








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 172-186.doi: 10.13560/j.cnki.biotech.bull.1985.2025-1277
Previous Articles Next Articles
YAN Chen-lin1( ), LI Fan1, YAN Chun-ting1, CHENG Jiao-wen1, HU Kai-lin1, YE Zhi-biao2, SONG Jian-wen1(
), LI Fan1, YAN Chun-ting1, CHENG Jiao-wen1, HU Kai-lin1, YE Zhi-biao2, SONG Jian-wen1( )
)
Received:2025-11-24
Online:2026-03-26
Published:2026-04-23
Contact:
SONG Jian-wen
E-mail:15170097160@163.com;songjianwen200@scau.edu.cn
YAN Chen-lin, LI Fan, YAN Chun-ting, CHENG Jiao-wen, HU Kai-lin, YE Zhi-biao, SONG Jian-wen. Advances in Genes Related to Tomato Fruit Morphogenesis[J]. Biotechnology Bulletin, 2026, 42(3): 172-186.
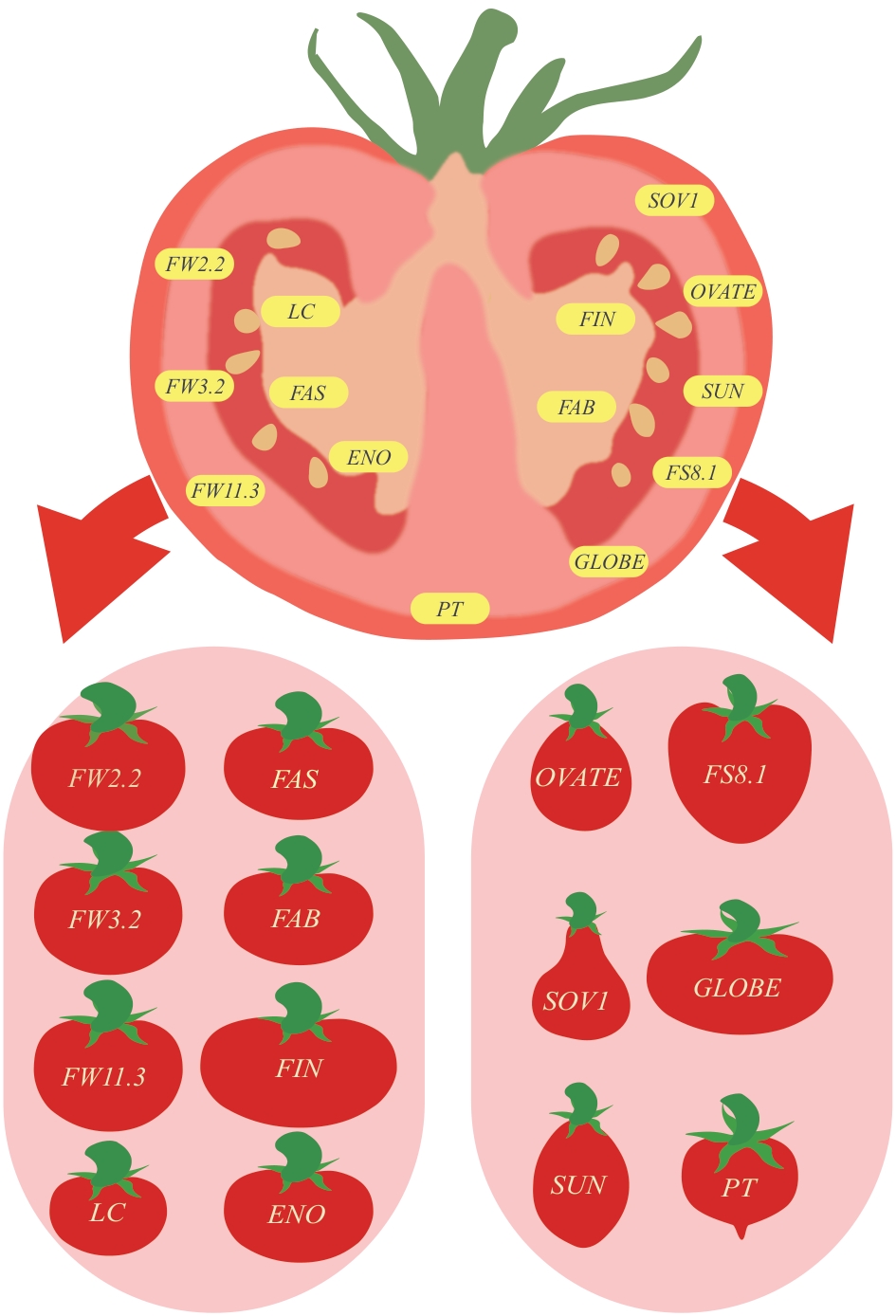
Fig. 2 Genes associated with fruit morphology development in tomatoesLeft: Genes and their phenotypes associated with tomato fruit size. Right: Genes and their phenotypes associated with tomato fruit shape
基因类型 Gene type | 基因编号 Gene ID | 调控机制 Regulatory mechanism | 参考文献 References |
|---|---|---|---|
| FW2.2 | Solyc02g090730 | 负调控胞间连丝处的胼胝质沉积,保证细胞间信号分子的流通 | [ |
| The negative regulation of callose deposition at plasmodesmata ensures the flow of signaling molecules between cells | |||
| FW3.2 | Solyc03g114940 | KLUH基因剂量提高,基因表达量显著提升 | [ |
| An increase in the dosage of the KLUH gene leads to a significant elevation in gene expressions | |||
| FW11.3 | Solyc11g071940 | 可能与细胞分化和维管发育有关,具体调控机制未知 | [ |
| It may be associated with cell differentiation and vascular development, although the precise regulatory mechanism remains unknown | |||
| LC | Solyc02g083950 | 调控WUS基因表达,影响WUS-CLV3途径 | [ |
| Regulation of WUS gene expression to influence the WUS-CLV3 pathway | |||
| FAS | Solyc11g071380 | 调控CLV3基因表达,影响WUS-CLV3途径 | [ |
| Regulation of CLV3 gene expression to influence the WUS-CLV3 pathway | |||
| FAB | Solyc04g081590 | 影响WUS-CLV3途径 | [ |
| Impact on the WUS-CLV3 route | |||
| FIN | Solyc11g064850 | 调控CLV3的糖基化修饰,影响WUS-CLV3途径 | [ |
| Modulation of CLV3 glycosylation modification influences the WUS-CLV3 pathway | |||
| ENO | Solyc03g117230 | 调控WUS基因表达,影响WUS-CLV3途径 | [ |
| Regulation of WUS gene expression influences the WUS-CLV3 pathway | |||
| OVATE | Solyc02g085500 | OVATE和OFP20通过与TRM互作,改变细胞分裂模式 | [ |
| OVATE and OFP20 modify cell division patterns by interacting with TRM | |||
| SOV1 | Solyc03g097060 | SOV1和TRM家族互作,调控细胞分裂模式,最终影响果实伸长 | [ |
| SOV1 interacts with the TRM family to regulate cell division patterns, thereby influencing fruit elongation | |||
| SUN | Solyc10g079240 | SUN通过CAM介导微管重排,影响细胞分裂 | [ |
| SUN influences cell division by intervening in the rearrangement of microtubules via CAM | |||
| GLOBE | Solyc12g006860 | 可能与油菜素内酯有关,具体调控机制未知 | [ |
| This could be related to brassinosteroids, although the precise regulatory mechanism remains unknown | |||
| FS8.1 | Solyc08g061910 | FS8.1编码蛋白与SlGT-16蛋白协同作用,调控子房壁与中轴的细胞增殖速率 | [ |
| The protein encoded by FS8.1 acts synergistically with the SIGT-16 protein to regulate cell proliferation rates in the ovarian wall and central axis | |||
| PT | Solyc05g054030 | 影响C2H2结构域数目,负调控FUL2的表达 | [ |
| Influencing the number of C2H2 domains, negatively regulating the expression of FUL2 |
Table 1 Genes and regulatory mechanisms involved in tomato fruit morphogenesis
基因类型 Gene type | 基因编号 Gene ID | 调控机制 Regulatory mechanism | 参考文献 References |
|---|---|---|---|
| FW2.2 | Solyc02g090730 | 负调控胞间连丝处的胼胝质沉积,保证细胞间信号分子的流通 | [ |
| The negative regulation of callose deposition at plasmodesmata ensures the flow of signaling molecules between cells | |||
| FW3.2 | Solyc03g114940 | KLUH基因剂量提高,基因表达量显著提升 | [ |
| An increase in the dosage of the KLUH gene leads to a significant elevation in gene expressions | |||
| FW11.3 | Solyc11g071940 | 可能与细胞分化和维管发育有关,具体调控机制未知 | [ |
| It may be associated with cell differentiation and vascular development, although the precise regulatory mechanism remains unknown | |||
| LC | Solyc02g083950 | 调控WUS基因表达,影响WUS-CLV3途径 | [ |
| Regulation of WUS gene expression to influence the WUS-CLV3 pathway | |||
| FAS | Solyc11g071380 | 调控CLV3基因表达,影响WUS-CLV3途径 | [ |
| Regulation of CLV3 gene expression to influence the WUS-CLV3 pathway | |||
| FAB | Solyc04g081590 | 影响WUS-CLV3途径 | [ |
| Impact on the WUS-CLV3 route | |||
| FIN | Solyc11g064850 | 调控CLV3的糖基化修饰,影响WUS-CLV3途径 | [ |
| Modulation of CLV3 glycosylation modification influences the WUS-CLV3 pathway | |||
| ENO | Solyc03g117230 | 调控WUS基因表达,影响WUS-CLV3途径 | [ |
| Regulation of WUS gene expression influences the WUS-CLV3 pathway | |||
| OVATE | Solyc02g085500 | OVATE和OFP20通过与TRM互作,改变细胞分裂模式 | [ |
| OVATE and OFP20 modify cell division patterns by interacting with TRM | |||
| SOV1 | Solyc03g097060 | SOV1和TRM家族互作,调控细胞分裂模式,最终影响果实伸长 | [ |
| SOV1 interacts with the TRM family to regulate cell division patterns, thereby influencing fruit elongation | |||
| SUN | Solyc10g079240 | SUN通过CAM介导微管重排,影响细胞分裂 | [ |
| SUN influences cell division by intervening in the rearrangement of microtubules via CAM | |||
| GLOBE | Solyc12g006860 | 可能与油菜素内酯有关,具体调控机制未知 | [ |
| This could be related to brassinosteroids, although the precise regulatory mechanism remains unknown | |||
| FS8.1 | Solyc08g061910 | FS8.1编码蛋白与SlGT-16蛋白协同作用,调控子房壁与中轴的细胞增殖速率 | [ |
| The protein encoded by FS8.1 acts synergistically with the SIGT-16 protein to regulate cell proliferation rates in the ovarian wall and central axis | |||
| PT | Solyc05g054030 | 影响C2H2结构域数目,负调控FUL2的表达 | [ |
| Influencing the number of C2H2 domains, negatively regulating the expression of FUL2 |
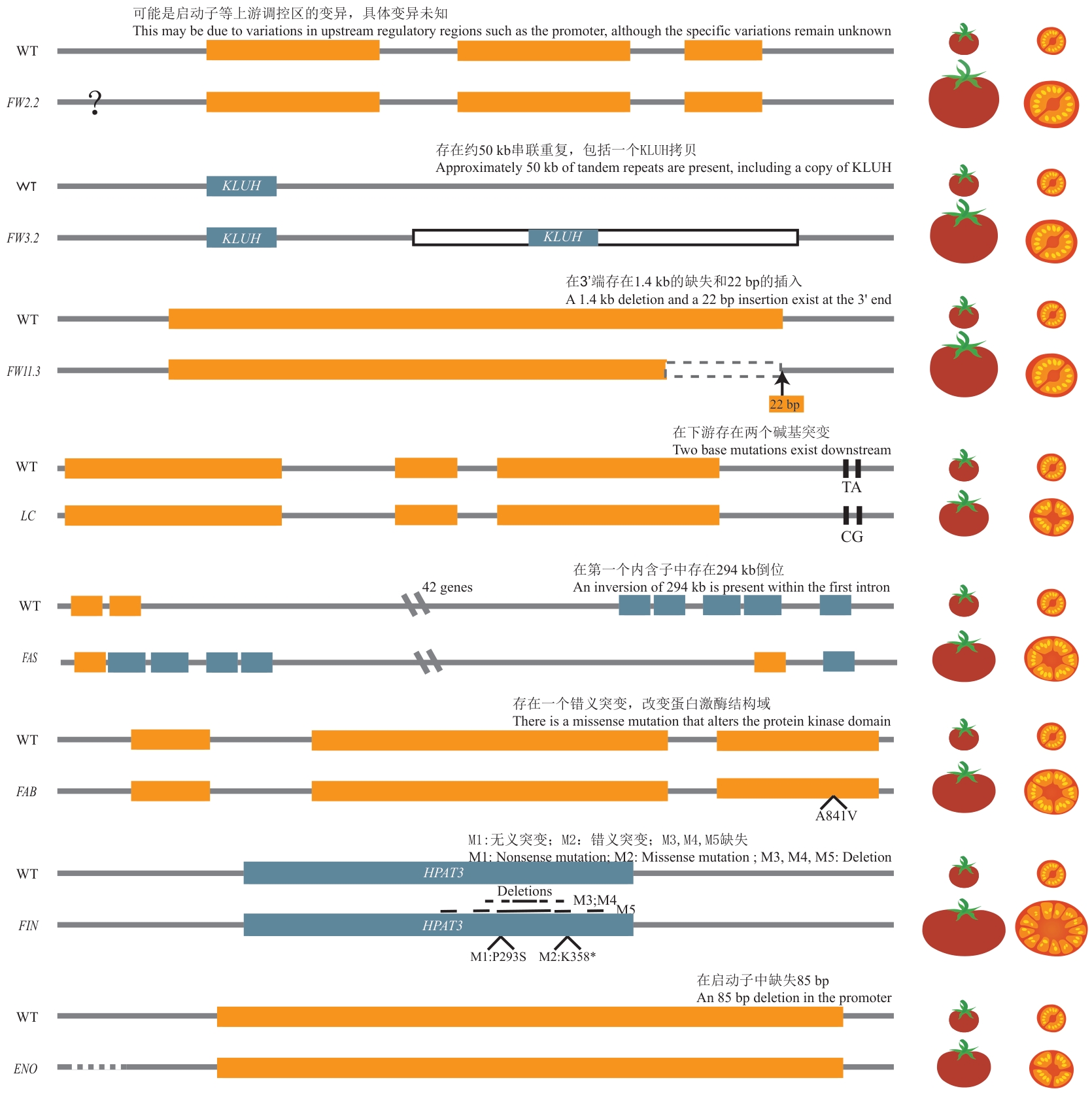
Fig. 3 Genetic structure of governing tomato fruit size︿Orange box: Exon; blue box: gene; white box: inserted nucleotide sequence; black box: single base site; grey box: early termination; : genetic mutation/amino acid; M: different allelic mutation; dotted line: promoter deletion︿
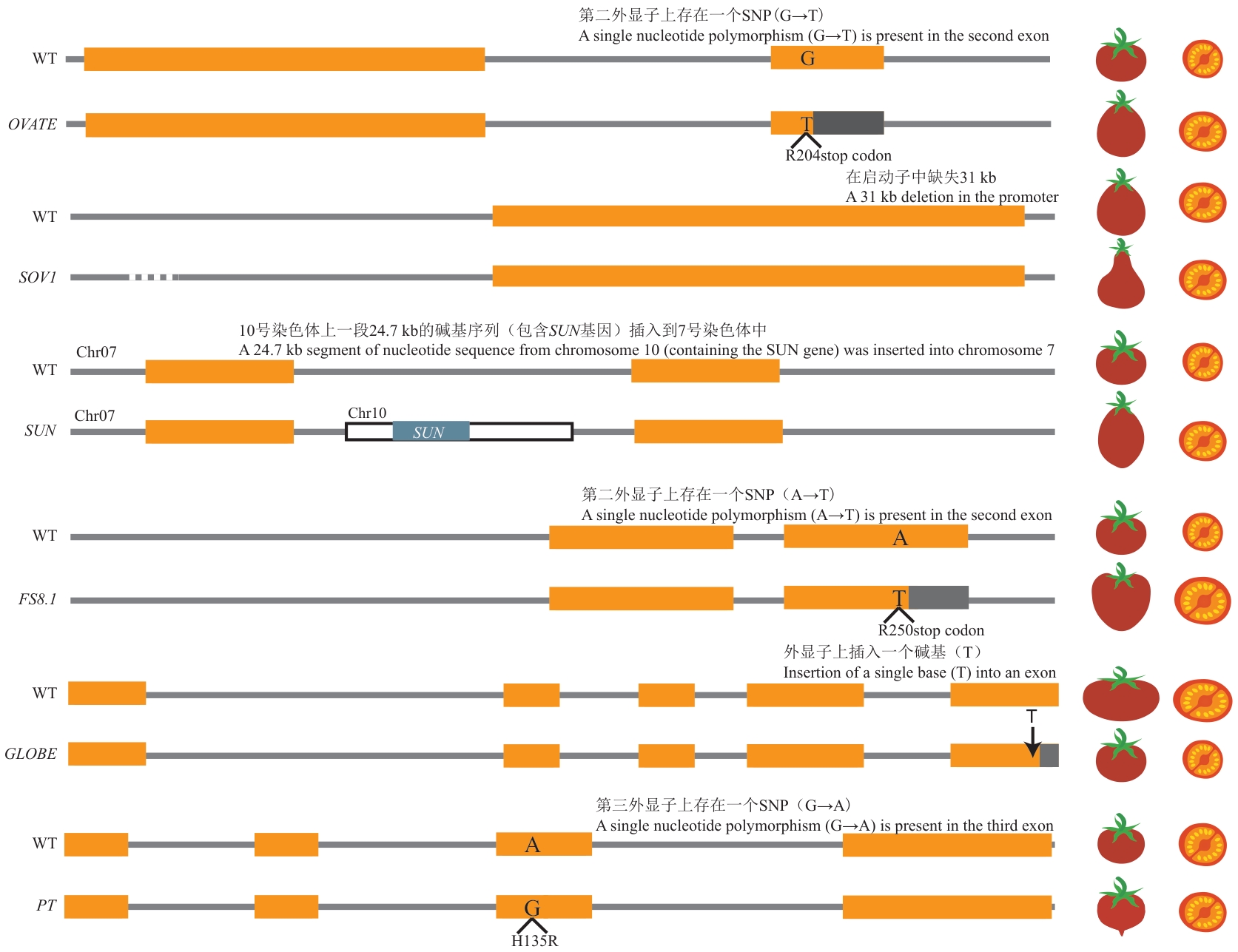
Fig. 4 Genetic structure governing tomato fruit shape︿Orange box: Exon; blue box: gene; white box: inserted nucleotide sequence; grey box: premature termination; : genetic mutation/amino acid; dashed line: promoter deletion; ↓: nucleotide insertion︿
| [1] | Blanca J, Montero-Pau J, Sauvage C, et al. Genomic variation in tomato, from wild ancestors to contemporary breeding accessions [J]. BMC Genomics, 2015, 16(1): 257. |
| [2] | Lin T, Zhu GT, Zhang JH, et al. Genomic analyses provide insights into the history of tomato breeding [J]. Nat Genet, 2014, 46(11): 1220-1226. |
| [3] | Causse M, Friguet C, Coiret C, et al. Consumer preferences for fresh tomato at the European scale: a common segmentation on taste and firmness [J]. J Food Sci, 2010, 75(9): S531-S541. |
| [4] | Simonne AH, Behe BK, Marshall MM. Consumers prefer low-priced and highlycopene-content fresh-market tomatoes [J]. HortTechnology, 2006, 16(4): 674-681. |
| [5] | Muños S, Ranc N, Botton E, et al. Increase in tomato locule number is controlled by two single-nucleotide polymorphisms located near WUSCHEL [J]. Plant Physiol, 2011, 156(4): 2244-2254. |
| [6] | Rodríguez GR, Muños S, Anderson C, et al. Distribution of SUN, OVATE, LC, and FAS in the tomato germplasm and the relationship to fruit shape diversity [J]. Plant Physiol, 2011, 156(1): 275-285. |
| [7] | Meyer RS, Purugganan MD. Evolution of crop species: genetics of domestication and diversification [J]. Nat Rev Genet, 2013, 14(12): 840-852. |
| [8] | Olsen KM, Wendel JF. A bountiful harvest: genomic insights into crop domestication phenotypes [J]. Annu Rev Plant Biol, 2013, 64: 47-70. |
| [9] | Doebley J, Stec A, Wendel J, et al. Genetic and morphological analysis of a maize-teosinte F2 population: implications for the origin of maize [J]. Proc Natl Acad Sci USA, 1990, 87(24): 9888-9892. |
| [10] | Paran I, van der Knaap E. Genetic and molecular regulation of fruit and plant domestication traits in tomato and pepper [J]. J Exp Bot, 2007, 58(14): 3841-3852. |
| [11] | Welty N, Radovich C, Meulia T, et al. Inflorescence development in two tomato species [J]. Can J Bot, 2007, 85(1): 111-118. |
| [12] | Xiao H, Radovich C, Welty N, et al. Integration of tomato reproductive developmental landmarks and expression profiles, and the effect of SUN on fruit shape [J]. BMC Plant Biol, 2009, 9: 49. |
| [13] | Du MM, Sun CL, Deng L, et al. Molecular breeding of tomato: Advances and challenges [J]. J Integr Plant Biol, 2025, 67(3): 669-721. |
| [14] | Chen KL, Wang YP, Zhang R, et al. CRISPR/cas genome editing and precision plant breeding in agriculture [J]. Annu Rev Plant Biol, 2019, 70: 667-697. |
| [15] | Rodríguez-Leal D, Lemmon ZH, Man J, et al. Engineering quantitative trait variation for crop improvement by genome editing [J]. Cell, 2017, 171(2): 470-480.e8. |
| [16] | Wallace JG, Rodgers-Melnick E, Buckler ES. On the road to breeding 4.0: unraveling the good, the bad, and the boring of crop quantitative genomics [J]. Annu Rev Genet, 2018, 52: 421-444. |
| [17] | Gillaspy G, Ben-David H, Gruissem W. Fruits: A developmental perspective [J]. Plant Cell, 1993, 5(10): 1439-1451. |
| [18] | Bohner J, Bangerth F. Effects of fruit set sequence and defoliation on cell number, cell size and hormone levels of tomato fruits (Lycopersicon esculentum Mill.) within a truss [J]. Plant Growth Regul, 1988, 7(3): 141-155. |
| [19] | Frary A, Nesbitt TC, Grandillo S, et al. fw2.2: a quantitative trait locus key to the evolution of tomato fruit size [J]. Science, 2000, 289(5476): 85-88. |
| [20] | Guo M, Rupe MA, Dieter JA, et al. Cell Number Regulator1 affects plant and organ size in maize: implications for crop yield enhancement and heterosis [J]. Plant Cell, 2010, 22(4): 1057-1073. |
| [21] | Cong B, Liu JP, Tanksley SD. Natural alleles at a tomato fruit size quantitative trait locus differ by heterochronic regulatory mutations [J]. Proc Natl Acad Sci USA, 2002, 99(21): 13606-13611. |
| [22] | Liu JP, Cong B, Tanksley SD. Generation and analysis of an artificial gene dosage series in tomato to study the mechanisms by which the cloned quantitative trait locus fw2.2 controls fruit size [J]. Plant Physiol, 2003, 132(1): 292-299. |
| [23] | Beauchet A, Bollier N, Grison M, et al. The CELL NUMBER REGULATOR FW2.2 protein regulates cell-to-cell communication in tomato by modulating callose deposition at plasmodesmata [J]. Plant Physiol, 2024, 196(2): 883-901. |
| [24] | Cong B, Tanksley SD. FW2.2 and cell cycle control in developing tomato fruit: a possible example of gene co-option in the evolution of a novel organ [J]. Plant Mol Biol, 2006, 62(6): 867-880. |
| [25] | Chakrabarti M, Zhang N, Sauvage C, et al. A cytochrome P450 regulates a domestication trait in cultivated tomato [J]. Proc Natl Acad Sci USA, 2013, 110(42): 17125-17130. |
| [26] | Alonge M, Wang XG, Benoit M, et al. Major impacts of widespread structural variation on gene expression and crop improvement in tomato [J]. Cell, 2020, 182(1): 145-161.e23. |
| [27] | Huang ZJ, van der Knaap E. Tomato fruit weight 11.3 maps close to fasciated on the bottom of chromosome 11 [J]. Theor Appl Genet, 2011, 123(3): 465-474. |
| [28] | Mu Q, Huang ZJ, Chakrabarti M, et al. Fruit weight is controlled by cell size regulator encoding a novel protein that is expressed in maturing tomato fruits [J]. PLoS Genet, 2017, 13(8): e1006930. |
| [29] | Liu XG, Kim YJ, Müller R, et al. AGAMOUS terminates floral stem cell maintenance in Arabidopsis by directly repressing WUSCHEL through recruitment of polycomb group proteins [J]. Plant Cell, 2011, 23(10): 3654-3670. |
| [30] | Chu YH, Jang JC, Huang ZJ, et al. Tomato locule number and fruit size controlled by natural alleles of lc and fas [J]. Plant Direct, 2019, 3(7): e00142. |
| [31] | Xu C, Liberatore KL, MacAlister CA, et al. A cascade of Arabinosyltransferases controls shoot meristem size in tomato [J]. Nat Genet, 2015, 47(7): 784-792. |
| [32] | Yuste-Lisbona FJ, Fernández-Lozano A, Pineda B, et al. ENO regulates tomato fruit size through the floral meristem development network [J]. Proc Natl Acad Sci USA, 2020, 117(14): 8187-8195. |
| [33] | Liu JP, Van Eck J, Cong B, et al. A new class of regulatory genes underlying the cause of pear-shaped tomato fruit [J]. Proc Natl Acad Sci USA, 2002, 99(20): 13302-13306. |
| [34] | Van Der Knaap E, Lippman ZB, Tanksley SD. Extremely elongated tomato fruit controlled by four quantitative trait loci with epistatic interactions [J]. Theor Appl Genet, 2002, 104(2/3): 241-247. |
| [35] | Wu S, Zhang BY, Keyhaninejad N, et al. A common genetic mechanism underlies morphological diversity in fruits and other plant organs [J]. Nat Commun, 2018, 9(1): 4734. |
| [36] | Xiao H, Jiang N, Schaffner E, et al. A retrotransposon-mediated gene duplication underlies morphological variation of tomato fruit [J]. Science, 2008, 319(5869): 1527-1530. |
| [37] | Bürstenbinder K, Möller B, Plötner R, et al. The IQD family of calmodulin-binding proteins links calcium signaling to microtubules, membrane subdomains, and the nucleus [J]. Plant Physiol, 2017, 173(3): 1692-1708. |
| [38] | Sierra-Orozco E, Shekasteband R, Illa-Berenguer E, et al. Identification and characterization of GLOBE a major gene controlling fruit shape and impacting fruit size and marketability in tomato [J]. Hortic Res, 2021, 8(1): 138. |
| [39] | Zhu Q, Deng L, Chen J, et al. Redesigning the tomato fruit shape for mechanized production [J]. Nat Plants, 2023, 9(10): 1659-1674. |
| [40] | Song JW, Shang LL, Li CX, et al. Variation in the fruit development gene POINTED TIP regulates protuberance of tomato fruit tip [J]. Nat Commun, 2022, 13(1): 5940. |
| [41] | Anastasiou E, Kenz S, Gerstung M, et al. Control of plant organ size by KLUH/CYP78A5-dependent intercellular signaling [J]. Dev Cell, 2007, 13(6): 843-856. |
| [42] | Lye ZN, Purugganan MD. Copy number variation in domestication [J]. Trends Plant Sci, 2019, 24(4): 352-365. |
| [43] | Shang LL, Tao JB, Song JW, et al. CRISPR/Cas9-mediated mutations of FANTASTIC FOUR gene family for creating early flowering mutants in tomato [J]. Plant Biotechnol J, 2024, 22(3): 774-784. |
| [44] | Barrero LS, Cong B, Wu F, et al. Developmental characterization of the fasciated locus and mapping of Arabidopsis candidate genes involved in the control of floral meristem size and carpel number in tomato [J]. Genome, 2006, 49(8): 991-1006. |
| [45] | Tanksley SD. The genetic, developmental, and molecular bases of fruit size and shape variation in tomato [J]. Plant Cell, 2004, 16(): S181-S189. |
| [46] | Fletcher JC, Brand U, Running MP, et al. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems [J]. Science, 1999, 283(5409): 1911-1914. |
| [47] | Laux T, Mayer KF, Berger J, et al. The WUSCHEL gene is required for shoot and floral meristem integrity in Arabidopsis [J]. Development, 1996, 122(1): 87-96. |
| [48] | Mayer KFX, Schoof H, Haecker A, et al. Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem [J]. Cell, 1998, 95(6): 805-815. |
| [49] | Brand U, Fletcher JC, Hobe M, et al. Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity [J]. Science, 2000, 289(5479): 617-619. |
| [50] | Schoof H, Lenhard M, Haecker A, et al. The stem cell population of Arabidopsis shoot meristems is maintained by a regulatory loop between the CLAVATA and WUSCHEL genes [J]. Cell, 2000, 100(6): 635-644. |
| [51] | Perales M, Rodriguez K, Snipes S, et al. Threshold-dependent transcriptional discrimination underlies stem cell homeostasis [J]. Proc Natl Acad Sci USA, 2016, 113(41): E6298-E6306. |
| [52] | Yadav RK, Perales M, Gruel J, et al. WUSCHEL protein movement mediates stem cell homeostasis in the Arabidopsis shoot apex [J]. Genes Dev, 2011, 25(19): 2025-2030. |
| [53] | Shinohara H, Matsubayashi Y. Reevaluation of the CLV3-receptor interaction in the shoot apical meristem: dissection of the CLV3 signaling pathway from a direct ligand-binding point of view [J]. Plant J, 2015, 82(2): 328-336. |
| [54] | Somssich M, Je BI, Simon R, et al. CLAVATA-WUSCHEL signaling in the shoot meristem [J]. Development, 2016, 143(18): 3238-3248. |
| [55] | Song SK, Lee MM, Clark SE. POL and PLL1 phosphatases are CLAVATA1 signaling intermediates required for Arabidopsis shoot and floral stem cells [J]. Development, 2006, 133(23): 4691-4698. |
| [56] | Betsuyaku S, Takahashi F, Kinoshita A, et al. Mitogen-activated protein kinase regulated by the CLAVATA receptors contributes to shoot apical meristem homeostasis [J]. Plant Cell Physiol, 2011, 52(1): 14-29. |
| [57] | Lippman Z, Tanksley SD. Dissecting the genetic pathway to extreme fruit size in tomato using a cross between the small-fruited wild species Lycopersicon pimpinellifolium and L. esculentum var. Giant Heirloom [J]. Genetics, 2001, 158(1): 413-422. |
| [58] | van der Knaap E, Tanksley SD. The making of a bell pepper-shaped tomato fruit: identification of loci controlling fruit morphology in Yellow Stuffer tomato [J]. Theor Appl Genet, 2003, 107(1): 139-147. |
| [59] | Lenhard M, Bohnert A, Jürgens G, et al. Termination of stem cell maintenance in Arabidopsis floral meristems by interactions between WUSCHEL and AGAMOUS [J]. Cell, 2001, 105(6): 805-814. |
| [60] | Lohmann JU, Hong RL, Hobe M, et al. A molecular link between stem cell regulation and floral patterning in Arabidopsis [J]. Cell, 2001, 105(6): 793-803. |
| [61] | Yanofsky MF, Ma H, Bowman JL, et al. The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors [J]. Nature, 1990, 346(6279): 35-39. |
| [62] | Cong B, Barrero LS, Tanksley SD. Regulatory change in YABBY-like transcription factor led to evolution of extreme fruit size during tomato domestication [J]. Nat Genet, 2008, 40(6): 800-804. |
| [63] | Barrero LS, Tanksley SD. Evaluating the genetic basis of multiple-locule fruit in a broad cross section of tomato cultivars [J]. Theor Appl Genet, 2004, 109(3): 669-679. |
| [64] | Fernández-Lozano A, Yuste-Lisbona FJ, Pérez-Martín F, et al. Mutation at the tomato EXCESSIVE NUMBER OF FLORAL ORGANS (ENO) locus impairs floral meristem development, thus promoting an increased number of floral organs and fruit size [J]. Plant Sci, 2015, 232: 41-48. |
| [65] | Franco-Zorrilla JM, López-Vidriero I, Carrasco JL, et al. DNA-binding specificities of plant transcription factors and their potential to define target genes [J]. Proc Natl Acad Sci USA, 2014, 111(6): 2367-2372. |
| [66] | Aguirre L, Hendelman A, Hutton SF, et al. Idiosyncratic and dose-dependent epistasis drives variation in tomato fruit size [J]. Science, 2023, 382(6668): 315-320. |
| [67] | van der Knaap E, Chakrabarti M, Chu YH, et al. What lies beyond the eye: the molecular mechanisms regulating tomato fruit weight and shape [J]. Front Plant Sci, 2014, 5: 227. |
| [68] | Hackbusch J, Richter K, Müller J, et al. A central role of Arabidopsis thaliana ovate family proteins in networking and subcellular localization of 3-aa loop extension homeodomain proteins [J]. Proc Natl Acad Sci USA, 2005, 102(13): 4908-4912. |
| [69] | Wang SC, Chang Y, Guo JJ, et al. Arabidopsis Ovate Family Protein 1 is a transcriptional repressor that suppresses cell elongation [J]. Plant J, 2007, 50(5): 858-872. |
| [70] | Tsaballa A, Pasentsis K, Darzentas N, et al. Multiple evidence for the role of an Ovate-like gene in determining fruit shape in pepper [J]. BMC Plant Biol, 2011, 11: 46. |
| [71] | van der Knaap E, Tanksley SD. Identification and characterization of a novel locus controlling early fruit development in tomato [J]. Theor Appl Genet, 2001, 103(2): 353-358. |
| [72] | van der Knaap E, Sanyal A, Jackson SA, et al. High-resolution fine mapping and fluorescence in situ hybridization analysis of sun, a locus controlling tomato fruit shape, reveals a region of the tomato genome prone to DNA rearrangements [J]. Genetics, 2004, 168(4): 2127-2140. |
| [73] | Levy M, Wang QM, Kaspi R, et al. Arabidopsis IQD1, a novel calmodulin-binding nuclear protein, stimulates glucosinolate accumulation and plant defense [J]. Plant J, 2005, 43(1): 79-96. |
| [74] | Zhang C, Fan XC, Liu CH, et al. Anatomical berry characteristics during the development of grape berries with different shapes [J]. Hortic Plant J, 2021, 7(4): 295-306. |
| [75] | Huang ZJ, Van Houten J, Gonzalez G, et al. Genome-wide identification, phylogeny and expression analysis of SUN, OFP and YABBY gene family in tomato [J]. Mol Genet Genomics, 2013, 288(3/4): 111-129. |
| [76] | Bao ZR, Xu ZJ, Zang JZ, et al. The morphological diversity of plant organs: manipulating the organization of microtubules may do the trick [J]. Front Cell Dev Biol, 2021, 9: 649626. |
| [77] | Bao ZR, Guo Y, Deng YL, et al. Microtubule-associated protein SlMAP70 interacts with IQ67-domain protein SlIQD21a to regulate fruit shape in tomato [J]. Plant Cell, 2023, 35(12): 4266-4283. |
| [78] | Bishop GJ, Yokota T. Plants steroid hormones, brassinosteroids: current highlights of molecular aspects on their synthesis/metabolism, transport, perception and response [J]. Plant Cell Physiol, 2001, 42(2): 114-120. |
| [79] | Ohnishi T, Nomura T, Watanabe B, et al. Tomato cytochrome P450 CYP734A7 functions in brassinosteroid catabolism [J]. Phytochemistry, 2006, 67(17): 1895-1906. |
| [80] | Shinozaki Y, Nicolas P, Fernandez-Pozo N, et al. High-resolution spatiotemporal transcriptome mapping of tomato fruit development and ripening [J]. Nat Commun, 2018, 9(1): 364. |
| [81] | Grandillo S, Tanksley SD. QTL analysis of horticultural traits differentiating the cultivated tomato from the closely related species Lycopersicon pimpinellifolium [J]. Theor Appl Genet, 1996, 92(8): 935-951. |
| [82] | Sun L, Rodriguez GR, Clevenger JP, et al. Candidate gene selection and detailed morphological evaluations of fs8.1, a quantitative trait locus controlling tomato fruit shape [J]. J Exp Bot, 2015, 66(20): 6471-6482. |
| [83] | Frary A, Fulton TM, Zamir D, et al. Advanced backcross QTL analysis of a Lycopersicon esculentum × L. pennellii cross and identification of possible orthologs in the Solanaceae [J]. Theor Appl Genet, 2004, 108(3): 485-496. |
| [84] | Ku HM, Grandillo S, Tanksley SD. fs8.1, a major QTL, sets the pattern of tomato carpel shape well before anthesis [J]. Theor Appl Genet, 2000, 101(5): 873-878. |
| [85] | Kaplan-Levy RN, Brewer PB, Quon T, et al. The trihelix family of transcription factors-light, stress and development [J]. Trends Plant Sci, 2012, 17(3): 163-171. |
| [86] | Zhou DX. Regulatory mechanism of plant gene transcription by GT-elements and GT-factors [J]. Trends Plant Sci, 1999, 4(6): 210-214. |
| [87] | Barten JHM, Scott JW, Gardner RG. Characterization of blossom-end morphology genes in tomato and their usefulness in breeding for smooth blossom-end scars [J]. J Amer Soc Hort Sci, 1994, 119(4): 798-803. |
| [88] | Langguth P, Mutschler E. Lipophilisation of hydrophilic compounds. Consequences on transepidermal and intestinal transport of trospium chloride [J]. Arzneimittelforschung, 1987, 37(12): 1362-1366. |
| [89] | Rick CM. Abortion of male and female gametes in the tomato determined by allelic interaction [J]. Genetics, 1966, 53(1): 85-96. |
| [90] | Rick CM, Butler L. Cytogenetics of the tomato [M]//Advances in Genetics. Amsterdam: Elsevier, 1956: 267-382. |
| [91] | de Jong M, Wolters-Arts M, García-Martínez JL, et al. The Solanum lycopersicum AUXIN RESPONSE FACTOR 7 (SlARF7) mediates cross-talk between auxin and gibberellin signalling during tomato fruit set and development [J]. J Exp Bot, 2011, 62(2): 617-626. |
| [92] | Mariotti L, Picciarelli P, Lombardi L, et al. Fruit-set and early fruit growth in tomato are associated with increases in indoleacetic acid, cytokinin, and bioactive gibberellin contents [J]. J Plant Growth Regul, 2011, 30(4): 405-415. |
| [93] | Su LY, Bassa C, Audran C, et al. The auxin Sl-IAA17 transcriptional repressor controls fruit size via the regulation of endoreduplication-related cell expansion [J]. Plant Cell Physiol, 2014, 55(11): 1969-1976. |
| [94] | Wang YP, Clevenger JP, Illa-Berenguer E, et al. A comparison of sun, ovate, fs8.1 and auxin application on tomato fruit shape and gene expression [J]. Plant Cell Physiol, 2019, 60(5): 1067-1081. |
| [95] | 艾国. BR信号调控番茄果实形状机制研究 [D]. 武汉: 华中农业大学, 2020. |
| Ai G. Functional dissection of BR signal transduction in regulating tomato fruit shape [D]. Wuhan: Huazhong Agricultural University, 2020. | |
| [96] | Zouine M, Fu YY, Chateigner-Boutin AL, et al. Characterization of the tomato ARF gene family uncovers a multi-levels post-transcriptional regulation including alternative splicing [J]. PLoS One, 2014, 9(1): e84203. |
| [97] | de Jong M, Wolters-Arts M, Schimmel BCJ, et al. Solanum lycopersicum AUXIN RESPONSE FACTOR 9 regulates cell division activity during early tomato fruit development [J]. J Exp Bot, 2015, 66(11): 3405-3416. |
| [98] | Liu SY, Zhang YW, Feng QS, et al. Tomato AUXIN RESPONSE FACTOR 5 regulates fruit set and development via the mediation of auxin and gibberellin signaling [J]. Sci Rep, 2018, 8(1): 2971. |
| [99] | Su LY, Audran C, Bouzayen M, et al. The Aux/IAA, Sl-IAA17 regulates quality parameters over tomato fruit development [J]. Plant Signal Behav, 2015, 10(11): e1071001. |
| [100] | Yu T, Ai G, Xie QM, et al. Regulation of tomato fruit elongation by transcription factor BZR1.7 through promotion of SUN gene expression [J]. Hortic Res, 2022, 9: uhac121. |
| [101] | Grandillo S, Ku HM, Tanksley SD. Identifying the loci responsible for natural variation in fruit size and shape in tomato [J]. Theor Appl Genet, 1999, 99(6): 978-987. |
| [102] | Li TD, Yang XP, Yu Y, et al. Domestication of wild tomato is accelerated by genome editing [J]. Nat Biotechnol, 2018, 36(12): 1160-1163. |
| [103] | Hickey LT, Hafeez A N, Robinson H, et al. Breeding crops to feed 10 billion [J]. Nat Biotechnol, 2019, 37(7): 744-754. |
| [104] | Purugganan MD. Evolutionary insights into the nature of plant domestication [J]. Curr Biol, 2019, 29(14): R705-R714. |
| [105] | Purugganan MD. What is domestication [J]. Trends Ecol Evol, 2022, 37(8): 663-671. |
| [106] | Benke K, Tomkins B. Future food-production systems: vertical farming and controlled-environment agriculture [J]. Sustain Sci Pract Policy, 2017, 13(1): 13-26. |
| [107] | Pearson LJ, Pearson L, Pearson CJ. Sustainable urban agriculture: stocktake and opportunities [J]. Int J Agric Sustain, 2010, 8(1/2): 7-19. |
| [108] | Kwon CT, Heo J, Lemmon ZH, et al. Rapid customization of Solanaceae fruit crops for urban agriculture [J]. Nat Biotechnol, 2020, 38(2): 182-188. |
| [109] | Bouidghaghen J, Moreau L, Beauchêne K, et al. Robotized indoor phenotyping allows genomic prediction of adaptive traits in the field [J]. Nat Commun, 2023, 14(1): 6603. |
| [110] | Jafar A, Bibi N, Ali Naqvi R, et al. Revolutionizing agriculture with artificial intelligence: plant disease detection methods, applications, and their limitations [J]. Front Plant Sci, 2024, 15: 1356260. |
| [111] | Jung J, Maeda M, Chang AJ, et al. The potential of remote sensing and artificial intelligence as tools to improve the resilience of agriculture production systems [J]. Curr Opin Biotechnol, 2021, 70: 15-22. |
| [112] | Pennisi E. Sowing the seeds for the ideal crop [J]. Science, 2010, 327(5967): 802-803. |
| [113] | Zhang P, Guo ZL, Ullah S, et al. Nanotechnology and artificial intelligence to enable sustainable and precision agriculture [J]. Nat Plants, 2021, 7(7): 864-876. |
| [114] | Xie Y, Zhang TH, Yang MH, et al. Engineering crop flower morphology facilitates robotization of cross-pollination and speed breeding [J]. Cell, 2025, 188(21): 5809-5830.e27. |
| [1] | LIU Miao, LIN Tao, JIA Le-song, HU Feng, LI Tao, LI Zhi-wan, LIU Mei-fang, ZHENG Fang-yan, CUI Long. From Wild to Cultivated: Evolution and Regulatory Mechanisms of Tomato Fruit Color [J]. Biotechnology Bulletin, 2026, 42(3): 187-202. |
| [2] | LI Ya-ni, HAN Hong-yu, GENG Meng-shuang, MI Ruo-lan, Wang Wei-qi, Yu Wen-jing, MENG Xian-wen, LI Chuan-you. Mechanistic Study on ChiC-mediated Regulation Mechanism of Tomato Resistance to Botrytis cinerea [J]. Biotechnology Bulletin, 2026, 42(3): 255-262. |
| [3] | CHENG Yun-xia, ZHANG Jun-hong, YE Jie. Advances in the Genetic Regulation of Soluble Solid Accumulation in Tomato Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 145-155. |
| [4] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [5] | DU Dan, GUO Xiang, HU Xin, PAN Yu. Advances in the Regulatory Mechanisms of Plastid Development on Fruit Ripening and Quality [J]. Biotechnology Bulletin, 2026, 42(3): 48-59. |
| [6] | JIANG Zhe-hui, WANG Xiao-long, WANG Shou-chuang, ZHOU Ke. Advances in the Elucidation of Metabolic Pathways and Molecular Breeding for Tomato Flavor [J]. Biotechnology Bulletin, 2026, 42(3): 60-78. |
| [7] | WANG Xiao-yi, LI Jin-yan, XING Xing, ZHU Hong-liang. Screening and Functional Analysis of Ethylene-responsive Genes Regulating Tomato Fruit Ripening and Respiration [J]. Biotechnology Bulletin, 2026, 42(3): 275-282. |
| [8] | LI Ying-hui, WANG Yang-bo-han, ZHOU Hao-bo, LU Xin-ru, ZHANG Ke-xin, YU Yang, LI Chuan-you, SUN Chuan-long. Identification of VPE Gene Family and Their Functional Analysis under Abiotic Stress in Tomato [J]. Biotechnology Bulletin, 2026, 42(3): 263-274. |
| [9] | SU Xiu-min, HAN Wen-qing, WANG Jiao, LI Peng, WANG Qiu-lan, LI Wan-xing, CAO Jin-jun. Isolation, Identification, Biological Characteristics and Biocontrol Effects of Trichoderma harzianum M408 against Tomato Early Blight [J]. Biotechnology Bulletin, 2025, 41(9): 277-288. |
| [10] | HOU Ya-tao, LI Ying-hui, DENG Lei, LI Chang-bao, LI Chuan-you, SUN Chuan-long. Development of Functional Molecular Markers for Tomato Fruit Weight Gene and Population Genotyping Analysis [J]. Biotechnology Bulletin, 2025, 41(4): 98-105. |
| [11] | LIU Jie, WANG Fei, TAO Ting, ZHANG Yu-jing, CHEN Hao-ting, ZHANG Rui-xing, SHI Yu, ZHANG Yi. Overexpression of SlWRKY41 Improves the Tolerance of Tomato Seedlings to Drought [J]. Biotechnology Bulletin, 2025, 41(2): 107-118. |
| [12] | CHEN Hui-ting, XIA Yi-nan, YAN Wei, ZHANG Meng-fei, CHEN Zhou-mei, HOU Li-xia, LIU Xin. Effect of Exogenous Chlorogenic Acid Treatment on Fruit Quality of Facility Tomato [J]. Biotechnology Bulletin, 2025, 41(2): 119-126. |
| [13] | ZHANG Yu-qing, DONG Li-xue, ZHANG Bao-yue, ZHANG Ying, LIU Xue-ao, XIONG Shuang-xi, ZHANG Hong-xia. Characterization of the Activation Domain of SlMYB80 in Tomato and Its Function Validation during Pollen Development in Arabidopsis [J]. Biotechnology Bulletin, 2025, 41(10): 222-232. |
| [14] | LIU Wen-zhi, HE Dan, LI Peng, FU Ying-lin, ZHANG Yi-xin, WEN Hua-jie, YU Wen-qing. Paenibacillus polymyxa New Strain X-11 and Its Growth-promoting Effects on Tomato and Rice [J]. Biotechnology Bulletin, 2024, 40(9): 249-259. |
| [15] | ZHAO Yao, WEN Lang, LUO Shao-dan, LI Zi-xing, LIU Chao-chao. Identification of HMA Gene Family and Cadmium Transport Function of SlHMA1 in Tomato [J]. Biotechnology Bulletin, 2024, 40(2): 212-222. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||