








Biotechnology Bulletin ›› 2023, Vol. 39 ›› Issue (12): 16-32.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0793
Previous Articles Next Articles
ZHU Ye-sheng( ), WU Guo-qiang(
), WU Guo-qiang( ), WEI Ming
), WEI Ming
Received:2023-08-14
Online:2023-12-26
Published:2024-01-11
Contact:
WU Guo-qiang
E-mail:1428209434@qq.com;gqwu@lut.edu.cn
ZHU Ye-sheng, WU Guo-qiang, WEI Ming. Roles of Plasma Membrane Na+/H+ Antiporter SOS1 in Maintaining Ionic Homeostasis of Plants[J]. Biotechnology Bulletin, 2023, 39(12): 16-32.
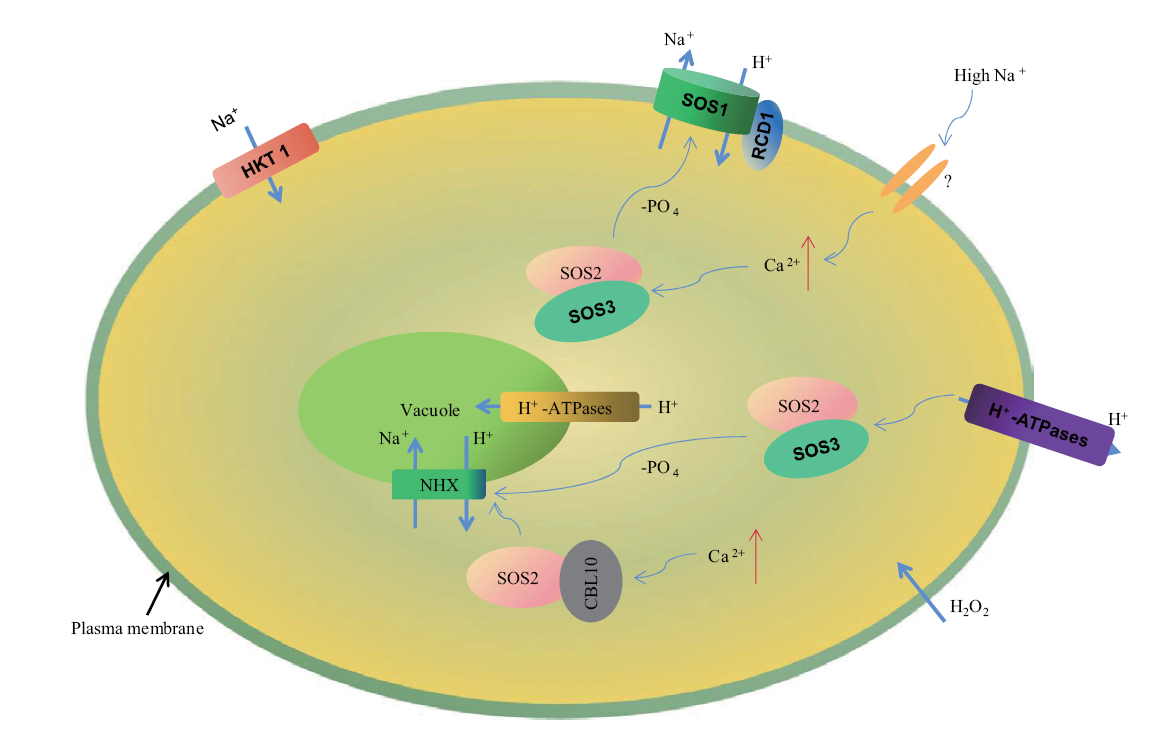
Fig. 1 Diagrammatic presentation of SOS1 response to salt stress Question marks indicate uncertainties or unknown components. Under high salt stress, the increase of Na+ level in cytoplasm can lead to the damage of multiple cellular processes. One of the main salt detoxification mechanisms in cells is calcium-activated SOS3-SOS2 protein complex, which activates SOS1 and is responsible for the efflux of Na+ out of cells. At the same time, the SOS3-SOS2 complex was also involved in the inhibition of HKT1(high-affinity K+ transporter 1), which transported Na+ under high salt conditions. Another member of the SOS3 family, CBL10, also formed a complex with SOS2. It is hypothesized that this complex can regulate Na+ efflux(by regulating SOS1)and the compartmentation of Na+ into the vacuole(activating NHX and pumping Na+ into the vacuole). SOS1 also interacts with RCD1 and mitigates the toxic effects of ROS and oxidative stress
| 物种名 Species name | 基因名称 Gene name | 基因登录号 GenBank accession No. | 氨基酸 Amino acid | 亚细胞定位 Sub-cellular localization | 跨膜结构域TMD | 亲水性平均值 GRAVY | 等电点 pI | 分子量 Mw/kD | 参考文献 Reference |
|---|---|---|---|---|---|---|---|---|---|
| 甜菜 Beta vulgaris | BvSOS1 | / | 1 162 | PM | 12 | 0.085 | 6.34 | 128.46 | Unpublished |
| 小盐芥thellungiella halophila | ThSOS1 | EF207775 | 1 146 | PM | 11 | 0.090 | 6.40 | 126.35 | [ |
| ThSOS1S | AB562331 | 473 | PM | 11 | 0.722 | 6.15 | 51.19 | [ | |
| 冰叶日中花 Mesembryanthemum crystallinum | McSOS1 | EF207776 | 1 115 | PM | 12 | 0.110 | 5.97 | 127.36 | [ |
| 花花柴 Karelinia caspia | KcSOS1 | MN481368 | 1 136 | PM | 12 | 0.070 | 5.97 | 126.02 | [ |
| 水稻Oryza sativa | OsSOS1-1 | AY785147 | 1 148 | PM | 12 | 0.046 | 6.77 | 127.92 | [ |
| OsSOS1-2 | MG602252 | 1 148 | PM | 13 | 0.047 | 6.77 | 127.90 | ||
| 小麦 Triticum aestivum | TaSOS1-1 | KJ563230 | 1 142 | PM | 12 | 0.110 | 7.00 | 126.22 | [ |
| TaSOS1-2 | AY326952 | 1 142 | PM | 13 | 0.106 | 7.40 | 126.18 | ||
| 大豆 Glycine max | GmSOS1 | JQ287499 | 1 143 | PM | 12 | 0.089 | 6.30 | 126.43 | [ |
| 文冠果Xanthoceras sorbifolium | XsSOS1 | / | 915 | PM | 6 | / | 6.19 | 102.50 | [ |
| 萝卜 Raphanus sativus | RsSOS1 | MZ484950 | 1 137 | PM | 12 | 0.093 | 6.71 | 125.62 | [ |
| 海马齿Sesuvium portulacastrum | SpSOS1 | JX674067 | 1 155 | PM | 12 | 0.063 | 6.08 | 127.95 | [ |
| 印度南瓜 Cucurbita pepo | CmaSOS1 | NW_019272028 | 1 142 | PM | 12 | 0.079 | 5.92 | 126.70 | [ |
| 陆地棉Gossypium hirsutum | GhSOS1 | KU994886 | 1 152 | PM | 12 | 0.061 | 6.32 | 128.18 | [ |
| 拟南芥Arabidopsis thaliana | AtSOS1 | AT2G01980 | 1 146 | PM | 13 | 0.098 | 7.62 | 127.19 | [ |
| 荞麦Fagopyrum esculentum | FeSOS1 | MH064255 | 1 132 | PM | 12 | 0.087 | 6.36 | 126.20 | [ |
| 白刺 Nitraria tangutorum | NtSOS1 | KC292267 | 1 163 | PM | 11 | 0.107 | 6.43 | 127.59 | [ |
| 黄花草木樨Melilotus officinalis | MoSOS1 | MG680455 | 957 | PM | 8 | 0.000 | 6.10 | 107.21 | [ |
| 黄花红砂Reaumuria trigyna | RtSOS1 | KC292265 | 1 145 | PM | 11 | 0.114 | 6.10 | 126.79 | [ |
| 木榄Bruguiera gymnorhiza | BgSOS1 | HM054521 | 1 153 | PM | 12 | 0.136 | 6.19 | 126.91 | [ |
| 盐生草Halogeton glomeratus | HgSOS1 | KT759142 | 1 165 | PM | 11 | 0.086 | 6.21 | 128.97 | / |
| 牡蒿 Artemisia japonica | AjSOS1 | KP896475 | 1 147 | PM | 12 | 0.061 | 6.08 | 127.04 | [ |
| 菠菜 Spinacia oleracea | SoSOS1 | HG799055 | 1 102 | / | 12 | 0.164 | 6.38 | 121.97 | [ |
| 葡萄 Vitis vinifera | VvSOS1 | NM_001281211 | 1 141 | / | 12 | 0.122 | 6.32 | 126.30 | [ |
| 黄瓜 Cucumis sativus | CsSOS1 | JQ655747 | 1 144 | PM | 11 | 0.078 | 6.30 | 127.27 | [ |
| 黑果枸杞Lycium ruthenicum | LrSOS1 | MH454108 | 1 160 | / | 12 | 0.062 | 5.89 | 128.61 | [ |
| 欧洲油菜 Brassica napus | BnSOS1 | EU487184 | 1 142 | PM | 11 | 0.109 | 6.45 | 126.01 | [ |
| 胡杨 Populus euphratica | PeSOS1 | KX132789 | 1 145 | PM | 12 | 0.141 | 6.43 | 126.84 | [ |
| 篦麻 Ricinus communis | RcSOS1 | KX943304 | 1 143 | PM | 12 | 0.069 | 6.62 | 126.49 | [ |
| 菊芋 Helianthus tuberosus | HtSOS1 | KC410809 | 1 130 | PM | 12 | 0.091 | 6.00 | 125.00 | [ |
| 盐地碱蓬 Suaeda salsa | SsSOS1 | KF914414.1 | 1 169 | PM | 12 | 0.023 | 6.52 | 129.20 | [ |
| 绿豆 Vigna radiata | VrSOS1 | KC855193 | 1 046 | / | 11 | 0.119 | 6.20 | 116.10 | / |
| 节节麦 Aegilops tauschii | AtASOS1 | FN356231 | 1 142 | / | 12 | 0.10 | 6.85 | 126.18 | / |
| 短柄草Brachypodium sylvaticum | BsSOS1 | KF671967 | 1 140 | / | 12 | 0.078 | 7.63 | 126.77 | / |
| 海滨锦葵Kosteletzkya virginica | KvSOS1 | KJ577576 | 1 147 | PM | 12 | 0.073 | 6.18 | 127.56 | [ |
| 山萮菜Eutrema salsugineum | EsSOS1 | KF671959 | 1 127 | / | 12 | 0.064 | 6.47 | 124.51 | [ |
| 一粒小麦Triticum monococcum | TmSOS1 | FN356229 | 1 142 | / | 12 | 0.108 | 6.87 | 126.33 | / |
| 大叶补血草Limonium gmelinii | LgSOS1 | EU780458 | 1 151 | PM | 13 | 0.163 | 6.15 | 126.52 | [ |
| 藜麦Chenopodium quinoa | CqSOS1 | EU024570 | 1 158 | PM | 11 | 0.094 | 6.27 | 127.90 | [ |
| 霸王Zygophyllum xanthoxylon | ZxSOS1 | GU177864 | 1 153 | PM | 12 | 0.083 | 6.20 | 127.88 | [ |
| 番茄Solanum lycopersicum | SlSOS1-1 | AB675690 | 1 151 | PM | 12 | 0.069 | 5.85 | 127.54 | [ |
| SlSOS1-2 | NM_001247769 | 1 151 | PM | 13 | 0.083 | 5.89 | 127.50 | ||
| 苦荞麦Fagopyrum tataricum | FtSOS1 | KY659584 | 1 131 | PM | 12 | 0.090 | 6.13 | 125.88 | [ |
| 盐角草Salicornia brachiata | SbSOS1 | EU879059 | 1 159 | / | 11 | 0.037 | 5.92 | 128.55 | [ |
| 山丹 Lilium pumilum | LpSOS1 | OM650676 | 1 161 | PM | 12 | 0.107 | 6.59 | 129.14 | [ |
| 甘蔗Saccharum officinarum | ScSOS1 | KT003285 | 423 | Nucleus | / | -0.396 | 9.12 | 47.60 | [ |
| 小花碱茅Puccinellia tenuiflora | PtSOS1-1 | EF440291 | 1 137 | PM | 11 | 0.107 | 6.48 | 125.50 | [ |
| PtSOS1-2 | GQ452778 | 1 143 | PM | 13 | 0.103 | 6.82 | 126.16 | ||
| 海岛棉Gossypium barbadense | GbSOS1 | / | 1 152 | PM | / | / | 6.42 | 128.04 | [ |
Table 1 SOS1 gene in different plant species
| 物种名 Species name | 基因名称 Gene name | 基因登录号 GenBank accession No. | 氨基酸 Amino acid | 亚细胞定位 Sub-cellular localization | 跨膜结构域TMD | 亲水性平均值 GRAVY | 等电点 pI | 分子量 Mw/kD | 参考文献 Reference |
|---|---|---|---|---|---|---|---|---|---|
| 甜菜 Beta vulgaris | BvSOS1 | / | 1 162 | PM | 12 | 0.085 | 6.34 | 128.46 | Unpublished |
| 小盐芥thellungiella halophila | ThSOS1 | EF207775 | 1 146 | PM | 11 | 0.090 | 6.40 | 126.35 | [ |
| ThSOS1S | AB562331 | 473 | PM | 11 | 0.722 | 6.15 | 51.19 | [ | |
| 冰叶日中花 Mesembryanthemum crystallinum | McSOS1 | EF207776 | 1 115 | PM | 12 | 0.110 | 5.97 | 127.36 | [ |
| 花花柴 Karelinia caspia | KcSOS1 | MN481368 | 1 136 | PM | 12 | 0.070 | 5.97 | 126.02 | [ |
| 水稻Oryza sativa | OsSOS1-1 | AY785147 | 1 148 | PM | 12 | 0.046 | 6.77 | 127.92 | [ |
| OsSOS1-2 | MG602252 | 1 148 | PM | 13 | 0.047 | 6.77 | 127.90 | ||
| 小麦 Triticum aestivum | TaSOS1-1 | KJ563230 | 1 142 | PM | 12 | 0.110 | 7.00 | 126.22 | [ |
| TaSOS1-2 | AY326952 | 1 142 | PM | 13 | 0.106 | 7.40 | 126.18 | ||
| 大豆 Glycine max | GmSOS1 | JQ287499 | 1 143 | PM | 12 | 0.089 | 6.30 | 126.43 | [ |
| 文冠果Xanthoceras sorbifolium | XsSOS1 | / | 915 | PM | 6 | / | 6.19 | 102.50 | [ |
| 萝卜 Raphanus sativus | RsSOS1 | MZ484950 | 1 137 | PM | 12 | 0.093 | 6.71 | 125.62 | [ |
| 海马齿Sesuvium portulacastrum | SpSOS1 | JX674067 | 1 155 | PM | 12 | 0.063 | 6.08 | 127.95 | [ |
| 印度南瓜 Cucurbita pepo | CmaSOS1 | NW_019272028 | 1 142 | PM | 12 | 0.079 | 5.92 | 126.70 | [ |
| 陆地棉Gossypium hirsutum | GhSOS1 | KU994886 | 1 152 | PM | 12 | 0.061 | 6.32 | 128.18 | [ |
| 拟南芥Arabidopsis thaliana | AtSOS1 | AT2G01980 | 1 146 | PM | 13 | 0.098 | 7.62 | 127.19 | [ |
| 荞麦Fagopyrum esculentum | FeSOS1 | MH064255 | 1 132 | PM | 12 | 0.087 | 6.36 | 126.20 | [ |
| 白刺 Nitraria tangutorum | NtSOS1 | KC292267 | 1 163 | PM | 11 | 0.107 | 6.43 | 127.59 | [ |
| 黄花草木樨Melilotus officinalis | MoSOS1 | MG680455 | 957 | PM | 8 | 0.000 | 6.10 | 107.21 | [ |
| 黄花红砂Reaumuria trigyna | RtSOS1 | KC292265 | 1 145 | PM | 11 | 0.114 | 6.10 | 126.79 | [ |
| 木榄Bruguiera gymnorhiza | BgSOS1 | HM054521 | 1 153 | PM | 12 | 0.136 | 6.19 | 126.91 | [ |
| 盐生草Halogeton glomeratus | HgSOS1 | KT759142 | 1 165 | PM | 11 | 0.086 | 6.21 | 128.97 | / |
| 牡蒿 Artemisia japonica | AjSOS1 | KP896475 | 1 147 | PM | 12 | 0.061 | 6.08 | 127.04 | [ |
| 菠菜 Spinacia oleracea | SoSOS1 | HG799055 | 1 102 | / | 12 | 0.164 | 6.38 | 121.97 | [ |
| 葡萄 Vitis vinifera | VvSOS1 | NM_001281211 | 1 141 | / | 12 | 0.122 | 6.32 | 126.30 | [ |
| 黄瓜 Cucumis sativus | CsSOS1 | JQ655747 | 1 144 | PM | 11 | 0.078 | 6.30 | 127.27 | [ |
| 黑果枸杞Lycium ruthenicum | LrSOS1 | MH454108 | 1 160 | / | 12 | 0.062 | 5.89 | 128.61 | [ |
| 欧洲油菜 Brassica napus | BnSOS1 | EU487184 | 1 142 | PM | 11 | 0.109 | 6.45 | 126.01 | [ |
| 胡杨 Populus euphratica | PeSOS1 | KX132789 | 1 145 | PM | 12 | 0.141 | 6.43 | 126.84 | [ |
| 篦麻 Ricinus communis | RcSOS1 | KX943304 | 1 143 | PM | 12 | 0.069 | 6.62 | 126.49 | [ |
| 菊芋 Helianthus tuberosus | HtSOS1 | KC410809 | 1 130 | PM | 12 | 0.091 | 6.00 | 125.00 | [ |
| 盐地碱蓬 Suaeda salsa | SsSOS1 | KF914414.1 | 1 169 | PM | 12 | 0.023 | 6.52 | 129.20 | [ |
| 绿豆 Vigna radiata | VrSOS1 | KC855193 | 1 046 | / | 11 | 0.119 | 6.20 | 116.10 | / |
| 节节麦 Aegilops tauschii | AtASOS1 | FN356231 | 1 142 | / | 12 | 0.10 | 6.85 | 126.18 | / |
| 短柄草Brachypodium sylvaticum | BsSOS1 | KF671967 | 1 140 | / | 12 | 0.078 | 7.63 | 126.77 | / |
| 海滨锦葵Kosteletzkya virginica | KvSOS1 | KJ577576 | 1 147 | PM | 12 | 0.073 | 6.18 | 127.56 | [ |
| 山萮菜Eutrema salsugineum | EsSOS1 | KF671959 | 1 127 | / | 12 | 0.064 | 6.47 | 124.51 | [ |
| 一粒小麦Triticum monococcum | TmSOS1 | FN356229 | 1 142 | / | 12 | 0.108 | 6.87 | 126.33 | / |
| 大叶补血草Limonium gmelinii | LgSOS1 | EU780458 | 1 151 | PM | 13 | 0.163 | 6.15 | 126.52 | [ |
| 藜麦Chenopodium quinoa | CqSOS1 | EU024570 | 1 158 | PM | 11 | 0.094 | 6.27 | 127.90 | [ |
| 霸王Zygophyllum xanthoxylon | ZxSOS1 | GU177864 | 1 153 | PM | 12 | 0.083 | 6.20 | 127.88 | [ |
| 番茄Solanum lycopersicum | SlSOS1-1 | AB675690 | 1 151 | PM | 12 | 0.069 | 5.85 | 127.54 | [ |
| SlSOS1-2 | NM_001247769 | 1 151 | PM | 13 | 0.083 | 5.89 | 127.50 | ||
| 苦荞麦Fagopyrum tataricum | FtSOS1 | KY659584 | 1 131 | PM | 12 | 0.090 | 6.13 | 125.88 | [ |
| 盐角草Salicornia brachiata | SbSOS1 | EU879059 | 1 159 | / | 11 | 0.037 | 5.92 | 128.55 | [ |
| 山丹 Lilium pumilum | LpSOS1 | OM650676 | 1 161 | PM | 12 | 0.107 | 6.59 | 129.14 | [ |
| 甘蔗Saccharum officinarum | ScSOS1 | KT003285 | 423 | Nucleus | / | -0.396 | 9.12 | 47.60 | [ |
| 小花碱茅Puccinellia tenuiflora | PtSOS1-1 | EF440291 | 1 137 | PM | 11 | 0.107 | 6.48 | 125.50 | [ |
| PtSOS1-2 | GQ452778 | 1 143 | PM | 13 | 0.103 | 6.82 | 126.16 | ||
| 海岛棉Gossypium barbadense | GbSOS1 | / | 1 152 | PM | / | / | 6.42 | 128.04 | [ |
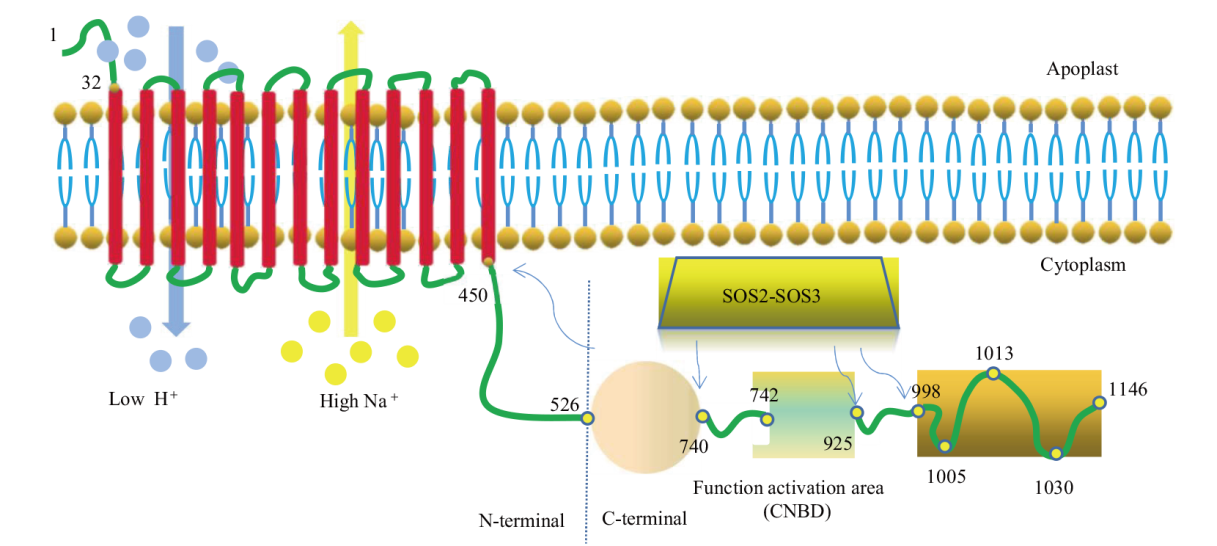
Fig. 2 Structural diagram of SOS1 protein The red box represents the TMD. The blue arrow indicates the conserved phosphorylation site recognized by SOS2-SOS3 complex

Fig. 3 Phylogenetic relationship of plant SOS1 gene family The source, name and accession number of the SOS1 family are as follows: Thellungiella salsuginea ThSOS1(ABN04857); Aeluropus littoralis AlSOS1(AEV89922); Aegilops speltoides AsSOS1(CAX83736); Betula platyphylla BpSOS1(ALV66191); Chrysanthemum crissum CcSOS1(BAR88076); Cymodocea nodosa CnoSOS1(CAD20320); Hesperis matronalis HmSOS1(AHL27892); Indosasa sinica IsSOS1(AGB06353); Populus pruinosa PpSOS1(AQN76710); Suaeda japonica SjSOS1(BAE95196); Solanum tuberosum StSOS1(XP_006364070); Triticum turgidum TtSOS1(ACB47885). Other species, name and accession number of the SOS genes are shown in Table 1
| [1] |
Ma L, Liu XH, Lv WJ, et al. Molecular mechanisms of plant responses to salt stress[J]. Front Plant Sci, 2022, 13: 934877.
doi: 10.3389/fpls.2022.934877 URL |
| [2] |
Foster KJ, Miklavcic SJ. A comprehensive biophysical model of ion and water transport in plant roots. II. clarifying the roles of SOS1 in the salt-stress response in Arabidopsis[J]. Front Plant Sci, 2019, 10: 1121.
doi: 10.3389/fpls.2019.01121 pmid: 31620152 |
| [3] |
Zhu JK. Salt and drought stress signal transduction in plants[J]. Annu Rev Plant Biol, 2002, 53: 247-273.
doi: 10.1146/arplant.2002.53.issue-1 URL |
| [4] |
Zhu M, Shabala L, Cuin TA, et al. Nax loci affect SOS1-like Na+/H+ exchanger expression and activity in wheat[J]. J Exp Bot, 2016, 67(3): 835-844.
doi: 10.1093/jxb/erv493 URL |
| [5] |
Ishikawa T, Shabala S. Control of xylem Na+ loading and transport to the shoot in rice and barley as a determinant of differential salinity stress tolerance[J]. Physiol Plant, 2019, 165(3): 619-631.
doi: 10.1111/ppl.12758 pmid: 29761494 |
| [6] |
Wu HH, Zhang XC, Giraldo JP, et al. It is not all about sodium: revealing tissue specificity and signalling roles of potassium in plant responses to salt stress[J]. Plant Soil, 2018, 431(1): 1-17.
doi: 10.1007/s11104-018-3770-y |
| [7] |
Isayenkov SV, Maathuis FJM. Plant salinity stress: many unanswered questions remain[J]. Front Plant Sci, 2019, 10: 80.
doi: 10.3389/fpls.2019.00080 pmid: 30828339 |
| [8] |
Rubio F, Nieves-Cordones M, Horie T, et al. Doing ‘business as usual’ comes with a cost: evaluating energy cost of maintaining plant intracellular K+ homeostasis under saline conditions[J]. New Phytol, 2020, 225(3): 1097-1104.
doi: 10.1111/nph.v225.3 URL |
| [9] |
Wu N, Li Z, Wu F, et al. Sex-specific photosynthetic capacity and Na+ homeostasis in Populus euphratica exposed to NaCl stress and AMF inoculation[J]. Front Plant Sci, 2022, 13: 1066954.
doi: 10.3389/fpls.2022.1066954 URL |
| [10] |
Gul Z, Tang ZH, Arif M, et al. An insight into abiotic stress and influx tolerance mechanisms in plants to cope in saline environments[J]. Biology, 2022, 11(4): 597.
doi: 10.3390/biology11040597 URL |
| [11] |
Gupta BK, Sahoo KK, Anwar K, et al. Silicon nutrition stimulates salt-overly sensitive(SOS)pathway to enhance salinity stress tolerance and yield in rice[J]. Plant Physiol Biochem, 2021, 166: 593-604.
doi: 10.1016/j.plaphy.2021.06.010 URL |
| [12] |
DeTar RA, Höhner R, Manavski N, et al. Loss of salt overly sensitive 1 prevents virescence in chloroplast K+/H+ efflux antiporter-deficient mutants[J]. Plant Physiol, 2022, 189(3): 1220-1225.
doi: 10.1093/plphys/kiac142 URL |
| [13] |
Wang YH, Pan CC, Chen QH, et al. Architecture and autoinhibitory mechanism of the plasma membrane Na+/H+ antiporter SOS1 in Arabidopsis[J]. Nat Commun, 2023, 14(1): 4487.
doi: 10.1038/s41467-023-40215-y |
| [14] |
Ponce KS, Meng LJ, Guo LB, et al. Advances in sensing, response and regulation mechanism of salt tolerance in rice[J]. Int J Mol Sci, 2021, 22(5): 2254.
doi: 10.3390/ijms22052254 URL |
| [15] |
Zhu JK, Liu J, Xiong L. Genetic analysis of salt tolerance in Arabidopsis. Evidence for a critical role of potassium nutrition[J]. Plant Cell, 1998, 10(7): 1181-1191.
doi: 10.1105/tpc.10.7.1181 pmid: 9668136 |
| [16] |
Shi H, Ishitani M, Kim C, et al. The Arabidopsis thaliana salt tolerance gene SOS1 encodes a putative Na+/H+ antiporter[J]. Proc Natl Acad Sci USA, 2000, 97(12): 6896-6901.
doi: 10.1073/pnas.120170197 pmid: 10823923 |
| [17] |
Wu SJ, Zhu JK. SOS1, a genetic locus essential for salt tolerance and potassium acquisition[J]. Plant Cell, 1996, 8(4): 617-627.
doi: 10.2307/3870339 URL |
| [18] |
Shi HZ, Quintero FJ, Pardo JM, et al. The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants[J]. Plant Cell, 2002, 14(2): 465-477.
doi: 10.1105/tpc.010371 URL |
| [19] |
Che BN, Cheng C, Fang JJ, et al. The recretohalophyte Tamarix TrSOS1 gene confers enhanced salt tolerance to transgenic hairy root composite cotton seedlings exhibiting virus-induced gene silencing of GhSOS1[J]. Int J Mol Sci, 2019, 20(12): 2930.
doi: 10.3390/ijms20122930 URL |
| [20] |
时丕彪, 洪立洲, 王军, 等. 印度南瓜Na+/H+逆向转运蛋白基因CmaSOS1的克隆与表达分析[J]. 核农学报, 2020, 34(12): 2638-2646.
doi: 10.11869/j.issn.100-8551.2020.12.2638 |
| Shi PB, Hong LZ, Wang J, et al. Cloning and expression analysis of a Na+/H+antiporter gene CmaSOS1 in Cucurbita maxima[J]. J Nucl Agric Sci, 2020, 34(12): 2638-2646. | |
| [21] |
Shahzad B, Shabala L, Zhou MX, et al. Comparing essentiality of SOS1-mediated Na+ exclusion in salinity tolerance between cultivated and wild rice species[J]. Int J Mol Sci, 2022, 23(17): 9900.
doi: 10.3390/ijms23179900 URL |
| [22] |
El Mahi H, Pérez-Hormaeche J, Luca AD, et al. A critical role of sodium flux via the plasma membrane Na+/H+ exchanger SOS1 in the salt tolerance of rice[J]. Plant Physiol, 2019, 180(2): 1046-1065.
doi: 10.1104/pp.19.00324 URL |
| [23] |
Tomita M, Yamashita M, Omichi A. Gene structure of three kinds of vacuolar-type Na+/H+ antiporters including TaNHX2 transcribed in bread wheat[J]. Genet Mol Biol, 2021, 44(1): e20200207.
doi: 10.1590/1678-4685-gmb-2020-0207 URL |
| [24] |
Jiang W, Pan R, Buitrago S, et al. Conservation and divergence of the TaSOS1 gene family in salt stress response in wheat(Triticum aestivum L.)[J]. Physiol Mol Biol Plants, 2021, 27(6): 1245-1260.
doi: 10.1007/s12298-021-01009-y |
| [25] |
Li SJ, Wu GQ, Lin LY. AKT1, HAK5, SKOR, HKT1;5, SOS1 and NHX1 synergistically control Na+ and K+ homeostasis in sugar beet(Beta vulgaris L.)seedlings under saline conditions[J]. J Plant Biochem Biotechnol, 2022, 31(1): 71-84.
doi: 10.1007/s13562-021-00656-2 |
| [26] |
Zhang WT, Li JX, Dong JH, et al. RsSOS1 responding to salt stress might be involved in regulating salt tolerance by maintaining Na+ homeostasis in radish(Raphanus sativus L.)[J]. Horticulturae, 2021, 7(11): 458.
doi: 10.3390/horticulturae7110458 URL |
| [27] | 张婷婷, 康宇乾, 李雨欣, 等. 非生物胁迫下海马齿SpSOS1基因的表达及响应ABA模式分析[J]. 分子植物育种, 2021, 19(13): 4371-4377. |
| Zhang TT, Kang YQ, Li YX, et al. Expression pattern analysis of SpSOS1 from Sesuvium portulacastrum under abiotic stresses and the response to ABA[J]. Mol Plant Breed, 2021, 19(13): 4371-4377. | |
| [28] |
Zhao CY, William D, Sandhu D. Isolation and characterization of salt overly sensitive family genes in spinach[J]. Physiol Plant, 2021, 171(4): 520-532.
doi: 10.1111/ppl.v171.4 URL |
| [29] |
Brindha C, Vasantha S, Raja AK, et al. Characterization of the salt overly sensitive pathway genes in sugarcane under salinity stress[J]. Physiol Plant, 2021, 171(4): 677-687.
doi: 10.1111/ppl.13245 pmid: 33063359 |
| [30] |
Yang YQ, Han XL, Ma L, et al. Dynamic changes of phosphatidylinositol and phosphatidylinositol 4-phosphate levels modulate H+-ATPase and Na+/H+ antiporter activities to maintain ion homeostasis in Arabidopsis under salt stress[J]. Mol Plant, 2021, 14(12): 2000-2014.
doi: 10.1016/j.molp.2021.07.020 URL |
| [31] |
Xie Q, Zhou Y, Jiang XY. Structure, function, and regulation of the plasma membrane Na+/H+ antiporter salt overly sensitive 1 in plants[J]. Front Plant Sci, 2022, 13: 866265.
doi: 10.3389/fpls.2022.866265 URL |
| [32] |
Zhang MH, Cao JF, Zhang TX, et al. A putative plasma membrane Na+/H+ antiporter GmSOS1 is critical for salt stress tolerance in Glycine max[J]. Front Plant Sci, 2022, 13: 870695.
doi: 10.3389/fpls.2022.870695 URL |
| [33] | 陈叶, 丁健, 阮成江, 等. 文冠果Na+/H+逆向转运蛋白基因XsSOS1的克隆与表达分析[J]. 分子植物育种, 2022. http://kns.cnki.net/kcms/detail/46.1068.S.20220803.1806.016.html. |
| Chen Y, Ding J, Ruan CJ, et al. Cloning and expression analysis of a Na+/H+ antiporter gene XsSOS1 in Xanthoceras sorbifolium[J]. Mol Plant Breed, 2022. http://kns.cnki.net/kcms/detail/46.1068.S.20220803.1806.016.html. | |
| [34] |
Arciniegas Vega JP, Melino VJ. Uncovering natural genetic variants of the SOS pathway to improve salinity tolerance in maize[J]. New Phytol, 2022, 236(2): 313-315.
doi: 10.1111/nph.18422 pmid: 35977055 |
| [35] |
Xu FC, Wang MJ, Guo YW, et al. The Na+/H+ antiporter GbSOS1 interacts with SIP5 and regulates salt tolerance in Gossypium barbadense[J]. Plant Sci, 2023, 330: 111658.
doi: 10.1016/j.plantsci.2023.111658 URL |
| [36] |
Liu ZY, Xie QJ, Tang FF, et al. The ThSOS3 gene improves the salt tolerance of transgenic Tamarix hispida and Arabidopsis thaliana[J]. Front Plant Sci, 2021, 11: 597480.
doi: 10.3389/fpls.2020.597480 URL |
| [37] |
Taji T, Komatsu K, Katori T, et al. Comparative genomic analysis of 1047 completely sequenced cDNAs from an Arabidopsis-related model halophyte, Thellungiella halophila[J]. BMC Plant Biol, 2010, 10: 261.
doi: 10.1186/1471-2229-10-261 URL |
| [38] |
Cosentino C, Fischer-Schliebs E, Bertl A, et al. Na+/H+ antiporters are differentially regulated in response to NaCl stress in leaves and roots of Mesembryanthemum crystallinum[J]. New Phytol, 2010, 186(3): 669-680.
doi: 10.1111/j.1469-8137.2010.03208.x pmid: 20298477 |
| [39] |
Guo Q, Meng L, Han JW, et al. SOS1 is a key systemic regulator of salt secretion and K+/Na+ homeostasis in the recretohalophyte Karelinia caspia[J]. Environ Exp Bot, 2020, 177: 104098.
doi: 10.1016/j.envexpbot.2020.104098 URL |
| [40] |
Chen XG, Lu XK, Shu N, et al. GhSOS1, a plasma membrane Na+/H+ antiporter gene from upland cotton, enhances salt tolerance in transgenic Arabidopsis thaliana[J]. PLoS One, 2017, 12(7): e0181450.
doi: 10.1371/journal.pone.0181450 URL |
| [41] |
郑琳琳, 张慧荣, 贺龙梅, 等. 唐古特白刺质膜Na+/H+逆向转运蛋白基因的克隆与表达分析[J]. 草业学报, 2013, 22(4): 179-186.
doi: 10.11686/cyxb20130422 |
| Zheng LL, Zhang HR, He LM, et al. Isolation and expression analysis of a plasma membrane Na+/H+ antiporter from Nitraria tangutorum[J]. Acta Prataculturae Sin, 2013, 22(4): 179-186. | |
| [42] |
黄坤勇, 李杉杉, 郭强, 等. 黄花草木樨MoSOS1基因克隆及表达分析[J]. 生物技术通报, 2017, 33(9): 120-130.
doi: 10.13560/j.cnki.biotech.bull.1985.2017-0364 |
| Huang KY, Li SS, Guo Q, et al. Cloning and expression analysis of MoSOS1 gene in Melilotus officinalis[J]. Biotechnol Bull, 2017, 33(9): 120-130. | |
| [43] |
Wang JY, Li Q, Zhang M, et al. The high pH value of alkaline salt destroys the root membrane permeability of Reaumuria trigyna and leads to its serious physiological decline[J]. J Plant Res, 2022, 135(6): 785-798.
doi: 10.1007/s10265-022-01410-y |
| [44] | 荆海瑜, 周扬, 郭晓颖, 等. 木榄细胞膜Na+/H+逆向运输蛋白BgSOS1功能的初步验证[J]. 海南大学学报: 自然科学版, 2014, 32(3): 252-259, 269. |
| Jing HY, Zhou Y, Guo XY, et al. Preliminary analysis of a Na+/H+ antiporter BgSOS1 at plasma membrane of Bruguiera gymnorrhiza[J]. Nat Sci J Hainan Univ, 2014, 32(3): 252-259, 269. | |
| [45] |
Gao JJ, Sun J, Cao PP, et al. Variation in tissue Na+ content and the activity of SOS1 genes among two species and two related Genera of Chrysanthemum[J]. BMC Plant Biol, 2016, 16: 98.
doi: 10.1186/s12870-016-0781-9 URL |
| [46] |
Venturini L, Ferrarini A, Zenoni S, et al. De novo transcriptome characterization of Vitis vinifera cv. Corvina unveils varietal diversity[J]. BMC Genomics, 2013, 14: 41.
doi: 10.1186/1471-2164-14-41 pmid: 23331995 |
| [47] |
Wang S, Li Z, Rui R, et al. Cloning and characterization of a plasma membrane Na+/H+ antiporter gene from Cucumis sativus[J]. Russ J Plant Physiol, 2013, 60(3): 330-336.
doi: 10.1134/S102144371303014X URL |
| [48] |
Hu J, Hu XK, Zhang HW, et al. Moderate NaCl alleviates osmotic stress in Lycium ruthenicum[J]. Plant Growth Regul, 2022, 96(1): 25-35.
doi: 10.1007/s10725-021-00754-0 |
| [49] |
Meng KB, Wu YX. Footprints of divergent evolution in two Na+/H+ type antiporter gene families(NHX and SOS1)in the genus Populus[J]. Tree Physiol, 2018, 38(6): 813-824.
doi: 10.1093/treephys/tpx173 URL |
| [50] | 李平, 王晓宇, 徐惠, 等. 蓖麻质膜型Na+/H+逆向转运蛋白基因(RcSOS1)克隆及表达载体构建[J]. 分子植物育种, 2018, 16(10): 3182-3189. |
| Li P, Wang XY, Xu H, et al. Cloning and expression vectors establishment of plasma membrane Na+/H+ antiporter gene(RcSOS1)in castor(Ricinus communis L.)[J]. Mol Plant Breed, 2018, 16(10): 3182-3189. | |
| [51] |
Li Q, Tang Z, Hu YB, et al. Functional analyses of a putative plasma membrane Na+/H+ antiporter gene isolated from salt tolerant Helianthus tuberosus[J]. Mol Biol Rep, 2014, 41(8): 5097-5108.
doi: 10.1007/s11033-014-3375-3 URL |
| [52] |
Guo JR, Dong XX, Han GL, et al. Salt-enhanced reproductive development of Suaeda salsa L. coincided with ion transporter gene upregulation in flowers and increased pollen K+ content[J]. Front Plant Sci, 2019, 10: 333.
doi: 10.3389/fpls.2019.00333 URL |
| [53] |
Wang WY, Liu YQ, Duan HR, et al. SsHKT1;1 is coordinated with SsSOS1 and SsNHX1 to regulate Na+ homeostasis in Suaeda salsa under saline conditions[J]. Plant Soil, 2020, 449(1/2): 117-131.
doi: 10.1007/s11104-020-04463-x |
| [54] | Wang HY, Tang XL, Shao CY, et al. Molecular cloning and bioinformatics analysis of a new plasma membrane Na+/H+ antiporter gene from the halophyte Kosteletzkya virginica[J]. Sci World J, 2014, 2014: 141675. |
| [55] |
Jarvis DE, Ryu CH, Beilstein MA, et al. Distinct roles for SOS1 in the convergent evolution of salt tolerance in Eutrema salsugineum and Schrenkiella parvula[J]. Mol Biol Evol, 2014, 31(8): 2094-2107.
doi: 10.1093/molbev/msu152 pmid: 24803640 |
| [56] | 马苏勇, 祝建波, 王爱英, 等. 大叶补血草质膜Na+/H+逆向转运蛋白基因SOS1的克隆及对番茄的遗传转化[J]. 生物技术通报, 2010(9): 111-115. |
| Ma SY, Zhu JB, Wang AY, et al. Cloning and genetic transformation of tomato with SOS1 gene from Limonium gmelinii[J]. Biotechnol Bull, 2010(9): 111-115. | |
| [57] |
Abd El-Moneim D, ELsarag EIS, Aloufi S, et al. Quinoa(Chenopodium quinoa Willd.): genetic diversity according to ISSR and SCoT markers, relative gene expression, and morpho-physiological variation under salinity stress[J]. Plants, 2021, 10(12): 2802.
doi: 10.3390/plants10122802 URL |
| [58] |
Ma Q, Li YX, Yuan HJ, et al. ZxSOS1 is essential for long-distance transport and spatial distribution of Na+ and K+ in the xerophyte Zygophyllum xanthoxylum[J]. Plant Soil, 2014, 374(1): 661-676.
doi: 10.1007/s11104-013-1891-x URL |
| [59] |
Wang Z, Hong YC, Li YM, et al. Natural variations in SlSOS1 contribute to the loss of salt tolerance during tomato domestication[J]. Plant Biotechnol J, 2021, 19(1): 20-22.
doi: 10.1111/pbi.v19.1 URL |
| [60] |
Lu QH, Wang YQ, Xu JP, et al. Effect of ABA on physiological characteristics and expression of salt tolerance-related genes in Tartary buckwheat[J]. Acta Physiol Plant, 2021, 43(5): 1-11.
doi: 10.1007/s11738-020-03172-3 |
| [61] |
Yadav NS, Shukla PS, Jha A, et al. The SbSOS1 gene from the extreme halophyte Salicornia brachiata enhances Na+ loading in xylem and confers salt tolerance in transgenic tobacco[J]. BMC Plant Biol, 2012, 12: 188.
doi: 10.1186/1471-2229-12-188 |
| [62] |
Yang Y, Xu LF, Li WX, et al. A Na+/H+ antiporter-encoding salt overly sensitive 1 gene, LpSOS1, involved in positively regulating the salt tolerance in Lilium pumilum[J]. Gene, 2023, 874: 147485.
doi: 10.1016/j.gene.2023.147485 URL |
| [63] | Kaewjiw N, Laksana C, Chanprame S. Cloning and identification of salt overly sensitive(SOS1)gene of sugarcane[J]. Int J Agric Biol, 2018, 20(7): 1569-1574. |
| [64] |
Han QQ, Wang YP, Li J, et al. The mechanistic basis of sodium exclusion in Puccinellia tenuiflora under conditions of salinity and potassium deprivation[J]. Plant J, 2022, 112(2): 322-338.
doi: 10.1111/tpj.v112.2 URL |
| [65] |
Núñez-Ramírez R, Sánchez-Barrena MJ, Villalta I, et al. Structural insights on the plant salt-overly-sensitive 1(SOS1)Na+/H+ antiporter[J]. J Mol Biol, 2012, 424(5): 283-294.
doi: 10.1016/j.jmb.2012.09.015 pmid: 23022605 |
| [66] |
Quintero FJ, Martinez-Atienza J, Villalta I, et al. Activation of the plasma membrane Na/H antiporter salt-overly-sensitive 1(SOS1)by phosphorylation of an auto-inhibitory C-terminal domain[J]. Proc Natl Acad Sci USA, 2011, 108(6): 2611-2616.
doi: 10.1073/pnas.1018921108 URL |
| [67] |
Goswami P, Paulino C, Hizlan D, et al. Structure of the archaeal Na+/H+ antiporter NhaP1 and functional role of transmembrane helix 1[J]. EMBO J, 2011, 30(2): 439-449.
doi: 10.1038/emboj.2010.321 URL |
| [68] |
Zheng M, Li JP, Zeng CW, et al. Subgenome-biased expression and functional diversification of a Na+/H+ antiporter homoeologs in salt tolerance of polyploid wheat[J]. Front Plant Sci, 2022, 13: 1072009.
doi: 10.3389/fpls.2022.1072009 URL |
| [69] |
Ma X, Li QH, Yu YN, et al. The CBL-CIPK pathway in plant response to stress signals[J]. Int J Mol Sci, 2020, 21(16): 5668.
doi: 10.3390/ijms21165668 URL |
| [70] |
Tang RJ, Wang C, Li KL, et al. The CBL-CIPK calcium signaling network: unified paradigm from 20 years of discoveries[J]. Trends Plant Sci, 2020, 25(6): 604-617.
doi: 10.1016/j.tplants.2020.01.009 URL |
| [71] |
Wu GQ, Wang JL, Li SJ. Genome-wide identification of Na+/H+ antiporter(NHX)genes in sugar beet(Beta vulgaris L.) and their regulated expression under salt stress[J]. Genes, 2019, 10(5): 401.
doi: 10.3390/genes10050401 URL |
| [72] |
Hao SH, Wang YR, Yan YX, et al. A review on plant responses to salt stress and their mechanisms of salt resistance[J]. Horticulturae, 2021, 7(6): 132.
doi: 10.3390/horticulturae7060132 URL |
| [73] |
Park HJ, Qiang Z, Kim WY, et al. Diurnal and circadian regulation of salt tolerance in Arabidopsis[J]. J Plant Biol, 2016, 59(6): 569-578.
doi: 10.1007/s12374-016-0317-8 URL |
| [74] |
Chen MX, She ZY, Aslam M, et al. Genomic insights of the WRKY genes in kenaf(Hibiscus cannabinus L.) reveal that HcWRKY44 improves the plant's tolerance to the salinity stress[J]. Front Plant Sci, 2022, 13: 984233.
doi: 10.3389/fpls.2022.984233 URL |
| [75] |
Wu X, Xu JN, Meng XN, et al. Linker histone variant HIS1-3 and WRKY1 oppositely regulate salt stress tolerance in Arabidopsis[J]. Plant Physiol, 2022, 189(3): 1833-1847.
doi: 10.1093/plphys/kiac174 URL |
| [76] |
Zhang HF, Guo JB, Chen XQ, et al. Pepper bHLH transcription factor CabHLH035 contributes to salt tolerance by modulating ion homeostasis and proline biosynthesis[J]. Hortic Res, 2022, 9: uhac203.
doi: 10.1093/hr/uhac203 URL |
| [77] |
Fu HQ, Yu X, Jiang YY, et al. SALT OVERLY SENSITIVE 1 is inhibited by clade D protein phosphatase 2C D6 and D7 in Arabidopsis thaliana[J]. Plant Cell, 2023, 35(1): 279-297.
doi: 10.1093/plcell/koac283 URL |
| [78] |
Duscha K, Martins Rodrigues C, Müller M, et al. 14-3-3 proteins and other candidates form protein-protein interactions with the cytosolic C-terminal end of SOS1 affecting its transport activity[J]. Int J Mol Sci, 2020, 21(9): 3334.
doi: 10.3390/ijms21093334 URL |
| [79] |
Kim WY, Ali Z, Park HJ, et al. Release of SOS2 kinase from sequestration with GIGANTEA determines salt tolerance in Arabidopsis[J]. Nat Commun, 2013, 4: 1352.
doi: 10.1038/ncomms2357 |
| [80] |
Li JF, Zhou HP, Zhang Y, et al. The GSK3-like kinase BIN2 is a molecular switch between the salt stress response and growth recovery in Arabidopsis thaliana[J]. Dev Cell, 2020, 55(3): 367-380.e6.
doi: 10.1016/j.devcel.2020.08.005 URL |
| [81] |
Amirbakhtiar N, Ismaili A, Ghaffari MR, et al. Transcriptome analysis of bread wheat leaves in response to salt stress[J]. PLoS One, 2021, 16(7): e0254189.
doi: 10.1371/journal.pone.0254189 URL |
| [82] |
Zhou M, Wang W. SOS1 safeguards plant circadian rhythm against daily salt fluctuations[J]. Proc Natl Acad Sci USA, 2022, 119(36): e2212950119.
doi: 10.1073/pnas.2212950119 URL |
| [83] |
Pei SY, Liu YT, Li WK, et al. OSCA1 is an osmotic specific sensor: a method to distinguish Ca2+-mediated osmotic and ionic perception[J]. New Phytol, 2022, 235(4): 1665-1678.
doi: 10.1111/nph.v235.4 URL |
| [84] |
Yuan F, Yang HM, Xue Y, et al. OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in Arabidopsis[J]. Nature, 2014, 514(7522): 367-371.
doi: 10.1038/nature13593 |
| [85] |
Jiang ZH, Zhou XP, Tao M, et al. Plant cell-surface GIPC sphingolipids sense salt to trigger Ca2+ influx[J]. Nature, 2019, 572(7769): 341-346.
doi: 10.1038/s41586-019-1449-z |
| [86] |
Yin XC, Xia YQ, Xie Q, et al. The protein kinase complex CBL10-CIPK8-SOS1 functions in Arabidopsis to regulate salt tolerance[J]. J Exp Bot, 2020, 71(6): 1801-1814.
doi: 10.1093/jxb/erz549 URL |
| [87] |
Zhu JK. Abiotic stress signaling and responses in plants[J]. Cell, 2016, 167(2): 313-324.
doi: 10.1016/j.cell.2016.08.029 URL |
| [88] |
Guo Y, Liu Y, Zhang Y, et al. Effects of exogenous calcium on adaptive growth, photosynthesis, ion homeostasis and phenolics of Gleditsia sinensis Lam. plants under salt stress[J]. Agriculture, 2021, 11(10): 978.
doi: 10.3390/agriculture11100978 URL |
| [89] |
Fraile-Escanciano A, Kamisugi Y, Cuming AC, et al. The SOS1 transporter of Physcomitrella patens mediates sodium efflux in planta[J]. New Phytol, 2010, 188(3): 750-761.
doi: 10.1111/j.1469-8137.2010.03405.x pmid: 20696009 |
| [90] |
Gupta A, Shaw BP, Sahu BB. Post-translational regulation of the membrane transporters contributing to salt tolerance in plants[J]. Funct Plant Biol, 2021, 48(12): 1199-1212.
doi: 10.1071/FP21153 pmid: 34665998 |
| [91] |
Quintero FJ, Ohta M, Shi HZ, et al. Reconstitution in yeast of the Arabidopsis SOS signaling pathway for Na+ homeostasis[J]. Proc Natl Acad Sci USA, 2002, 99(13): 9061-9066.
doi: 10.1073/pnas.132092099 URL |
| [92] |
Zhou Y, Lai ZS, Yin XC, et al. Hyperactive mutant of a wheat plasma membrane Na+/H+ antiporter improves the growth and salt tolerance of transgenic tobacco[J]. Plant Sci, 2016, 253: 176-186.
doi: 10.1016/j.plantsci.2016.09.016 URL |
| [93] |
Xie Q, Yang Y, Wang Y, et al. The calcium sensor CBL10 negatively regulates plasma membrane H+-ATPase activity and alkaline stress response in Arabidopsis[J]. Environ Exp Bot, 2022, 194: 104752.
doi: 10.1016/j.envexpbot.2021.104752 URL |
| [94] |
Park HJ, Kim WY, Yun DJ. A new insight of salt stress signaling in plant[J]. Mol Cells, 2016, 39(6): 447-459.
doi: 10.14348/molcells.2016.0083 pmid: 27239814 |
| [95] |
Du WM, Lin HX, Chen S, et al. Phosphorylation of SOS3-like calcium-binding proteins by their interacting SOS2-like protein kinases is a common regulatory mechanism in Arabidopsis[J]. Plant Physiol, 2011, 156(4): 2235-2243.
doi: 10.1104/pp.111.173377 URL |
| [96] |
谢玲玲, 王金龙, 伍国强. 植物CBL-CIPK信号系统响应非生物胁迫的调控机制[J]. 植物学报, 2021, 56(5): 614-626.
doi: 10.11983/CBB21024 |
| Xie LL, Wang JL, Wu GQ. Regulatory mechanisms of the plant CBL-CIPK signaling system in response to abiotic stress[J]. Chin Bull Bot, 2021, 56(5): 614-626. | |
| [97] |
Wang Q, Guan C, Wang P, et al. The effect of AtHKT1;1 or AtSOS1 mutation on the expressions of Na+ or K+ transporter genes and ion homeostasis in Arabidopsis thaliana under salt stress[J]. Int J Mol Sci, 2019, 20(5): 1085.
doi: 10.3390/ijms20051085 URL |
| [98] |
Venkataraman G, Shabala S, Véry AA, et al. To exclude or to accumulate? Revealing the role of the sodium HKT1;5 transporter in plant adaptive responses to varying soil salinity[J]. Plant Physiol Biochem, 2021, 169: 333-342.
doi: 10.1016/j.plaphy.2021.11.030 URL |
| [99] |
Riedelsberger J, Miller JK, Valdebenito-Maturana B, et al. Plant HKT channels: an updated view on structure, function and gene regulation[J]. Int J Mol Sci, 2021, 22(4): 1892.
doi: 10.3390/ijms22041892 URL |
| [100] | Gu WT, Zhou LB, Liu RY, et al. Synergistic responses of NHX, AKT1, and SOS1 in the control of Na+ homeostasis in sweet sorghum mutants induced by 12C6+-ion irradiation[J]. Nucl Sci Tech, 2018, 29(1): 10. |
| [101] |
Sun QA, Yamada T, Han YL, et al. Differential responses of NHX1 and SOS1 gene expressions to salinity in two Miscanthus sinensis anderss. accessions with different salt tolerance[J]. Phyton, 2021, 90(3): 827-836.
doi: 10.32604/phyton.2021.013805 URL |
| [102] |
Olías R, Eljakaoui Z, Li J, et al. The plasma membrane Na+/H+ antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of Na+ between plant organs[J]. Plant Cell Environ, 2009, 32(7): 904-916.
doi: 10.1111/pce.2009.32.issue-7 URL |
| [103] |
Ishikawa T, Shabala L, Zhou MX, et al. Comparative analysis of root Na+ relation under salinity between Oryza sativa and Oryza coarctata[J]. Plants, 2022, 11(5): 656.
doi: 10.3390/plants11050656 URL |
| [104] |
Fan YF, Yin XC, Xie Q, et al. Co-expression of SpSOS1 and SpAHA1 in transgenic Arabidopsis plants improves salinity tolerance[J]. BMC Plant Biol, 2019, 19(1): 74.
doi: 10.1186/s12870-019-1680-7 |
| [105] | Shabala S, Munns R. Salinity stress: physiological constraints and adaptive mechanisms[M]// Plant Stress Physiology. UK: CABI, 2017: 24-63. |
| [106] |
Katiyar-Agarwal S, Zhu JH, Kim K, et al. The plasma membrane Na+/H+ antiporter SOS1 interacts with RCD1 and functions in oxidative stress tolerance in Arabidopsis[J]. Proc Natl Acad Sci USA, 2006, 103(49): 18816-18821.
pmid: 17023541 |
| [107] |
Feki K, Tounsi S, Masmoudi K, et al. The durum wheat plasma membrane Na+/H+ antiporter SOS1 is involved in oxidative stress response[J]. Protoplasma, 2017, 254(4): 1725-1734.
doi: 10.1007/s00709-016-1066-8 URL |
| [108] |
Zhou Y, Yin XC, Wan SM, et al. The Sesuvium portulacastrum plasma membrane Na+/H+ antiporter SpSOS1 complemented the salt sensitivity of transgenic Arabidopsis sos1 mutant plants[J]. Plant Mol Biol Rep, 2018, 36(4): 553-563.
doi: 10.1007/s11105-018-1099-6 |
| [109] |
Xu XD, Yuan L, Xie QG. The circadian clock ticks in plant stress responses[J]. Stress Biol, 2022, 2(1): 15.
doi: 10.1007/s44154-022-00040-7 pmid: 37676516 |
| [110] |
Yang YQ, Wu YJ, Ma L, et al. The Ca2+ sensor SCaBP3/CBL7 modulates plasma membrane H+-ATPase activity and promotes alkali tolerance in Arabidopsis[J]. Plant Cell, 2019, 31(6): 1367-1384.
doi: 10.1105/tpc.18.00568 URL |
| [111] |
Oh DH, Lee SY, Bressan RA, et al. Intracellular consequences of SOS1 deficiency during salt stress[J]. J Exp Bot, 2010, 61(4): 1205-1213.
doi: 10.1093/jxb/erp391 URL |
| [112] |
Zhou JY, Hao DL, Yang GZ. Regulation of cytosolic pH: the contributions of plant plasma membrane H+-ATPases and multiple transporters[J]. Int J Mol Sci, 2021, 22(23): 12998.
doi: 10.3390/ijms222312998 URL |
| [113] | Guo KM, Babourina O, Rengel Z. Na+/H+ antiporter activity of the SOS1 gene: lifetime imaging analysis and electrophysiological studies on Arabidopsis seedlings[J]. Physiol Plant, 2009, 137(2): 155-165. |
| [114] |
Cha JY, Kim J, Jeong SY, et al. The Na+/H+ antiporter salt overly sensitive 1 regulates salt compensation of circadian rhythms by stabilizing gigantea in arabidopsis[J]. Proc Natl Acad Sci USA, 2022, 119(33): e2207275119.
doi: 10.1073/pnas.2207275119 URL |
| [115] |
Ma YC, Wang L, Wang JY, et al. Isolation and expression analysis of salt overly sensitive gene family in grapevine(Vitis vinifera)in response to salt and PEG stress[J]. PLoS One, 2019, 14(3): e0212666.
doi: 10.1371/journal.pone.0212666 URL |
| [116] |
Shahzad B, Yun P, Rasouli F, et al. Root K+ homeostasis and signalling as a determinant of salinity stress tolerance in cultivated and wild rice species[J]. Environ Exp Bot, 2022, 201: 104944.
doi: 10.1016/j.envexpbot.2022.104944 URL |
| [117] |
Chai HX, Guo JF, Zhong YL, et al. The plasma-membrane polyamine transporter PUT3 is regulated by the Na+/H+ antiporter SOS1 and protein kinase SOS2[J]. New Phytol, 2020, 226(3): 785-797.
doi: 10.1111/nph.v226.3 URL |
| [118] |
Farooq M, Park JR, Jang YH, et al. Rice cultivars under salt stress show differential expression of genes related to the regulation of Na+/K+ balance[J]. Front Plant Sci, 2021, 12: 680131.
doi: 10.3389/fpls.2021.680131 URL |
| [119] |
Yue YS, Zhang MC, Zhang JC, et al. SOS1 gene overexpression increased salt tolerance in transgenic tobacco by maintaining a higher K+/Na+ ratio[J]. J Plant Physiol, 2012, 169(3): 255-261.
doi: 10.1016/j.jplph.2011.10.007 URL |
| [120] |
Akrimi R, Hajlaoui H, Batelli G, et al. Electromagnetic water enhanced metabolism and agro-physiological responses of potato(Solanum tuberosum L)under saline conditions[J]. J Agron Crop Sci, 2021, 207(1): 44-58.
doi: 10.1111/jac.v207.1 URL |
| [121] |
Awaji SM, Hanjagi PS, Pushpa BN, et al. Overexpression of plasma membrane Na+/H+ antiporter OsSOS1 gene improves salt tolerance in transgenic rice plants[J]. Oryza, 2020, 57(4): 277-287.
doi: 10.35709/ory.2019.56.2 URL |
| [122] |
Rao YR, Ansari MW, Sahoo RK, et al. Salicylic acid modulates ACS, NHX1, sos1 and HKT1;2 expression to regulate ethylene overproduction and Na+ ions toxicity that leads to improved physiological status and enhanced salinity stress tolerance in tomato plants cv. Pusa Ruby[J]. Plant Signal Behav, 2021, 16(11): 1950888.
doi: 10.1080/15592324.2021.1950888 URL |
| [123] |
Chang PJ, Wu Z, Song NN, et al. Identification MdeSOS1 in Magnolia denudata and its function in response to salt stress[J]. J Plant Interact, 2020, 15(1): 417-426.
doi: 10.1080/17429145.2020.1843721 URL |
| [124] |
Hichri I, Muhovski Y, Clippe A, et al. SlDREB2, a tomato dehydration-responsive element-binding 2 transcription factor, mediates salt stress tolerance in tomato and Arabidopsis[J]. Plant Cell Environ, 2016, 39(1): 62-79.
doi: 10.1111/pce.v39.1 URL |
| [1] | KANG Ling-yun, HAN Lu-lu, HAN De-ping, CHEN Jian-sheng, GAN Han-ling, XING Kai, MA You-ji, CUI Kai. Effect of Melatonin on Protecting the Jejunum Mucosal Epithelial Cells from Oxidative Stress Damage [J]. Biotechnology Bulletin, 2023, 39(9): 291-299. |
| [2] | HAN Zhi-yang, JIA Zi-miao, LIANG Qiu-ju, WANG Ke, TANG Hua-li, YE Xing-guo, ZHANG Shuang-xi. Salt Tolerance at Seedling Stage and Analysis of Selenium and Folic Acid Content in Seeds in Two Sets of Wheat-Dasypyrum villosum Chromosom Additional Lines [J]. Biotechnology Bulletin, 2023, 39(8): 185-193. |
| [3] | WANG Shuai, FENG Yu-mei, BAI Miao, DU Wei-jun, YUE Ai-qin. Functional Analysis of Soybean Gene GmHMGR Responding to Exogenous Hormones and Abiotic Stresses [J]. Biotechnology Bulletin, 2023, 39(7): 131-142. |
| [4] | LI Zhi-qi, YUAN Yue, MIAO Rong-qing, PANG Qiu-ying, ZHANG Ai-qin. Melatonin Contents in Eutrema salsugineum and Arabidopsis thaliana Under Salt Stress, and Expression Pattern Analysis of Synthesis Related Genes [J]. Biotechnology Bulletin, 2023, 39(5): 142-151. |
| [5] | WANG Qi, HU Zhe, FU Wei, LI Guang-zhe, HAO Lin. Regulation of Burkholderia sp. GD17 on the Drought Tolerance of Cucumber Seedlings [J]. Biotechnology Bulletin, 2023, 39(3): 163-175. |
| [6] | CHEN Yi-bo, YANG Wan-ming, YUE Ai-qin, WANG Li-xiang, DU Wei-jun, WANG Min. Construction of Soybean Genetic Map Based on SLAF Markers and QTL Mapping Analysis of Salt Tolerance at Seedling Stage [J]. Biotechnology Bulletin, 2023, 39(2): 70-79. |
| [7] | YAN Xiong-ying, WANG Zhen, WANG Xia, YANG Shi-hui. Microbial Sulfur Metabolism and Stress Resistance [J]. Biotechnology Bulletin, 2023, 39(11): 150-167. |
| [8] | ZHOU Heng, XIE Yan-jie. Recent Progress in Oxidative Stress Signaling and Response in Plants [J]. Biotechnology Bulletin, 2023, 39(11): 36-43. |
| [9] | LIN Rong, ZHENG Yue-ping, XU Xue-zhen, LI Dan-dan, ZHENG Zhi-fu. Functional Analysis of ACOL8 Gene in the Ethylene Synthesis and Response in Arabidopsis thaliana [J]. Biotechnology Bulletin, 2023, 39(1): 157-165. |
| [10] | LIU Jia-xin, ZHANG Hui-long, ZOU Rong-song, YANG Xiu-yan, ZHU Jian-feng, ZHANG Hua-xin. Research Progress in Na+ Antiport and Physiological Growth Mechanisms of Differernt Halophytes Adapted to Salt Stress [J]. Biotechnology Bulletin, 2023, 39(1): 59-72. |
| [11] | GAO Xiao-rong, DING Yao, LV Jun. Effects of Pseudomonas sp. PR3,a Pyrene-degrading Bacterium with Plant Growth-promoting Properties,on Rice Growth Under Pyrene Stress [J]. Biotechnology Bulletin, 2022, 38(9): 226-236. |
| [12] | CHEN Hong-yan, LI Xiao-er, LI Zhong-guang. Sugar Signaling and Its Role in Plant Response to Environmental Stress [J]. Biotechnology Bulletin, 2022, 38(7): 80-89. |
| [13] | XUE Xian-li, WANG Jing-ran, BI Hang-hang, WANG De-pei. Effect of Spt7 Overexpression of on the Growth and Stress Resistance of Aspergillus niger [J]. Biotechnology Bulletin, 2022, 38(5): 112-122. |
| [14] | ZU Guo-qiang, HU Zhe, WANG Qi, LI Guang-zhe, HAO Lin. Regulatory Role of Burkholderia sp. GD17 in Rice Seedling’s Responses to Cadmium Stress [J]. Biotechnology Bulletin, 2022, 38(4): 153-162. |
| [15] | YIPARE·Paerhati , ZULIHUMAER·Rouzi , TIAN Yong-zhi, ZHU Yan-lei, LI Yuan-ting, MA Xiao-lin. Research Progress in Diversity of Endophytes Microbial Communities Isolated from Desert Plants and Their Strengthening Effects on Drought and Salt Tolerance in Crops [J]. Biotechnology Bulletin, 2022, 38(12): 88-99. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||