








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (4): 227-238.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0740
Previous Articles Next Articles
NI Fei-fei1,2( ), CHEN Yi-cun1,2, GAO Ming1,2, ZHANG Sheng-jiao3, PENG Fang-you3, CHEN Tao-mei3, ZHAO Yun-xiao1,2(
), CHEN Yi-cun1,2, GAO Ming1,2, ZHANG Sheng-jiao3, PENG Fang-you3, CHEN Tao-mei3, ZHAO Yun-xiao1,2( ), WANG Yang-dong1,2(
), WANG Yang-dong1,2( )
)
Received:2025-07-09
Online:2026-04-26
Published:2026-04-30
Contact:
ZHAO Yun-xiao, WANG Yang-dong
E-mail:nifeifei2042634075@163.com;zyx_yunxiao@caf.ac.cn;wangyangdong@caf.ac.cn
NI Fei-fei, CHEN Yi-cun, GAO Ming, ZHANG Sheng-jiao, PENG Fang-you, CHEN Tao-mei, ZHAO Yun-xiao, WANG Yang-dong. Identification of Litsea cubebaTCP Genes and Their Roles in the Regulation of Terpenoid Biosynthesis[J]. Biotechnology Bulletin, 2026, 42(4): 227-238.
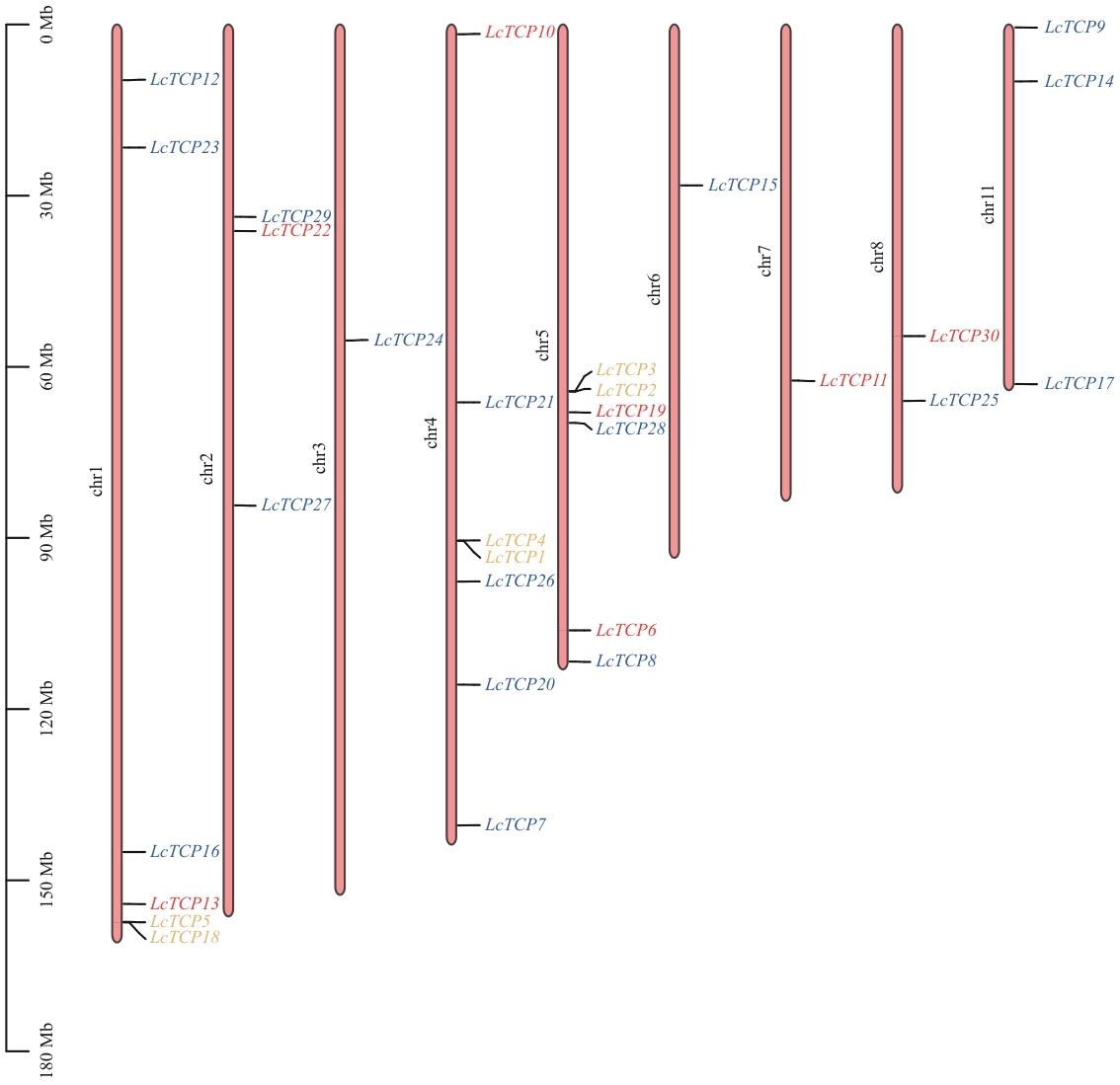
Fig. 1 Chromosomal localization of LcTCP genesBlue indicates the PCF subfamily, yellow indicates the CYC/TB1 subfamily, and red indicates the CIN subfamily
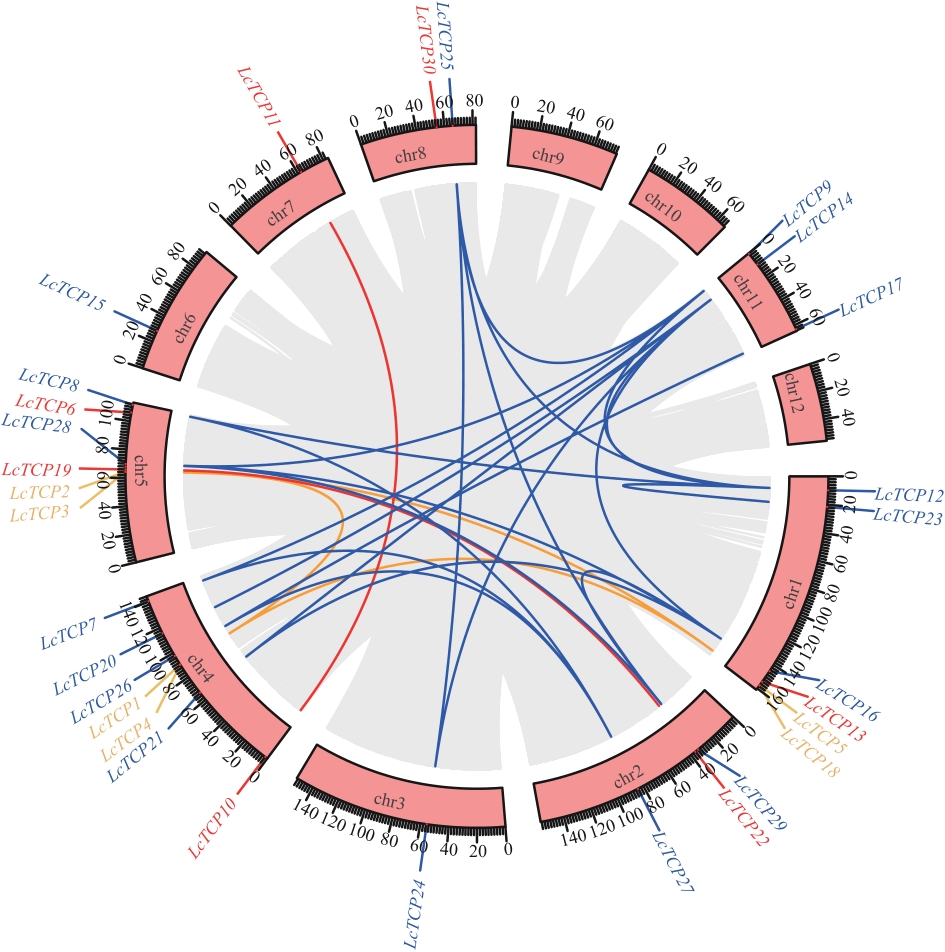
Fig. 4 Synteny relationship of LcTCP genesPink boxes indicate chromosomes, blue lines indicate syntenic relationships among PCF subfamily genes, red lines indicate those of the CIN subfamily, and yellow lines indicate those of the CYC/TB1 subfamily. The gray background indicates segmentally duplicated genes across the L. cubeba genome
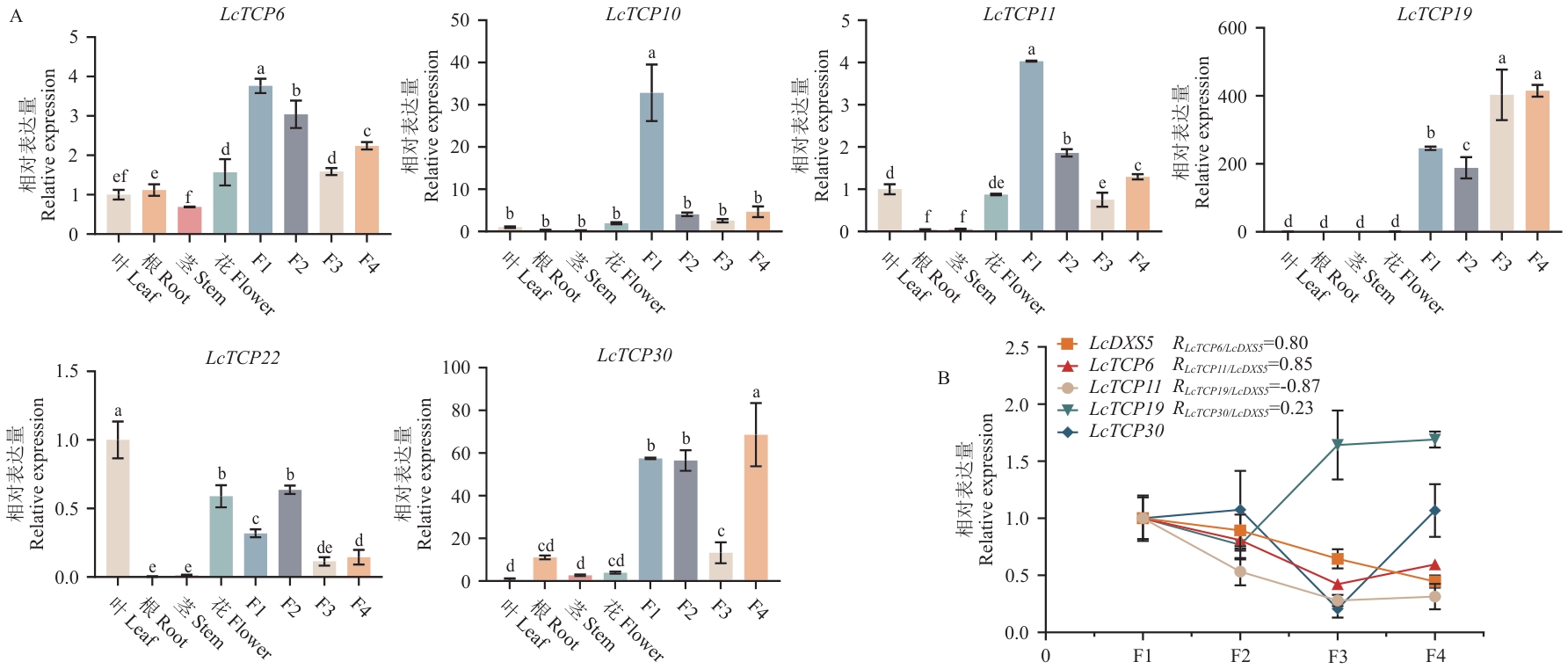
Fig. 6 Expression patterns in different tissues and correlation of expression at different fruit developmental stagesA: Expression patterns of 6 CIN subfamily members from L. cubeba at different fruit developmental stages and in various tissues. B: Expression correlation among LcTCP6, LcTCP11, LcTCP19, LcTCP30, and LcDXS5 at different fruit developmental stages. F1: 30 d after flowering; F2: 60 d after flowering, the peak period of essential oil synthesis; F3: 90 d after flowering; F4: 120 d after flowering. Different lower letters indicate significant differences (P<0.05)
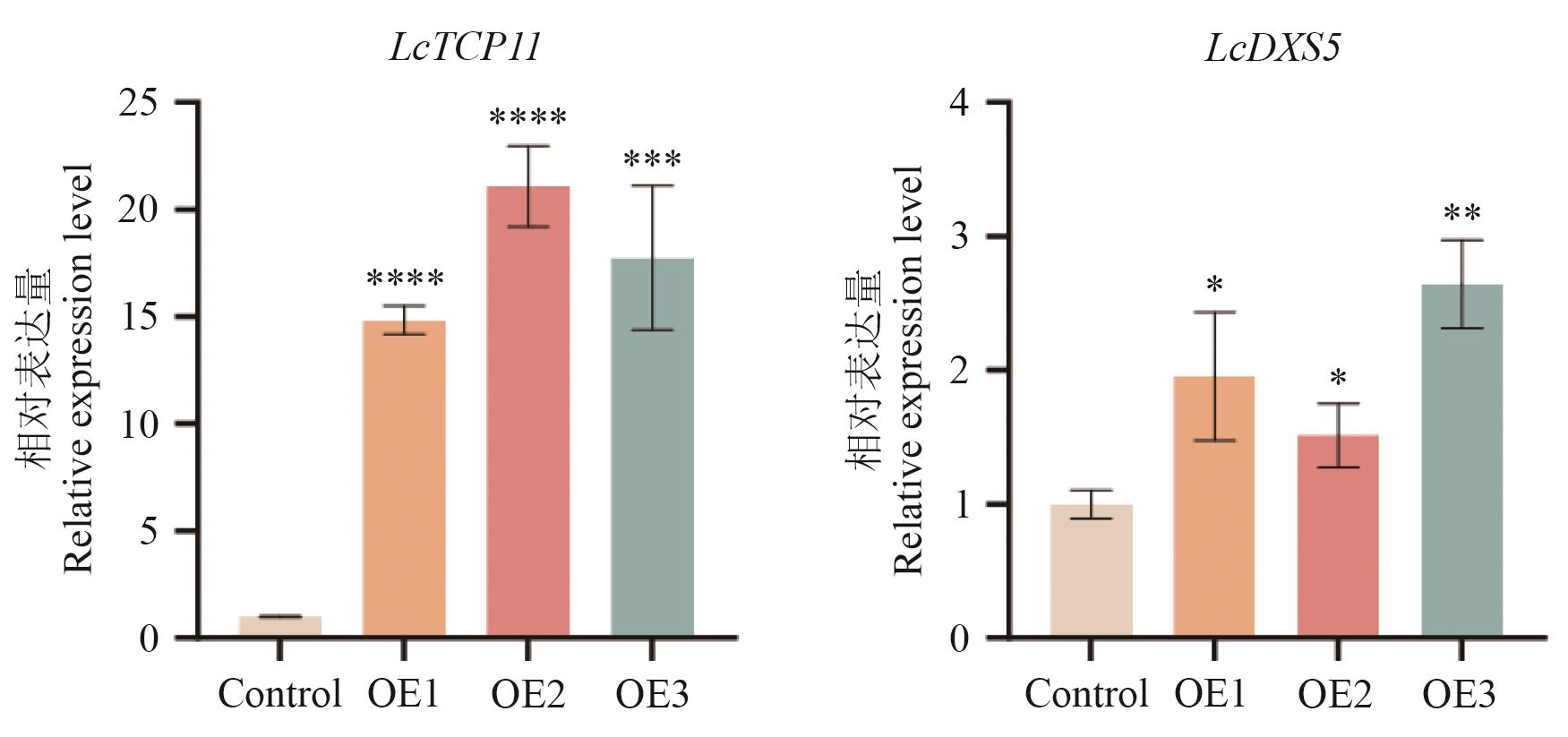
Fig. 7 Expression analysis of LcTCP11 and LcDXS5 in transiently transformed L. cubeba leavesOE1, OE2, and OE3 refer to three overexpression lines. Data are expressed as mean±SD from three replicates (*P<0.05, **P<0.01, ***P<0.001,and ****P<0.000 1)
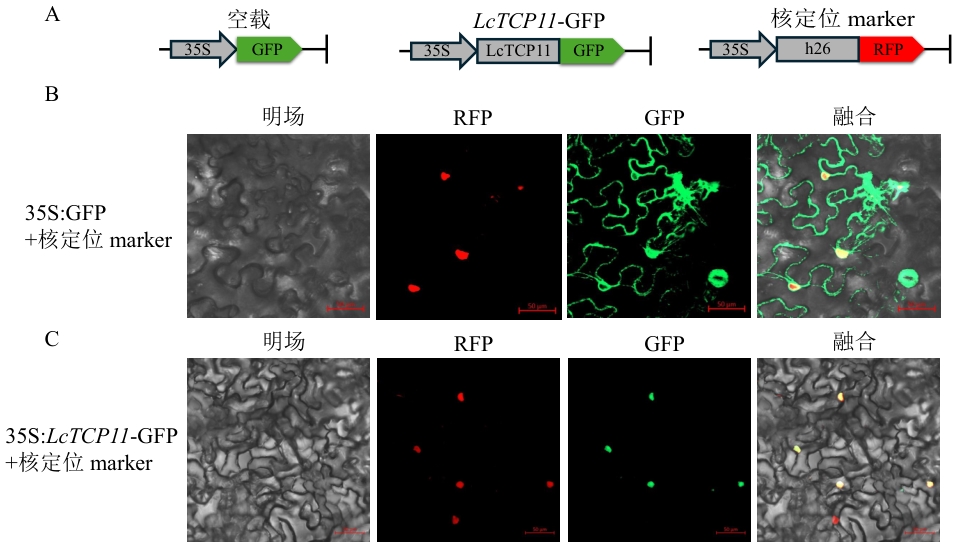
Fig. 8 Subcellular localization of LcTCP11A: Schematic diagram of the vector. B: Subcellular localization of the empty vector. C: Localization of LcTCP11-GFP fusion expression
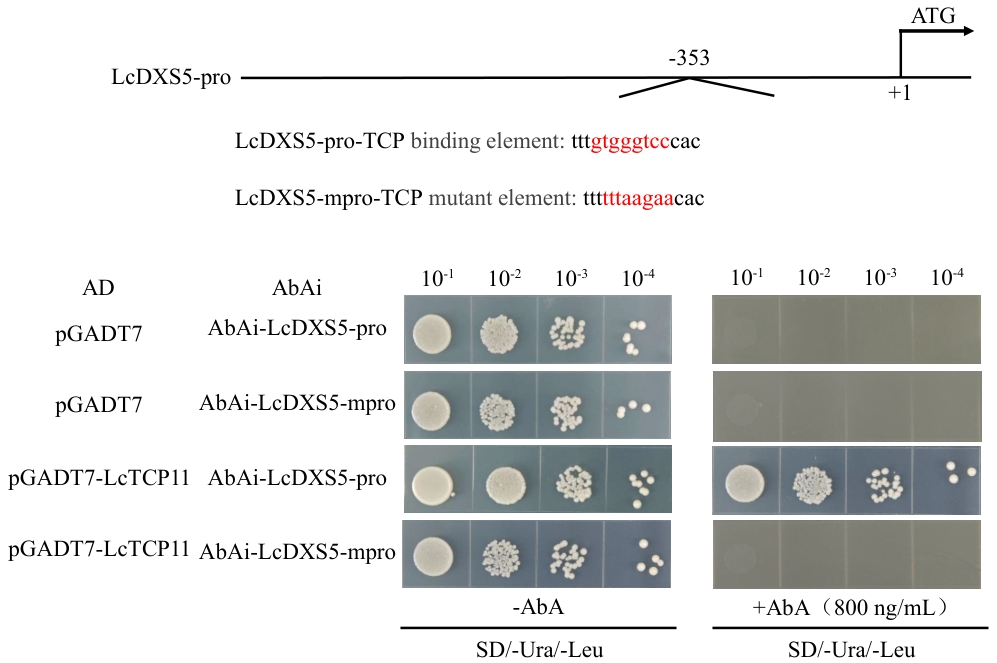
Fig. 9 Y1H assays verifying the recognition of LcTCP11 to the TCP element in LcDXS5 promoterThe red text in the figure indicates the binding element sequences on the promoter and the corresponding mutated element sequences, respectively. Yeast cells co-expressing pGADT7-LcTCP11 and pAbAi-3×gtgggtcc were cultured on selective medium containing AbA (SD/-Trp/-Ura) at 30 ℃ for 48 h. Once single colonies had grown, they were diluted in water to 10¹-10⁴ fold, and 10 μL of each dilution was spotted onto fresh selective medium containing AbA (SD/-Trp/-Ura) for a further 48 h of cultivation
| [1] | Lan JQ, Qin GJ. The regulation of CIN-like TCP transcription factors [J]. Int J Mol Sci, 2020, 21(12): 4498. |
| [2] | Cubas P, Lauter N, Doebley J, et al. The TCP domain: a motif found in proteins regulating plant growth and development [J]. Plant J, 1999, 18(2): 215-222. |
| [3] | Doebley J, Stec A, Hubbard L. The evolution of apical dominance in maize [J]. Nature, 1997, 386(6624): 485-488. |
| [4] | Luo D, Carpenter R, Vincent C, et al. Origin of floral asymmetry in Antirrhinum [J]. Nature, 1996, 383(6603): 794-799. |
| [5] | Shunichi Kosugi YO. PCF1 and PCF2 specifically bind to cis elements in the rice proliferating cell nuclear antigen gene [J]. Plant Cell, 1997, 9(9): 1607-1619. |
| [6] | Navaud O, Dabos P, Carnus E, et al. TCP transcription factors predate the emergence of land plants [J]. J Mol Evol, 2007, 65(1): 23-33. |
| [7] | 马亚男. TCP类转录因子调控青蒿素生物合成的分子机制研究 [D]. 上海: 上海交通大学, 2019. |
| Ma YN. The studies on molecular mechanism of TCP transcription factors regulating artemisinin biosynthesis in Artemisia annua L. [D]. Shanghai: Shanghai Jiao Tong University, 2019. | |
| [8] | Martín-Trillo M, Cubas P. TCP genes: a family snapshot ten years later [J]. Trends Plant Sci, 2010, 15(1): 31-39. |
| [9] | O’Connor C, Varshosaz P, Moise AR. Mechanisms of feedback regulation of vitamin a metabolism [J]. Nutrients, 2022, 14(6): 1312. |
| [10] | Ma N, Zhang ZY, Liao FL, et al. The birth of artemisinin [J]. Pharmacol Ther, 2020, 216: 107658. |
| [11] | Costa EF, Magalhães WV, Di Stasi LC. Recent advances in herbal-derived products with skin anti-aging properties and cosmetic applications [J]. Molecules, 2022, 27(21): 7518. |
| [12] | Thoppil RJ, Bishayee A. Terpenoids as potential chemopreventive and therapeutic agents in liver cancer [J]. World J Hepatol, 2011, 3(9): 228-249. |
| [13] | Bakkali F, Averbeck S, Averbeck D, et al. Biological effects of essential oils-A review [J]. Food Chem Toxicol, 2008, 46(2): 446-475. |
| [14] | Liu XM, Zhang WW, Tang N, et al. Genomic-wide identification and expression analysis of AP2/ERF transcription factors in Zanthoxylum armatum reveals the candidate genes for the biosynthesis of terpenoids [J]. Plant Genome, 2024, 17(1): e20422. |
| [15] | Guo YH, Guo ZY, Zhong J, et al. Positive regulatory role of R2R3 MYBs in terpene biosynthesis in Lilium ‘Siberia’ [J]. Hortic Plant J, 2023, 9(5): 1024-1038. |
| [16] | Li HW, Chen NN, Zhang HB, et al. Multidimensional regulation of transcription factors: decoding the comprehensive signals of plant secondary metabolism [J]. Front Plant Sci, 2025, 16: 1522278. |
| [17] | Liu C, Lv TT, Shen YH, et al. Genome-wide identification and integrated analysis of TCP genes controlling ginsenoside biosynthesis in Panax ginseng [J]. BMC Plant Biol, 2024, 24(1): 47. |
| [18] | Ma YN, Xu DB, Yan X, et al. Jasmonate- and abscisic acid-activated AaGSW1-AaTCP15/AaORA transcriptional cascade promotes artemisinin biosynthesis in Artemisia annua [J]. Plant Biotechnol J, 2021, 19(7): 1412-1428. |
| [19] | Ma YN, Xu DB, Li L, et al. Jasmonate promotes artemisinin biosynthesis by activating the TCP14-ORA complex in Artemisia annua [J]. Sci Adv, 2018, 4(11): eaas9357. |
| [20] | 詹忠根. 转录因子调控青蒿素生物合成的作用机制研究进展 [J]. 中草药, 2022, 53(19): 6258-6272. |
| Zhan ZG. Research progress on transcriptional regulation mechanism of artemisinin biosynthesis [J]. Chin Tradit Herb Drugs, 2022, 53(19): 6258-6272. | |
| [21] | Gao J, Chen YC, Gao M, et al. LcWRKY17, a WRKY transcription factor from Litsea cubeba, effectively promotes monoterpene synthesis [J]. Int J Mol Sci, 2023, 24(8): 7210. |
| [22] | Si LL, Chen YC, Han XJ, et al. Chemical composition of essential oils of Litsea cubeba harvested from its distribution areas in China [J]. Molecules, 2012, 17(6): 7057-7066. |
| [23] | Gao M, Lin LY, Chen YC, et al. Digital gene expression profiling to explore differentially expressed genes associated with terpenoid biosynthesis during fruit development in Litsea cubeba [J]. Molecules, 2016, 21(9): 1251. |
| [24] | Wang MY, Gao M, Zhao YX, et al. LcERF19, an AP2/ERF transcription factor from Litsea cubeba, positively regulates geranial and neral biosynthesis [J]. Hortic Res, 2022, 9: uhac093. |
| [25] | Zhao YX, Wang MY, Chen YC, et al. LcERF134 increases the production of monoterpenes by activating the terpene biosynthesis pathway in Litsea cubeba [J]. Int J Biol Macromol, 2023, 232: 123378. |
| [26] | Ni FF, Zhao YX, Gao M, et al. LcMYB43 enhances monoterpene biosynthesis by activating 1-deoxy-D-xylulose-5-phosphate synthase gene expression in Litsea cubeba [J]. Int J Biol Macromol, 2025, 304(Pt 1): 140860. |
| [27] | Zhao YX, Chen YC, Gao M, et al. Alcohol dehydrogenases regulated by a MYB44 transcription factor underlie Lauraceae citral biosynthesis [J]. Plant Physiol, 2024, 194(3): 1674-1691. |
| [28] | Yang JH, Chen YC, Gao M, et al. Comprehensive identification of bHLH transcription factors in Litsea cubeba reveals candidate gene involved in the monoterpene biosynthesis pathway [J]. Front Plant Sci, 2022, 13: 1081335. |
| [29] | Li ST, Zachgo S. TCP3 interacts with R2R3-MYB proteins, promotes flavonoid biosynthesis and negatively regulates the auxin response in Arabidopsis thaliana [J]. Plant J, 2013, 76(6): 901-913. |
| [30] | 宋姝熠, 蒋开秀, 刘欢艳, 等. ‘红阳’猕猴桃TCP基因家族鉴定及其在果实中的表达分析 [J]. 生物技术通报, 2025, 41(3): 190-201. |
| Song SY, Jiang KX, Liu HY, et al. Identification of the TCP gene family in Actinidia chinensis var. Hongyang and their expression analysis in fruit [J]. Biotechnol Bull, 2025, 41(3): 190-201. | |
| [31] | Chen YC, Li Z, Zhao YX, et al. The Litsea genome and the evolution of the laurel family [J]. Nat Commun, 2020, 11(1): 1675. |
| [32] | Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets [J]. Mol Biol Evol, 2016, 33(7): 1870-1874. |
| [33] | Viola IL, Camoirano A, Gonzalez DH. Redox-dependent modulation of anthocyanin biosynthesis by the TCP transcription factor TCP15 during exposure to high light intensity conditions in Arabidopsis [J]. Plant Physiol, 2016, 170(1): 74-85. |
| [34] | Liu DH, Luo Y, Han H, et al. Genome-wide analysis of citrus TCP transcription factors and their responses to abiotic stresses [J]. BMC Plant Biol, 2022, 22(1): 325. |
| [35] | Zhou Y, Zhang DZ, An JX, et al. TCP transcription factors regulate shade avoidance via directly mediating the expression of both PHYTOCHROME INTERACTING FACTORs and auxin biosynthetic genes [J]. Plant Physiol, 2018, 176(2): 1850-1861. |
| [36] | 唐羽翔, 高旭, 崔亚宁, 等. 植物TCP转录因子研究进展 [J]. 科学通报, 2022, 67(33): 3964-3975. |
| Tang YX, Gao X, Cui YN, et al. Research progress of plant TCP transcription factors [J]. Chin Sci Bull, 2022, 67(33): 3964-3975. | |
| [37] | Viola IL, Gonzalez DH. TCP transcription factors in plant reproductive development: juggling multiple roles [J]. Biomolecules, 2023, 13(5): 750. |
| [38] | Aguilar-Martínez JA, Poza-Carrión C, Cubas P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds [J]. Plant Cell, 2007, 19(2): 458-472. |
| [39] | Koyama T, Mitsuda N, Seki M, et al. TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis [J]. Plant Cell, 2010, 22(11): 3574-3588. |
| [40] | Miao-Ying Wang PZ. The cotton transcription factor TCP14 functions in auxin-mediated epidermal cell differentiation and elongation[J]. Plant Physiol, 2013, 162(3): 1669-1680. |
| [41] | Tian C, Zhai LS, Zhu WJ, et al. Characterization of the TCP gene family in Chrysanthemum nankingense and the role of CnTCP4 in cold tolerance [J]. Plants, 2022, 11(7): 936. |
| [42] | Chen SY, Chen B, Xu XN. Genome-wide identification and expression analysis of TCP transcription factors in Chrysanthemum indicum reveals their critical role in the response to various abiotic stresses [J]. BMC Plant Biol, 2025, 25(1): 631. |
| [43] | Xu YX, Ma L, Zeng XC, et al. Genome-wide identification and analysis of BrTCP transcription factor family genes involved in cold stress tolerance in winter rapeseed (Brassica rapa L.) [J]. Int J Mol Sci, 2024, 25(24): 13592. |
| [44] | Sprenger GA, Schörken U, Wiegert T, et al. Identification of a thiamin-dependent synthase in Escherichia coli required for the formation of the 1-deoxy-D-xylulose 5-phosphate precursor to isoprenoids, thiamin, and pyridoxol [J]. Proc Natl Acad Sci USA, 1997, 94(24): 12857-12862. |
| [45] | Yuan Y, Ren SY, Liu XF, et al. SlWRKY35 positively regulates carotenoid biosynthesis by activating the MEP pathway in tomato fruit [J]. New Phytol, 2022, 234(1): 164-178. |
| [46] | Tian L, Shi JW, Yang L, et al. Molecular cloning and functional analysis of DXS and FPS genes from Zanthoxylum bungeanum Maxim [J]. Foods, 2022, 11(12): 1746. |
| [1] | SUN Ting, ZHANG Yan, LIU Yu-shan, FENG Yuan-yuan, QIN Heng-shan, ZHANG Jun, HE Xiao-gang, ZHANG Jing-rong. Cloning and Functional Analysis of AeF3H Gene in Okra [J]. Biotechnology Bulletin, 2026, 42(4): 153-160. |
| [2] | FENG Cheng-hao, DANG Yu-le, WANG Zhi-ze, NIE Wei-dan, YANG Zhong-min, DU Chong. Screening of Differentially Expressed Genes in Tomato Defense against Meloidogyne incognita Based on RNA-seq and Expression Characteristics Analysis of RPP13 Gene [J]. Biotechnology Bulletin, 2026, 42(4): 141-152. |
| [3] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [4] | ZHAO Yan-xia, LI Qian, SUN Jia-bo, LIANG Hong-min, LI Bing-bing. Key Regulatory Genes and Molecular Networks Dissection Underlying Strawberry Fruit Quality Formation [J]. Biotechnology Bulletin, 2026, 42(3): 111-132. |
| [5] | XU Ze, ZHOU Chen-ping, KUANG Rui-bin, WU Xia-ming, YANG Min, LIU Chuan-he, HE Han, WEI Yue-rong. Identification of PG Gene Family and Their Roles in Papaya Fruit Softening [J]. Biotechnology Bulletin, 2026, 42(3): 349-361. |
| [6] | NONG Wei-you, ZHAO Chang-zu, QIAN Zhen-feng, DING Qian, WANG Yu-jie, CHEN Shu-ying, HE Li-lian, LI Fu-sheng. Identification of the EfBBX Gene Family in Erianthus fulvus and Analysis of Its Expression Patterns Under Cold Stress [J]. Biotechnology Bulletin, 2026, 42(2): 267-277. |
| [7] | ZHANG Dong-ling, ZHANG Yin-sheng, WANG Jian-jun, YE Fei-yu, LU Zi-han, MA Chen-chen, LIU Hua-feng, HU De-sheng, DENG Ya-zhou, CAO Li-ru. Expression Characteristics and Functions of HSFs Transcription Factors Family in Maize under Drought Stress [J]. Biotechnology Bulletin, 2026, 42(2): 178-187. |
| [8] | LONG Lin-xi, ZENG Yin-ping, WANG Qian, DENG Yu-ping, GE Min-qian, CHEN Yan-zhuo, LI Xin-juan, YANG Jun, ZOU Jian. Identification of Sunflower GH3 Gene Family and Analysis of Their Function in Flower Development [J]. Biotechnology Bulletin, 2026, 42(1): 125-138. |
| [9] | YANG Juan, FENG Hui, JI Nai-zhe, SUN Li-ping, WANG Yun, ZHANG Jia-nan, ZHAO Shi-wei. Cloning and Functional Analysis of AP2/ERF Transcription Factors RcERF4 and RcRAP2-12 in Rose [J]. Biotechnology Bulletin, 2026, 42(1): 150-160. |
| [10] | ZHANG Chi-hao, LIU Jin-nan, CHAO Yue-hui. Cloning and Functional Analysis of a bZIP Transcription Factor MtbZIP29 from Medicago truncatula [J]. Biotechnology Bulletin, 2026, 42(1): 241-250. |
| [11] | WU Cui-cui, CHEN Deng-ke, LAN Gang, XIA Zhi, LI Peng-bo. Bioinformatics Analysis of Peanut Transcription Factor AhHDZ70 and Its Tolerances to Salt and Drought [J]. Biotechnology Bulletin, 2026, 42(1): 198-207. |
| [12] | LIU Jia-li, SONG Jing-rong, ZHAO Wen-yu, ZHANG Xin-yuan, ZHAO Zi-yang, CAO Yi-bo, ZHANG Ling-yun. Identification of the R2R3-MYB Gene and Expression Analysis of Flavonoid Regulatory Genes in Blueberry [J]. Biotechnology Bulletin, 2025, 41(9): 124-138. |
| [13] | LI Yu-zhen, LI Meng-dan, ZHANG Wei, PENG Ting. Functional Study of RmEXPB2 Genein Rosa multiflora Based on the Identification of the Expansin Gene Family in Rosa sp. [J]. Biotechnology Bulletin, 2025, 41(9): 182-194. |
| [14] | ZHANG Chao-chao, HAN Kai-yuan, WANG Tong, CHEN Zhong. Cloning and Functional Analysis of PtoYABBY2 and PtoYABBY12 in Populus tomentosa [J]. Biotechnology Bulletin, 2025, 41(9): 256-264. |
| [15] | SHI Fa-chao, JIANG Yong-hua, LIU Hai-lun, WEN Ying-jie, YAN Qian. Cloning and Functional Analysis of LcTFL1 Gene in Litchi chinensis Sonn. [J]. Biotechnology Bulletin, 2025, 41(9): 159-167. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||