gene in different tissues of tomatoData are presented as mean ± SE from four biological replicates (n=4). Different lowercase letters indicate significant differences (P<0.05)
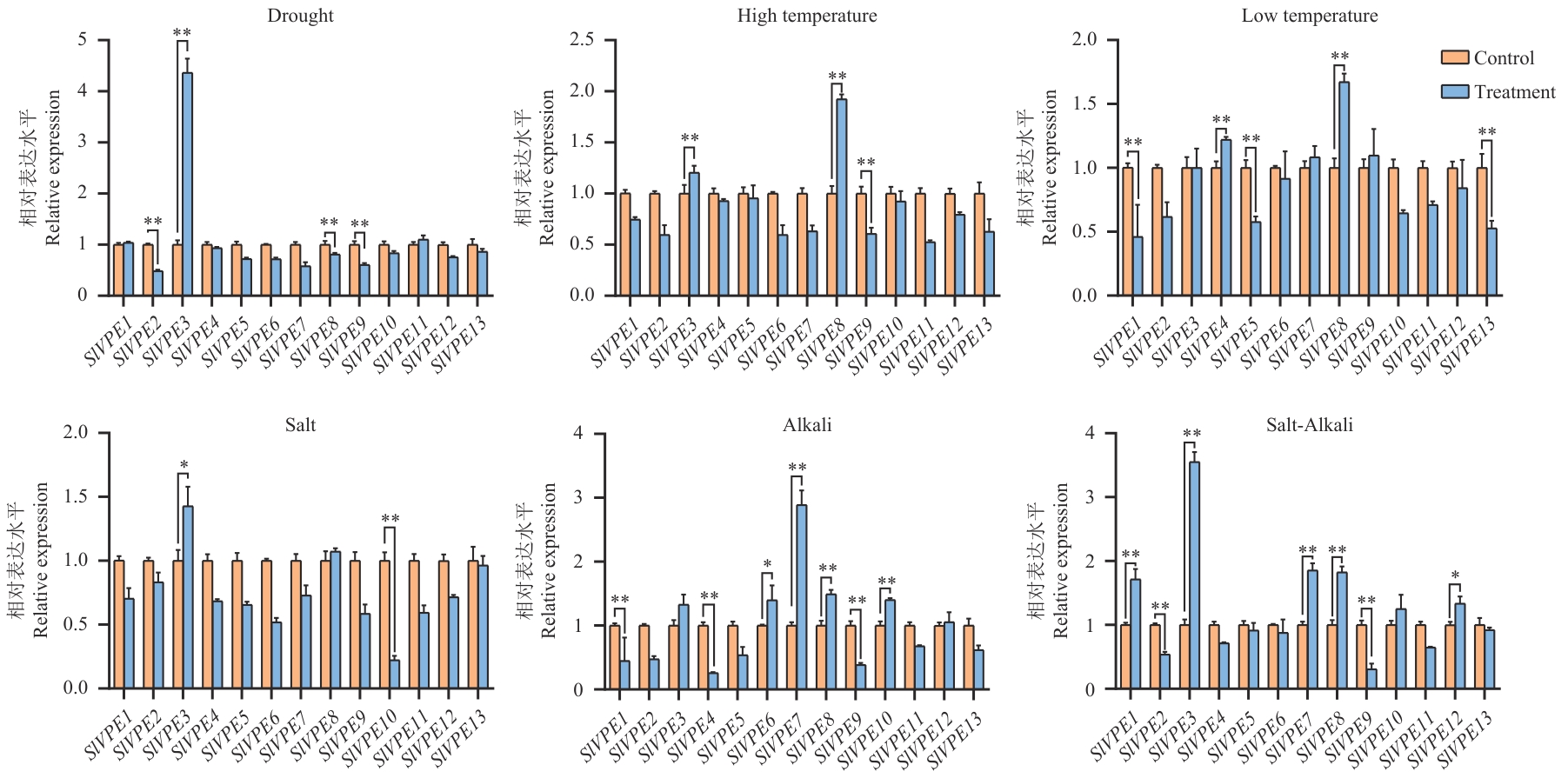
图6 逆境胁迫下SlVPE基因表达分析干旱胁迫:20% PEG6000;盐胁迫:150 mmol/L NaCl;碱胁迫:100 mmol/L NaHCO3;盐碱复合胁迫:60 mmol/L NaCl+60 mmol/L NaHCO3;低温胁迫:12 ℃;高温胁迫:38 ℃。取样时间点为胁迫处理后7 d。数据以3个生物学重复(n=3)的均值±标准误(SE)表示;每个生物学重复包含3个技术重复。*P<0.05,**P<0.01
Fig. 6 Expression analysis of SlVPE genes under various abiotic stressesDrought stress: 20% PEG6000; salt stress: 150 mmol/L NaCl; alkali stress: 100 mmol/L NaHCO₃; salt-alkali combined stress: 60 mmol/L NaCl+60 mmol/L NaHCO3; low-temperature stress: 12 ℃; high-temperature stress: 38 ℃. Samples were collected at 7 d after stress treatment. Data are presented as mean±SE of three biological replicates (n=3), each with three technical replicates. *P<0.05, **P<0.01
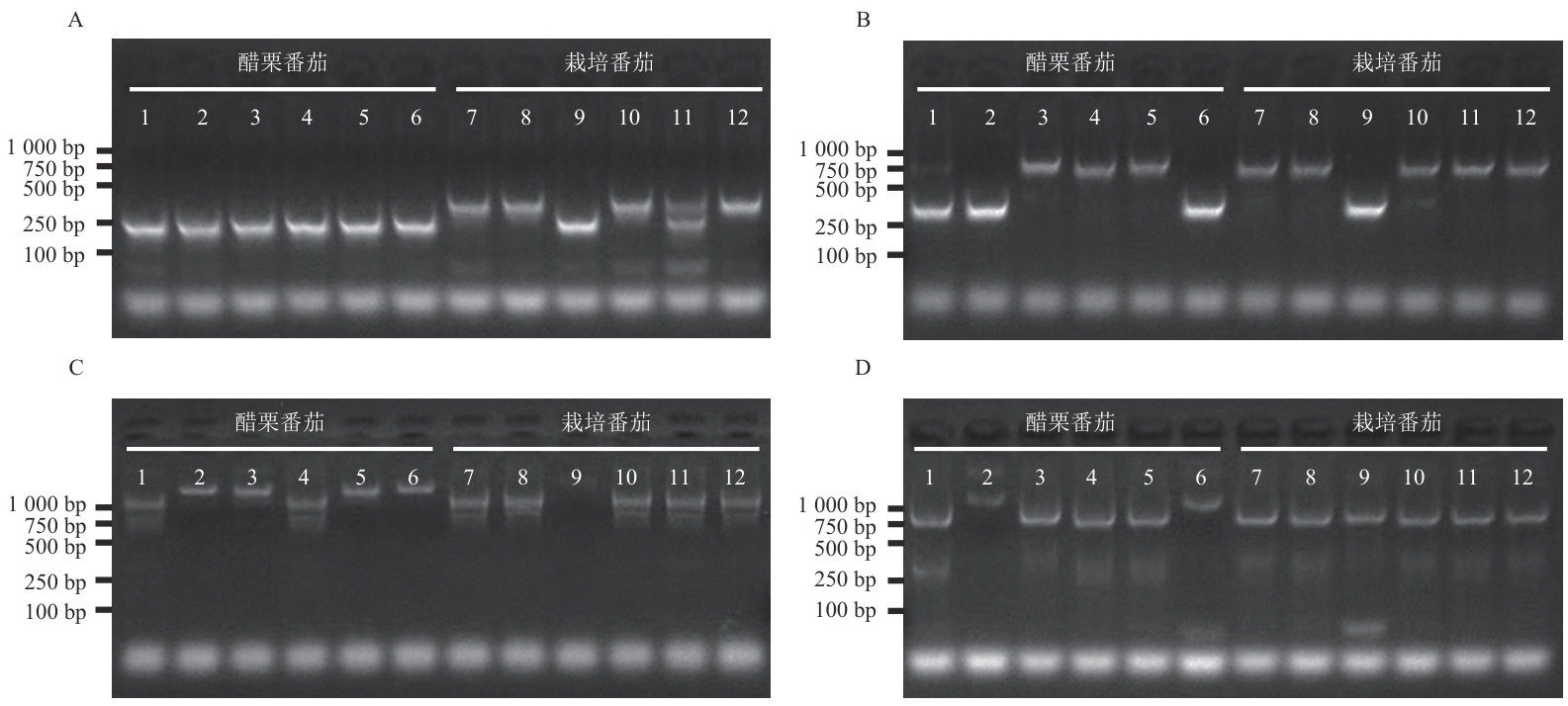
图7 SlVPEs基因分子标记的多态性检测利用琼脂糖凝胶电泳检测SlVPE6启动子区(A)、SlVPE11基因下游(B)、SlVPE13基因启动子区(C)、SlVPE10基因启动子区(D)在醋栗番茄和栽培番茄间的多态性。泳道1-6:醋栗番茄种质,依次为LA0533、LA1375、LA1420、LA2873、LA4763、LA1589;泳道7-12:栽培番茄材料,依次为AC、M82、TS-418、Hawaii7996、MoneyMaker、Heinz1706
Fig. 7 Polymorphism detection of SlVPE gene molecular markersAgarose gel electrophoresis was used to examine polymorphisms between Solanum pimpinellifolium and cultivated tomato in the promoter region of SlVPE6 (A), the downstream region of SlVPE11 (B), the promoter region of SlVPE13 (C), and the promoter region of SlVPE10 (D). Lane 1-6: S. pimpinellifolium accessions, LA0533, LA1375, LA1420, LA2873, LA4763, and LA1589; lane 7-12: cultivated tomato materials, AC, M82, TS-418, Hawaii7996, MoneyMaker, and Heinz1706
| [1] | Hara-Nishimura I, Hatsugai N. The role of vacuole in plant cell death [J]. Cell Death Differ, 2011, 18(8): 1298-1304. |
| [2] | Teper-Bamnolker P, Danieli R, Peled-Zehavi H, et al. Vacuolar processing enzyme translocates to the vacuole through the autophagy pathway to induce programmed cell death [J]. Autophagy, 2021, 17(10): 3109-3123. |
| [3] | Sarwar R, Yu J, Zekraoui M, et al. Unraveling the molecular functions of multifaced plant-vacuolar processing enzymes [J]. Physiol Plant, 2024, 176: e14131. |
| [4] | Gao XX, Tang YL, Shi QY, et al. Vacuolar processing enzyme positively modulates plant resistance and cell death in response to Phytophthora parasitica infection [J]. J Integr Agric, 2023, 22(5): 1424-1433. |
| [5] | Duan XB, Zhang K, Yu Y, et al. Genome-wide analysis of soybean vacuolar processing enzyme gene family reveals their roles in plant development and response to stress conditions [J]. J Plant Growth Regul, 2024, 43(8): 2817-2829. |
| [6] | Shimada T, Yamada K, Kataoka M, et al. Vacuolar processing enzymes are essential for proper processing of seed storage proteins in Arabidopsis thaliana [J]. J Biol Chem, 2003, 278(34): 32292-32299. |
| [7] | Tang CN, Wan Abdullah WMAN, Wee CY, et al. Promoter cis-element analyses reveal the function of αVPE in drought stress response of Arabidopsis [J]. Biology, 2023, 12(3): 430. |
| [8] | Nakaune S, Yamada K, Kondo M, et al. A vacuolar processing enzyme, δVPE, is involved in seed coat formation at the early stage of seed development [J]. Plant Cell, 2005, 17(3): 876-887. |
| [9] | Kuroyanagi M, Yamada K, Hatsugai N, et al. Vacuolar processing enzyme is essential for mycotoxin-induced cell death in Arabidopsis thaliana [J]. J Biol Chem, 2005, 280(38): 32914-32920. |
| [10] | Lu WY, Deng MJ, Guo F, et al. Suppression of OsVPE3 enhances salt tolerance by attenuating vacuole rupture during programmed cell death and affects stomata development in rice [J]. Rice, 2016, 9: 65. |
| [11] | Deng HB, Cao S, Zhang GL, et al. OsVPE2, a member of vacuolar processing enzyme family, decreases chilling tolerance of rice [J]. Rice, 2024, 17: 5. |
| [12] | Zhang JH, Yue Y, Hu MJ, et al. Dynamic transcriptome landscape of maize pericarp development [J]. Plant J, 2024, 117(5): 1574-1591. |
| [13] | Wang TH, Guan MH, Zheng YH, et al. Genome-wide molecular characterization and expression profiling of the cysteine protease gene family in maize [J]. BMC Genomics, 2025, 26(1): 789. |
| [14] | Quadros IPS, Madeira NN, Loriato VAP, et al. Cadmium-mediated toxicity in plant cells is associated with the DCD/NRP-mediated cell death response [J]. Plant Cell Environ, 2022, 45(2): 556-571. |
| [15] | Radchuk V, Tran V, Radchuk R, et al. Vacuolar processing enzyme 4 contributes to maternal control of grain size in barley by executing programmed cell death in the pericarp [J]. New Phytol, 2018, 218(3): 1127-1142. |
| [16] | Wang BK, Li N, Huang SY, et al. Enhanced soluble sugar content in tomato fruit using CRISPR/Cas9-mediated SlINVINH1 and SlVPE5 gene editing [J]. PeerJ, 2021, 9: e12478. |
| [17] | Wang WH, Cai JH, Wang PW, et al. Post-transcriptional regulation of fruit ripening and disease resistance in tomato by the vacuolar protease SlVPE3 [J]. Genome Biol, 2017, 18: 47. |
| [18] | Zhu LF, Wang XP, Tian J, et al. Genome-wide analysis of VPE family in four Gossypium species and transcriptional expression of VPEs in the upland cotton seedlings under abiotic stresses [J]. Funct Integr Genomics, 2022, 22(2): 179-192. |
| [19] | Song JF, Yang F, Xun M, et al. Genome-wide identification and characterization of vacuolar processing enzyme gene family and diverse expression under stress in apple (Malus × domestic) [J]. Front Plant Sci, 2020, 11: 626. |
| [20] | Bombarely A, Menda N, Tecle IY, et al. The Sol Genomics Network (solgenomics.net): growing tomatoes using Perl [J]. Nucleic Acids Res, 2011, 39(): D1149-D1155. |
| [21] | Consortium The Tomato Genome. The tomato genome sequence provides insights into fleshy fruit evolution [J]. Nature, 2012, 485(7400): 635-641. |
| [22] | Finn RD, Bateman A, Clements J, et al. Pfam: the protein families database [J]. Nucleic Acids Res, 2014, 42(D1): D222-D230. |
| [23] | Berardini TZ, Reiser L, Li DH, et al. The Arabidopsis information resource: Making and mining the “gold standard” annotated reference plant genome [J]. Genesis, 2015, 53(8): 474-485. |
| [24] | Bertioli DJ, Cannon SB, Froenicke L, et al. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut [J]. Nat Genet, 2016, 48(4): 438-446. |
| [25] | Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms [J]. Mol Biol Evol, 2018, 35(6): 1547-1549. |
| [26] | Chen CJ, Chen H, Zhang Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data [J]. Mol Plant, 2020, 13(8): 1194-1202. |
| [27] | Bailey TL, Boden M, Buske FA, et al. MEME Suite: tools for motif discovery and searching [J]. Nucleic Acids Res, 2009, 37(): W202-W208. |
| [28] | Lescot M, Déhais P, Thijs G, et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences [J]. Nucleic Acids Res, 2002, 30(1): 325-327. |
| [29] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method [J]. Methods, 2001, 25(4): 402-408. |
| [30] | Wleklik K, Borek S. Vacuolar processing enzymes in plant programmed cell death and autophagy [J]. Int J Mol Sci, 2023, 24(2): 1198. |
| [31] | Pat Heslop-Harrison JS, Schwarzacher T, Liu Q. Polyploidy: its consequences and enabling role in plant diversification and evolution [J]. Ann Bot, 2023, 131(1): 1-10. |
| [32] | Fang YP, Jiang JM, Hou XL, et al. Plant protein-coding gene families: Their origin and evolution [J]. Front Plant Sci, 2022, 13: 995746. |
| [33] | Zheng P, Zheng CY, Otegui MS, et al. Endomembrane mediated-trafficking of seed storage proteins: from Arabidopsis to cereal crops [J]. J Exp Bot, 2022, 73(5): 1312-1326. |
| [34] | Baral R, Vainer A, Melzer S, et al. ‘Bud to fruit’—hormonal interactions governing early fruit development [J]. J Exp Bot, 2025, 76(22): 6657-6673. |
| [35] | Wang RF, de Maagd RA. Transcriptional control of tomato fruit development and ripening [J]. J Exp Bot, 2025, 76(21): 6311-6326. |
| [36] | Wei SB, Li X, Lu ZF, et al. A transcriptional regulator that boosts grain yields and shortens the growth duration of rice [J]. Science, 2022, 377(6604): eabi8455. |
| [37] | Tan WR, Chen JH, Yue XL, et al. The heat response regulators HSFA1s promote Arabidopsis thermomorphogenesis via stabilizing PIF4 during the day [J]. Sci Adv, 2023, 9(44): eadh1738. |
| [38] | Xu L, Yang LJ, Li AP, et al. An AP2/ERF transcription factor confers chilling tolerance in rice [J]. Sci Adv, 2024, 10(35): eado4788. |
| [39] | Zhu JK. Abiotic stress signaling and responses in plants [J]. Cell, 2016, 167(2): 313-324. |
| [1] | 殷亚龙, 张明洋, 王洁敏, 苗雪雪, 陈劲, 王伟平. 水稻非生物胁迫协同耐受机制研究进展[J]. 生物技术通报, 2026, 42(4): 26-37. |
| [2] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [3] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [4] | 杜丹, 郭翔, 胡鑫, 潘宇. 质体发育调控果实成熟与品质的研究进展[J]. 生物技术通报, 2026, 42(3): 48-59. |
| [5] | 姜喆卉, 王小龙, 王守创, 周科. 番茄风味物质代谢途径解析与分子育种研究进展[J]. 生物技术通报, 2026, 42(3): 60-78. |
| [6] | 王潇奕, 李金焱, 邢醒, 朱鸿亮. 基于乙烯响应筛选调控番茄成熟且影响呼吸的基因及其功能分析[J]. 生物技术通报, 2026, 42(3): 275-282. |
| [7] | 刘淼, 林涛, 贾乐松, 胡丰, 李涛, 李志万, 刘美芳, 郑方燕, 崔龙. 从野生到栽培:番茄果实色泽的演化与调控机制[J]. 生物技术通报, 2026, 42(3): 187-202. |
| [8] | 李亚妮, 韩鸿宇, 耿梦爽, 米若兰, 王韦琪, 于文静, 孟宪文, 李传友. ChiC基因调控番茄灰霉病抗性的机制研究[J]. 生物技术通报, 2026, 42(3): 255-262. |
| [9] | 程云霞, 张俊红, 叶杰. 番茄果实可溶性固形物积累的遗传调控研究进展[J]. 生物技术通报, 2026, 42(3): 145-155. |
| [10] | 颜晨琳, 李凡, 闫春婷, 程蛟文, 胡开林, 叶志彪, 宋建文. 番茄果实形态发育相关基因研究进展[J]. 生物技术通报, 2026, 42(3): 172-186. |
| [11] | 李天源, 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏. 欧李SPL基因家族的鉴定及在果实发育过程中的表达分析[J]. 生物技术通报, 2026, 42(3): 362-373. |
| [12] | 龙林茜, 曾银萍, 王茜, 邓玉萍, 葛敏茜, 陈彦灼, 李鑫娟, 杨军, 邹建. 向日葵GH3基因家族鉴定及其在花发育中的功能分析[J]. 生物技术通报, 2026, 42(1): 125-138. |
| [13] | 李正, 邱炜玥, 孙瑞雪, 赵筱. 不同启动子过表达xylR基因提高大肠杆菌的葡萄糖-木糖共利用能力[J]. 生物技术通报, 2026, 42(1): 329-337. |
| [14] | 张驰昊, 刘晋囡, 晁跃辉. 蒺藜苜蓿bZIP转录因子MtbZIP29的克隆及功能分析[J]. 生物技术通报, 2026, 42(1): 241-250. |
| [15] | 吕镇, 甘恬, 霍思羽, 赵晨笛, 赵梦瑶, 李亚涛, 马玉超, 耿玉清. 产Surfactin贝莱斯芽胞杆菌C5A-1的鉴定和所产Surfactin对植物的促生效果[J]. 生物技术通报, 2025, 41(9): 265-276. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||