








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 250-266.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0481
Previous Articles Next Articles
ZHUANG Na( ), HE Hong-li, ZHANG Xing-zheng(
), HE Hong-li, ZHANG Xing-zheng( )
)
Received:2025-05-13
Online:2026-02-26
Published:2026-03-17
Contact:
ZHANG Xing-zheng
E-mail:m15589101772@163.com;zxz20051986@163.com
ZHUANG Na, HE Hong-li, ZHANG Xing-zheng. Genome-wide Identification and Expression Analysis of the SMXL Gene Family in Medicago truncatula[J]. Biotechnology Bulletin, 2026, 42(2): 250-266.
| 基因名称 Gene name | 基因ID Gene ID | 染色体定位 Location | 基因链 Strand | 外显子数 Exon | 编码区长度 CDS (bp) |
|---|---|---|---|---|---|
| MtSMXL1 | Medtr1g057150 | chr1:25117416..25121709 | + | 3 | 2 427 |
| MtSMXL2 | Medtr1g090640 | chr1:40585935..40589246 | - | 3 | 2 550 |
| MtSMXL3 | Medtr2g038200 | chr2:16610767..16615642 | + | 3 | 3 030 |
| MtSMXL4 | Medtr3g461000 | chr3:24065055..24068423 | + | 8 | 2 449 |
| MtSMXL5 | Medtr3g464150 | chr3:25772408..25778862 | + | 3 | 3 243 |
| MtSMXL6 | Medtr3g070850 | chr3:31780989..31785558 | + | 3 | 3 246 |
| MtSMXL7 | Medtr3g098310 | chr3:44871095..44878208 | - | 13 | 2 859 |
| MtSMXL8 | Medtr3g498725 | chr3:45109729..45116929 | + | 11 | 2 769 |
| MtSMXL9 | Medtr3g102830 | chr3:47377876..47383774 | + | 10 | 2 784 |
| MtSMXL10 | Medtr3g106160 | chr3:49006958..49015432 | - | 10 | 2 943 |
| MtSMXL11 | Medtr4g109450 | chr4:45465442..45469308 | + | 5 | 2 739 |
| MtSMXL12 | Medtr4g127290 | chr4:52803794..52807296 | - | 3 | 2 463 |
| MtSMXL13 | Medtr4g129230 | chr4:53817793..53821564 | + | 3 | 3 126 |
| MtSMXL14 | Medtr5g071060 | chr5:30105997..30111708 | + | 3 | 3 279 |
| MtSMXL15 | Medtr6g013660 | chr6:4395062..4403563 | - | 10 | 2 925 |
| MtSMXL16 | Medtr7g012820 | chr7:3513549..3521223 | - | 10 | 2 937 |
| MtSMXL17 | Medtr8g100040 | chr8:40241178..40246683 | + | 10 | 2 781 |
Table 1 Basic information of MtSMXL genes
| 基因名称 Gene name | 基因ID Gene ID | 染色体定位 Location | 基因链 Strand | 外显子数 Exon | 编码区长度 CDS (bp) |
|---|---|---|---|---|---|
| MtSMXL1 | Medtr1g057150 | chr1:25117416..25121709 | + | 3 | 2 427 |
| MtSMXL2 | Medtr1g090640 | chr1:40585935..40589246 | - | 3 | 2 550 |
| MtSMXL3 | Medtr2g038200 | chr2:16610767..16615642 | + | 3 | 3 030 |
| MtSMXL4 | Medtr3g461000 | chr3:24065055..24068423 | + | 8 | 2 449 |
| MtSMXL5 | Medtr3g464150 | chr3:25772408..25778862 | + | 3 | 3 243 |
| MtSMXL6 | Medtr3g070850 | chr3:31780989..31785558 | + | 3 | 3 246 |
| MtSMXL7 | Medtr3g098310 | chr3:44871095..44878208 | - | 13 | 2 859 |
| MtSMXL8 | Medtr3g498725 | chr3:45109729..45116929 | + | 11 | 2 769 |
| MtSMXL9 | Medtr3g102830 | chr3:47377876..47383774 | + | 10 | 2 784 |
| MtSMXL10 | Medtr3g106160 | chr3:49006958..49015432 | - | 10 | 2 943 |
| MtSMXL11 | Medtr4g109450 | chr4:45465442..45469308 | + | 5 | 2 739 |
| MtSMXL12 | Medtr4g127290 | chr4:52803794..52807296 | - | 3 | 2 463 |
| MtSMXL13 | Medtr4g129230 | chr4:53817793..53821564 | + | 3 | 3 126 |
| MtSMXL14 | Medtr5g071060 | chr5:30105997..30111708 | + | 3 | 3 279 |
| MtSMXL15 | Medtr6g013660 | chr6:4395062..4403563 | - | 10 | 2 925 |
| MtSMXL16 | Medtr7g012820 | chr7:3513549..3521223 | - | 10 | 2 937 |
| MtSMXL17 | Medtr8g100040 | chr8:40241178..40246683 | + | 10 | 2 781 |
基因名称 Gene name | 氨基酸数 Peptide (aa) | 分子质量 Mass of molecule (kD) | 等电点 Theoretical pI | 不稳定系数 Instability index | 脂肪指数 Aliphatic index | 平均亲水指数 GRAVY | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| MtSMXL1 | 808 | 90.50 | 5.74 | 46.88 | 82.86 | -0.502 | 叶绿体 |
| MtSMXL2 | 849 | 95.17 | 6.03 | 54.29 | 82.99 | -0.362 | 叶绿体 |
| MtSMXL3 | 1 009 | 113.46 | 6.41 | 51.71 | 78.12 | -0.509 | 细胞核 |
| MtSMXL4 | 810 | 90.80 | 7.33 | 37.38 | 97.32 | -0.233 | 细胞质 |
| MtSMXL5 | 1 080 | 120.931 | 6.27 | 50.68 | 85.24 | -0.307 | 细胞核 |
| MtSMXL6 | 1 081 | 119.74 | 6.34 | 50.45 | 84.38 | -0.356 | 细胞核 |
| MtSMXL7 | 963 | 107.09 | 7.04 | 47.16 | 89.41 | -0.287 | 叶绿体 |
| MtSMXL8 | 922 | 102.71 | 6.35 | 40.68 | 95.40 | -0.341 | 叶绿体 |
| MtSMXL9 | 927 | 103.07 | 6.41 | 40.14 | 93.50 | -0.360 | 叶绿体 |
| MtSMXL10 | 980 | 109.89 | 6.53 | 38.67 | 92.85 | -0.385 | 线粒体 |
| MtSMXL11 | 912 | 101.35 | 5.85 | 36.33 | 96.23 | -0.402 | 细胞质 |
| MtSMXL12 | 820 | 92.33 | 8.18 | 58.46 | 77.52 | -0.524 | 叶绿体 |
| MtSMXL13 | 1 027 | 112.90 | 6.34 | 46.61 | 83.50 | -0.471 | 细胞质 |
| MtSMXL14 | 1 092 | 120.61 | 6.05 | 43.61 | 81.12 | -0.254 | 细胞核 |
| MtSMXL15 | 974 | 109.96 | 6.65 | 42.89 | 89.30 | -0.474 | 叶绿体 |
| MtSMXL16 | 978 | 110.28 | 6.26 | 37.80 | 90.14 | -0.455 | 线粒体 |
| MtSMXL17 | 926 | 102.78 | 6.44 | 40.51 | 94.04 | -0.357 | 叶绿体 |
Table 2 Prediction of physical and chemical properties of MtSMXLproteins
基因名称 Gene name | 氨基酸数 Peptide (aa) | 分子质量 Mass of molecule (kD) | 等电点 Theoretical pI | 不稳定系数 Instability index | 脂肪指数 Aliphatic index | 平均亲水指数 GRAVY | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| MtSMXL1 | 808 | 90.50 | 5.74 | 46.88 | 82.86 | -0.502 | 叶绿体 |
| MtSMXL2 | 849 | 95.17 | 6.03 | 54.29 | 82.99 | -0.362 | 叶绿体 |
| MtSMXL3 | 1 009 | 113.46 | 6.41 | 51.71 | 78.12 | -0.509 | 细胞核 |
| MtSMXL4 | 810 | 90.80 | 7.33 | 37.38 | 97.32 | -0.233 | 细胞质 |
| MtSMXL5 | 1 080 | 120.931 | 6.27 | 50.68 | 85.24 | -0.307 | 细胞核 |
| MtSMXL6 | 1 081 | 119.74 | 6.34 | 50.45 | 84.38 | -0.356 | 细胞核 |
| MtSMXL7 | 963 | 107.09 | 7.04 | 47.16 | 89.41 | -0.287 | 叶绿体 |
| MtSMXL8 | 922 | 102.71 | 6.35 | 40.68 | 95.40 | -0.341 | 叶绿体 |
| MtSMXL9 | 927 | 103.07 | 6.41 | 40.14 | 93.50 | -0.360 | 叶绿体 |
| MtSMXL10 | 980 | 109.89 | 6.53 | 38.67 | 92.85 | -0.385 | 线粒体 |
| MtSMXL11 | 912 | 101.35 | 5.85 | 36.33 | 96.23 | -0.402 | 细胞质 |
| MtSMXL12 | 820 | 92.33 | 8.18 | 58.46 | 77.52 | -0.524 | 叶绿体 |
| MtSMXL13 | 1 027 | 112.90 | 6.34 | 46.61 | 83.50 | -0.471 | 细胞质 |
| MtSMXL14 | 1 092 | 120.61 | 6.05 | 43.61 | 81.12 | -0.254 | 细胞核 |
| MtSMXL15 | 974 | 109.96 | 6.65 | 42.89 | 89.30 | -0.474 | 叶绿体 |
| MtSMXL16 | 978 | 110.28 | 6.26 | 37.80 | 90.14 | -0.455 | 线粒体 |
| MtSMXL17 | 926 | 102.78 | 6.44 | 40.51 | 94.04 | -0.357 | 叶绿体 |
蛋白质 Protein | α螺旋 Alpha helix | 延伸链 Extended strand | β转角 β-turn | 无规则卷曲 Random coil | ||||
|---|---|---|---|---|---|---|---|---|
长度 Length (aa) | 所占比例 Proportion (%) | 长度 Length (aa) | 所占比例 Proportion (%) | 长度 Length (aa) | 所占比例 Proportion (%) | 长度 Length (aa) | 所占比例 Proportion (%) | |
| MtSMXL1 | 382 | 47.28 | 68 | 8.42 | 21 | 2.60 | 358 | 44.31 |
| MtSMXL2 | 366 | 43.11 | 53 | 6.24 | 13 | 1.53 | 430 | 50.65 |
| MtSMXL3 | 440 | 43.61 | 77 | 7.63 | 17 | 1.68 | 492 | 48.76 |
| MtSMXL4 | 417 | 51.48 | 87 | 10.74 | 51 | 6.30 | 306 | 37.78 |
| MtSMXL5 | 443 | 41.02 | 88 | 8.15 | 14 | 1.30 | 549 | 50.83 |
| MtSMXL6 | 406 | 37.56 | 99 | 9.16 | 19 | 1.76 | 576 | 53.28 |
| MtSMXL7 | 506 | 52.54 | 86 | 8.93 | 51 | 5.30 | 371 | 38.53 |
| MtSMXL8 | 496 | 53.80 | 87 | 9.44 | 51 | 5.53 | 339 | 36.77 |
| MtSMXL9 | 523 | 56.42 | 85 | 9.17 | 50 | 5.39 | 319 | 34.41 |
| MtSMXL10 | 563 | 57.45 | 82 | 8.37 | 44 | 4.49 | 335 | 34.18 |
| MtSMXL11 | 550 | 60.31 | 84 | 9.21 | 45 | 4.93 | 278 | 30.48 |
| MtSMXL12 | 363 | 44.27 | 53 | 6.46 | 7 | 0.85 | 404 | 49.27 |
| MtSMXL13 | 512 | 49.85 | 70 | 6.82 | 20 | 1.95 | 445 | 43.33 |
| MtSMXL14 | 471 | 43.13 | 84 | 7.69 | 16 | 1.47 | 537 | 49.18 |
| MtSMXL15 | 554 | 56.88 | 84 | 8.62 | 46 | 4.72 | 336 | 34.50 |
| MtSMXL16 | 545 | 55.73 | 86 | 8.79 | 45 | 4.60 | 347 | 35.48 |
| MtSMXL17 | 507 | 54.75 | 92 | 9.94 | 49 | 5.29 | 327 | 35.31 |
Table 3 Secondary structure prediction of the MtSMXLs protein
蛋白质 Protein | α螺旋 Alpha helix | 延伸链 Extended strand | β转角 β-turn | 无规则卷曲 Random coil | ||||
|---|---|---|---|---|---|---|---|---|
长度 Length (aa) | 所占比例 Proportion (%) | 长度 Length (aa) | 所占比例 Proportion (%) | 长度 Length (aa) | 所占比例 Proportion (%) | 长度 Length (aa) | 所占比例 Proportion (%) | |
| MtSMXL1 | 382 | 47.28 | 68 | 8.42 | 21 | 2.60 | 358 | 44.31 |
| MtSMXL2 | 366 | 43.11 | 53 | 6.24 | 13 | 1.53 | 430 | 50.65 |
| MtSMXL3 | 440 | 43.61 | 77 | 7.63 | 17 | 1.68 | 492 | 48.76 |
| MtSMXL4 | 417 | 51.48 | 87 | 10.74 | 51 | 6.30 | 306 | 37.78 |
| MtSMXL5 | 443 | 41.02 | 88 | 8.15 | 14 | 1.30 | 549 | 50.83 |
| MtSMXL6 | 406 | 37.56 | 99 | 9.16 | 19 | 1.76 | 576 | 53.28 |
| MtSMXL7 | 506 | 52.54 | 86 | 8.93 | 51 | 5.30 | 371 | 38.53 |
| MtSMXL8 | 496 | 53.80 | 87 | 9.44 | 51 | 5.53 | 339 | 36.77 |
| MtSMXL9 | 523 | 56.42 | 85 | 9.17 | 50 | 5.39 | 319 | 34.41 |
| MtSMXL10 | 563 | 57.45 | 82 | 8.37 | 44 | 4.49 | 335 | 34.18 |
| MtSMXL11 | 550 | 60.31 | 84 | 9.21 | 45 | 4.93 | 278 | 30.48 |
| MtSMXL12 | 363 | 44.27 | 53 | 6.46 | 7 | 0.85 | 404 | 49.27 |
| MtSMXL13 | 512 | 49.85 | 70 | 6.82 | 20 | 1.95 | 445 | 43.33 |
| MtSMXL14 | 471 | 43.13 | 84 | 7.69 | 16 | 1.47 | 537 | 49.18 |
| MtSMXL15 | 554 | 56.88 | 84 | 8.62 | 46 | 4.72 | 336 | 34.50 |
| MtSMXL16 | 545 | 55.73 | 86 | 8.79 | 45 | 4.60 | 347 | 35.48 |
| MtSMXL17 | 507 | 54.75 | 92 | 9.94 | 49 | 5.29 | 327 | 35.31 |
| 基序 Motif | Sites | 长度 Length (aa) | 序列 Sequence | Logo |
|---|---|---|---|---|
| 1 | 9 | 50 | MSEYMERHTVSRLIGAPPGYVGYEEGGQLTEAVRRRPYTVVLFDEIEKAH |  |
| 2 | 9 | 50 | GATTLDEYRKHIEKDPALERRFQPVKVPEPSVEDTISILKGLRERYEJHH |  |
| 3 | 9 | 50 | GKLDPVIGRDEZIERVIQILSRRTKNNPVLIGEPGVGKTAIAEGLAQRIV |  |
| 4 | 17 | 41 | NPNRPIASFMFLGPTGVGKTELAKALAEYVFGSEEALIRJD |  |
| 5 | 15 | 35 | LQIJEDGRLTDSQGRTVSFKNTIIIMTSNVGSSVI |  |
| 6 | 17 | 38 | QZELTEEAAKVJKLAVEVARRRGHAQVTPLHIASALLS |  |
| 7 | 9 | 50 | KVRYTDDALVAAAELSDRYISDRFLPDKAIDLIDEAGSKVRLZHASKPEE |  |
| 8 | 7 | 50 | VLKEVTESSGZIILFIDEIHTVIGAGATEGAMDAANJLKPMLARGELRCI |  |
| 9 | 9 | 49 | WTGIPVSKLSQDERERLLKLEDTLHKRVIGQDEAVEAIARAIRRSRVGL |  |
| 10 | 9 | 50 | KDSSYNRIKSLVMEELRQYFRPEFLNRLDEMIVFRPLTKLZVKEIADJML |  |
| 11 | 9 | 50 | VTERARDLVVDEGYBPSYGARPLRRAIQRLVEBELAEKMLRREIKEGDSV |  |
| 12 | 8 | 29 | VKVELEQLIJSILDDPSVSRVMREAGFSS |  |
| 13 | 9 | 24 | PPVSNALMAAJKRAQAHQRRGPIE |  |
| 14 | 17 | 29 | DVPETLHPLQLIALDLGLLVALAKYPGEF |  |
| 15 | 4 | 50 | ARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGE |  |
Table 4 Conserved motifs identification of MtSMXLs
| 基序 Motif | Sites | 长度 Length (aa) | 序列 Sequence | Logo |
|---|---|---|---|---|
| 1 | 9 | 50 | MSEYMERHTVSRLIGAPPGYVGYEEGGQLTEAVRRRPYTVVLFDEIEKAH |  |
| 2 | 9 | 50 | GATTLDEYRKHIEKDPALERRFQPVKVPEPSVEDTISILKGLRERYEJHH |  |
| 3 | 9 | 50 | GKLDPVIGRDEZIERVIQILSRRTKNNPVLIGEPGVGKTAIAEGLAQRIV |  |
| 4 | 17 | 41 | NPNRPIASFMFLGPTGVGKTELAKALAEYVFGSEEALIRJD |  |
| 5 | 15 | 35 | LQIJEDGRLTDSQGRTVSFKNTIIIMTSNVGSSVI |  |
| 6 | 17 | 38 | QZELTEEAAKVJKLAVEVARRRGHAQVTPLHIASALLS |  |
| 7 | 9 | 50 | KVRYTDDALVAAAELSDRYISDRFLPDKAIDLIDEAGSKVRLZHASKPEE |  |
| 8 | 7 | 50 | VLKEVTESSGZIILFIDEIHTVIGAGATEGAMDAANJLKPMLARGELRCI |  |
| 9 | 9 | 49 | WTGIPVSKLSQDERERLLKLEDTLHKRVIGQDEAVEAIARAIRRSRVGL |  |
| 10 | 9 | 50 | KDSSYNRIKSLVMEELRQYFRPEFLNRLDEMIVFRPLTKLZVKEIADJML |  |
| 11 | 9 | 50 | VTERARDLVVDEGYBPSYGARPLRRAIQRLVEBELAEKMLRREIKEGDSV |  |
| 12 | 8 | 29 | VKVELEQLIJSILDDPSVSRVMREAGFSS |  |
| 13 | 9 | 24 | PPVSNALMAAJKRAQAHQRRGPIE |  |
| 14 | 17 | 29 | DVPETLHPLQLIALDLGLLVALAKYPGEF |  |
| 15 | 4 | 50 | ARQLGHNYIGSEHLLLGLLREGEGVAARVLENLGADPTNIRTQVIRMVGE |  |
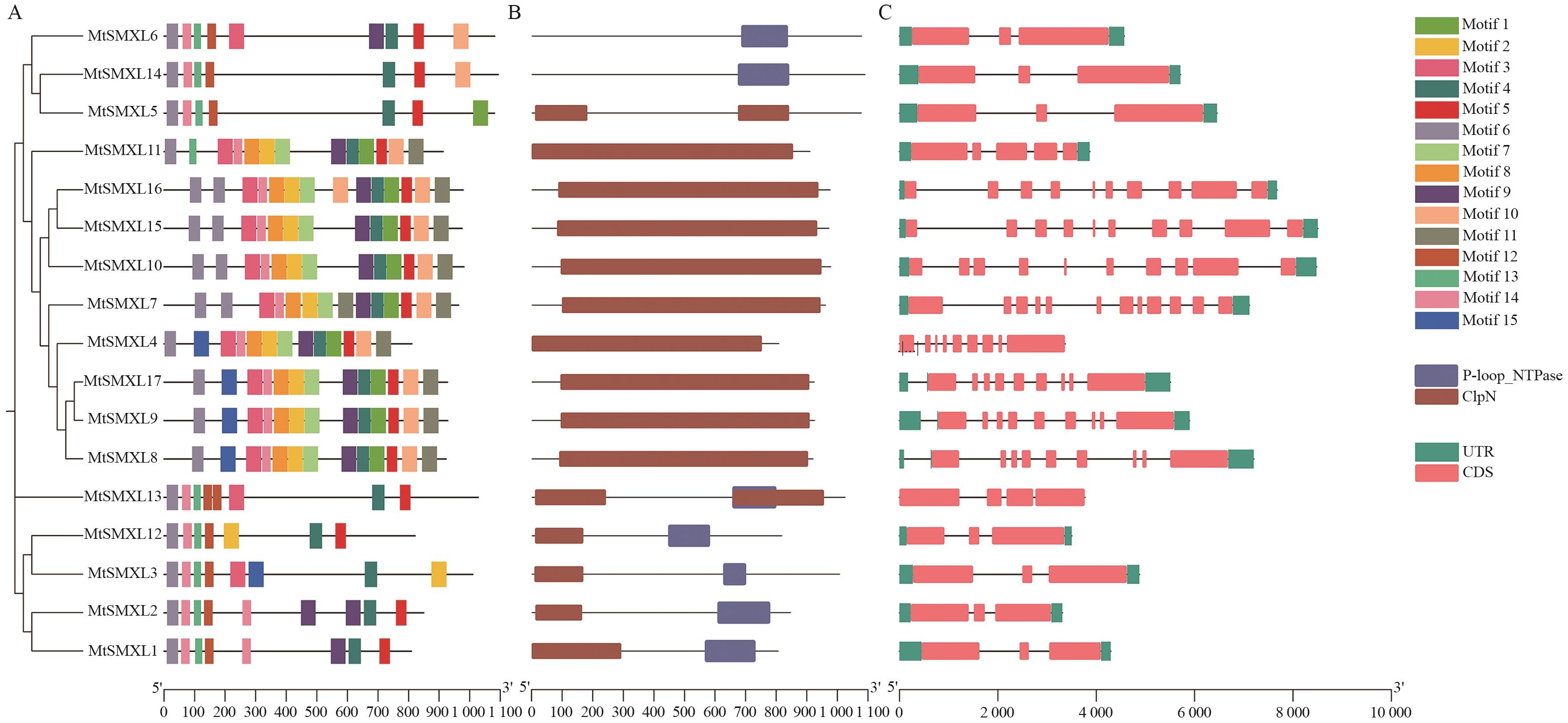
Fig. 3 Conserved motifs, protein conserved domains, and gene structure of MtSMXLs in M. truncatulaA: Conserved motif of MtSMXL protein, different squares indicate different motif. B: MtSMXL protein domain, different squares indicate different protein domains. C: Gene structure of MtSMXL genes
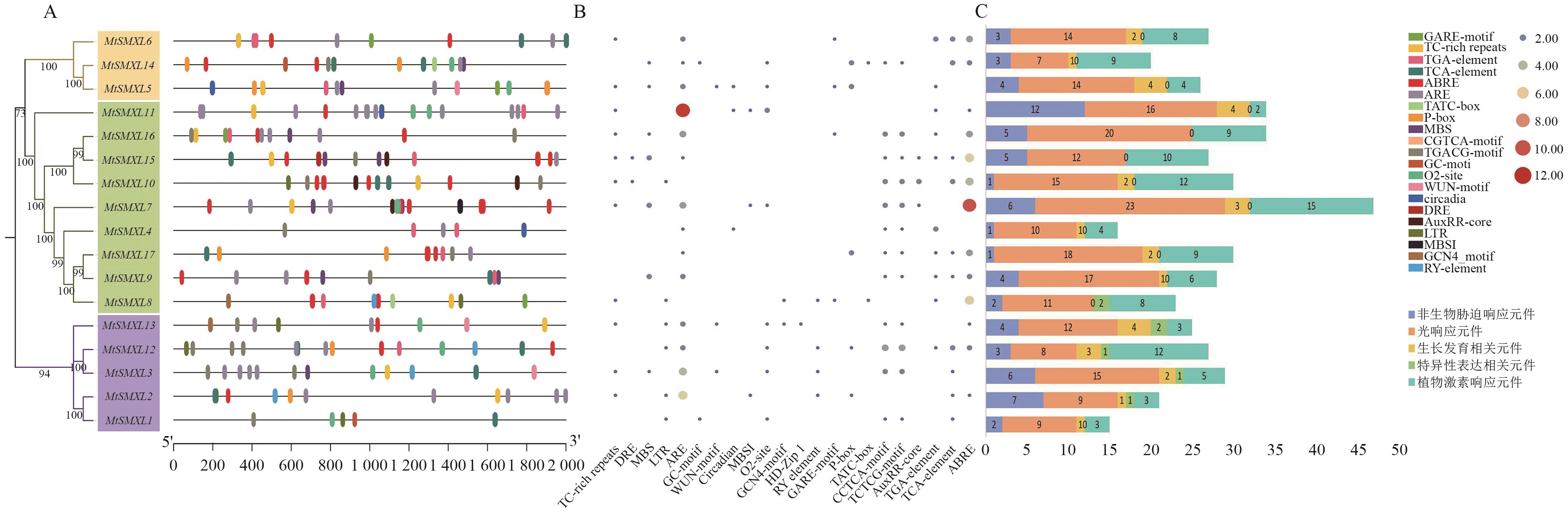
Fig. 4 Cis-acting elements in the promoter of the SMXL genesA: Distribution of cis-acting element in the promoter region of the MtSMXL gene, with different colored ovals representing different cis-elements. B: Heatmap of the number of cis-acting elements in the promoter region of M . C: Stacked statistical chart of various cis-acting elements in the promoter region of the MtSMXL gene. Due to the overlap of promoter positions and space limitations, light-responsive elements and some growth- and development-related elements irrelevant to this study are not shown in the promoter position map A and the heatmap B
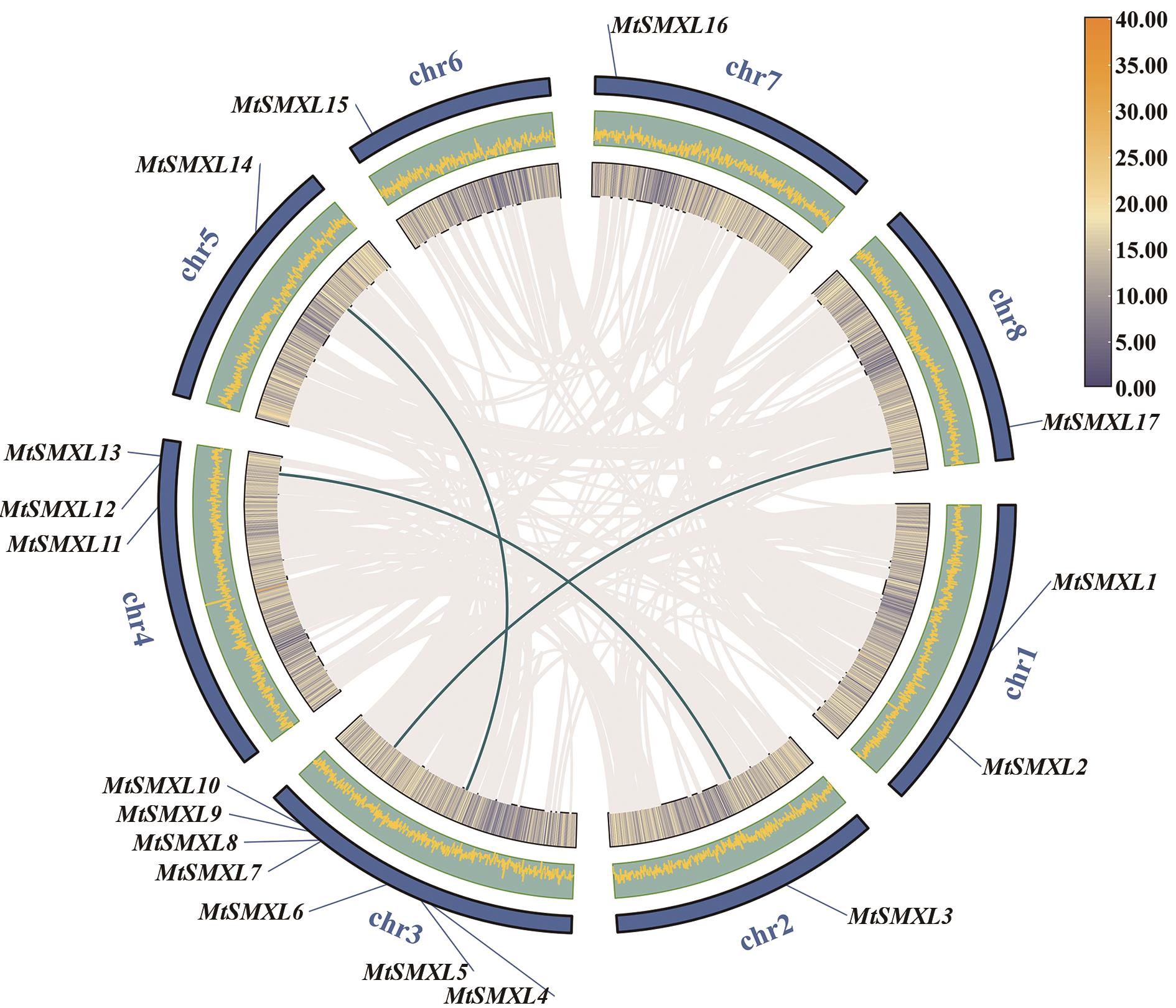
Fig. 5 Intraspecific collinearity of the MtSMXL genesThe gray lines indicate collinearity gene pairs in the genome, and the green lines indicate gene pairs ofcollinear MtSMXL. The inner and middle circles indicate the gene density of the M. truncatula genome, and the outermost circle indicates the chromosomes. The legend indicates the gene density in the inner circle, where yellow shows high gene density and blue shows low gene density
| 基因对Gene pairs | Ka | Ks | Ka/Ks | 分离时间Estimated time(Mya) | |
|---|---|---|---|---|---|
| MtSMXL3 | MtSMXL12 | 0.253 | 0.919 | 0.275 | 75.3 |
| MtSMXL5 | MtSMXL14 | 0.255 | 0.721 | 0.354 | 59.08 |
| MtSMXL8 | MtSMXL17 | 0.056 | 0.68 | 0.082 | 55.74 |
Table 5 Estimated separation time of MtSMXL gene pairs
| 基因对Gene pairs | Ka | Ks | Ka/Ks | 分离时间Estimated time(Mya) | |
|---|---|---|---|---|---|
| MtSMXL3 | MtSMXL12 | 0.253 | 0.919 | 0.275 | 75.3 |
| MtSMXL5 | MtSMXL14 | 0.255 | 0.721 | 0.354 | 59.08 |
| MtSMXL8 | MtSMXL17 | 0.056 | 0.68 | 0.082 | 55.74 |
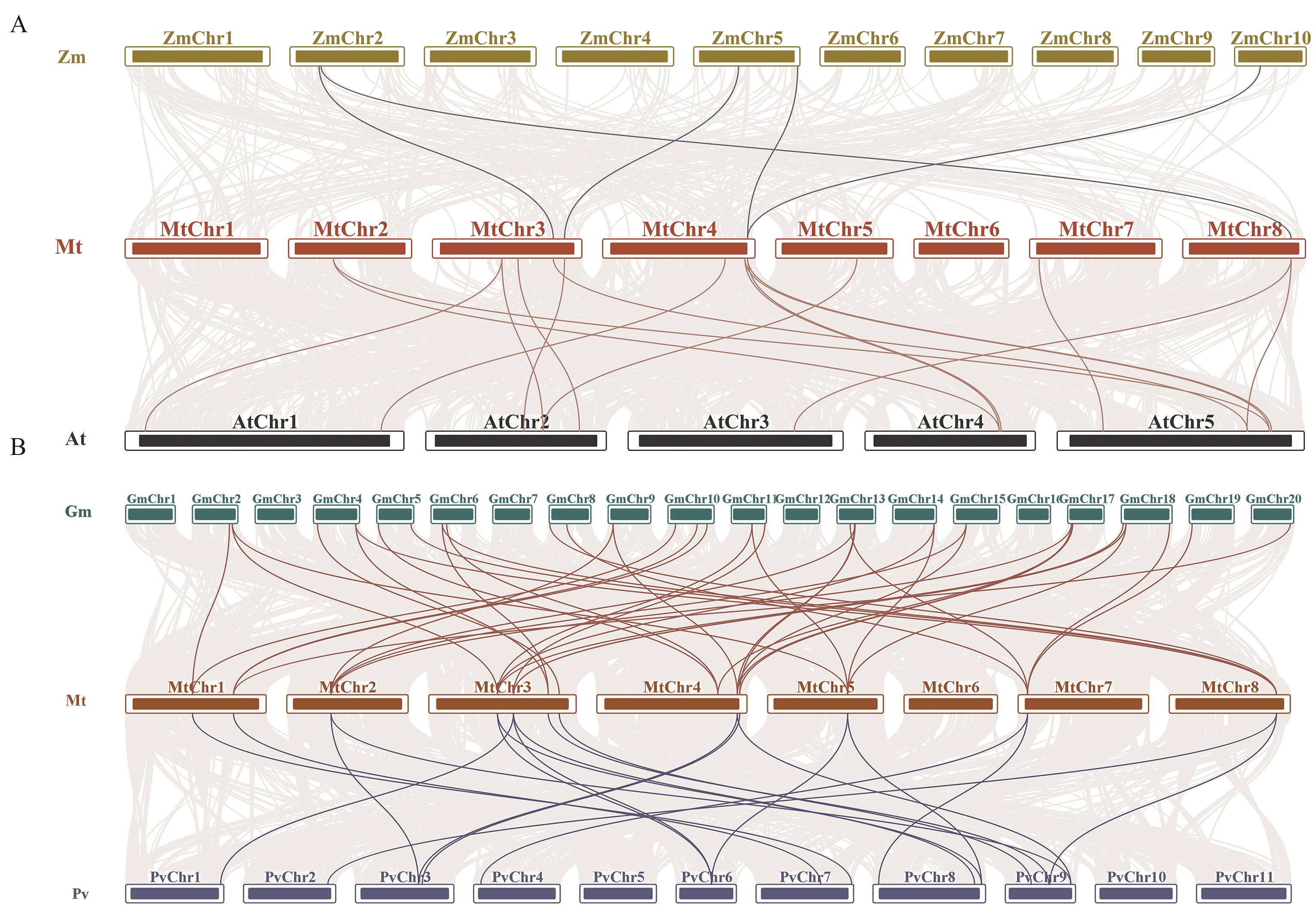
Fig. 6 Collinearity analysis of SMXL genes between M. truncatula and other speciesA: Collinearity analysis of SMXL family genes in M. truncatula, Glycine max and common bean. B: Collinearity analysis of SMXL family genes in M. truncatula, Z. mays and Arabidopsis. The collinear regions in the genomes of M. truncatula and other species are shown by gray lines, and the collinear SMXL gene pairs with other species are highlighted by bright lines
蒺藜苜蓿 M. truncatula | 拟南芥 A. thaliana | 玉米 Z. mays | 菜豆 P. vulgaris | 大豆 G. max | 蒺藜苜蓿 M. truncatula | 拟南芥 A. thaliana | 玉米 Z. mays | 菜豆 P. vulgaris | 大豆 G. max |
|---|---|---|---|---|---|---|---|---|---|
| MtSMXL1 | ESW17795 | Glyma.02G198000.1 | MtSMXL11 | AT1G74310.1 | Glyma.05G022200.1 | ||||
| Glyma.10G079000.1 | Glyma.06G202200.2 | ||||||||
| MtSMXL2 | ESW16225 | Glyma.10G121542.2 | Glyma.17G077500.1 | ||||||
| Glyma.10G196300.1 | MtSMXL12 | AT4G29920.2 | ESW10759 | Glyma.09G060500.1 | |||||
| Glyma.20G193900.1 | AT5G57130.1 | ESW26697 | Glyma.13G101300.1 | ||||||
| MtSMXL3 | AT4G29920.2 | ESW10759 | Glyma.09G060500.1 | Glyma.15G167000.1 | |||||
| AT5G57130.1 | ESW26697 | Glyma.13G101300.1 | Glyma.17G058400.1 | ||||||
| Glyma.15G167000.1 | MtSMXL13 | AT4G30350.1 | Zm00001eb412800_T002 | ESW26813 | Glyma.13G092700.1 | ||||
| Glyma.17G058400.1 | AT5G57710.1 | Zm00001eb257430_T002 | Glyma.17G067700.1 | ||||||
| MtSMXL4 | MtSMXL14 | AT2G29970.1 | ESW14200 | Glyma.02G226900.6 | |||||
| MtSMXL5 | AT1G07200.2 | ESW14200 | Glyma.02G226900.6 | ESW18766 | Glyma.11G174500.4 | ||||
| AT2G29970.1 | ESW18766 | Glyma.11G174500.4 | Glyma.14G193900.2 | ||||||
| Glyma.14G193900.2 | Glyma.18G062300.1 | ||||||||
| Glyma.18G062300.1 | MtSMXL15 | ||||||||
| MtSMXL6 | AT2G40130.2 | ESW13755 | Glyma.11G230700.1 | MtSMXL16 | AT5G15450.1 | ESW11501 | Glyma.08G242100.1 | ||
| ESW18766 | Glyma.18G026600.3 | ESW23286 | Glyma.13G058600.1 | ||||||
| ESW35546 | Glyma.18G062300.1 | Glyma.18G264400.1 | |||||||
| MtSMXL7 | AT5G51070.1 | Zm00001eb084420_T004 | ESW09825 | Glyma.04G203300.1 | Glyma.19G027900.1 | ||||
| Glyma.06G162200.1 | MtSMXL17 | AT3G48870.3 | Zm00001eb084890_T001 | ESW09858 | Glyma.04G200400.8 | ||||
| MtSMXL8 | AT5G50920.1 | ESW31965 | Glyma.05G201100.4 | ||||||
| MtSMXL9 | Glyma.06G165200.6 | ||||||||
| MtSMXL10 | AT2G25140.1 | Zm00001eb234160_T001 | ESW08954 | Glyma.04G062200.1 | Glyma.08G008400.1 |
Table 6 Synteny analysis of MdSMXLs and AtSMXLs, GmSMXLs, PvSMXLs, and ZmSMXLs gene
蒺藜苜蓿 M. truncatula | 拟南芥 A. thaliana | 玉米 Z. mays | 菜豆 P. vulgaris | 大豆 G. max | 蒺藜苜蓿 M. truncatula | 拟南芥 A. thaliana | 玉米 Z. mays | 菜豆 P. vulgaris | 大豆 G. max |
|---|---|---|---|---|---|---|---|---|---|
| MtSMXL1 | ESW17795 | Glyma.02G198000.1 | MtSMXL11 | AT1G74310.1 | Glyma.05G022200.1 | ||||
| Glyma.10G079000.1 | Glyma.06G202200.2 | ||||||||
| MtSMXL2 | ESW16225 | Glyma.10G121542.2 | Glyma.17G077500.1 | ||||||
| Glyma.10G196300.1 | MtSMXL12 | AT4G29920.2 | ESW10759 | Glyma.09G060500.1 | |||||
| Glyma.20G193900.1 | AT5G57130.1 | ESW26697 | Glyma.13G101300.1 | ||||||
| MtSMXL3 | AT4G29920.2 | ESW10759 | Glyma.09G060500.1 | Glyma.15G167000.1 | |||||
| AT5G57130.1 | ESW26697 | Glyma.13G101300.1 | Glyma.17G058400.1 | ||||||
| Glyma.15G167000.1 | MtSMXL13 | AT4G30350.1 | Zm00001eb412800_T002 | ESW26813 | Glyma.13G092700.1 | ||||
| Glyma.17G058400.1 | AT5G57710.1 | Zm00001eb257430_T002 | Glyma.17G067700.1 | ||||||
| MtSMXL4 | MtSMXL14 | AT2G29970.1 | ESW14200 | Glyma.02G226900.6 | |||||
| MtSMXL5 | AT1G07200.2 | ESW14200 | Glyma.02G226900.6 | ESW18766 | Glyma.11G174500.4 | ||||
| AT2G29970.1 | ESW18766 | Glyma.11G174500.4 | Glyma.14G193900.2 | ||||||
| Glyma.14G193900.2 | Glyma.18G062300.1 | ||||||||
| Glyma.18G062300.1 | MtSMXL15 | ||||||||
| MtSMXL6 | AT2G40130.2 | ESW13755 | Glyma.11G230700.1 | MtSMXL16 | AT5G15450.1 | ESW11501 | Glyma.08G242100.1 | ||
| ESW18766 | Glyma.18G026600.3 | ESW23286 | Glyma.13G058600.1 | ||||||
| ESW35546 | Glyma.18G062300.1 | Glyma.18G264400.1 | |||||||
| MtSMXL7 | AT5G51070.1 | Zm00001eb084420_T004 | ESW09825 | Glyma.04G203300.1 | Glyma.19G027900.1 | ||||
| Glyma.06G162200.1 | MtSMXL17 | AT3G48870.3 | Zm00001eb084890_T001 | ESW09858 | Glyma.04G200400.8 | ||||
| MtSMXL8 | AT5G50920.1 | ESW31965 | Glyma.05G201100.4 | ||||||
| MtSMXL9 | Glyma.06G165200.6 | ||||||||
| MtSMXL10 | AT2G25140.1 | Zm00001eb234160_T001 | ESW08954 | Glyma.04G062200.1 | Glyma.08G008400.1 |
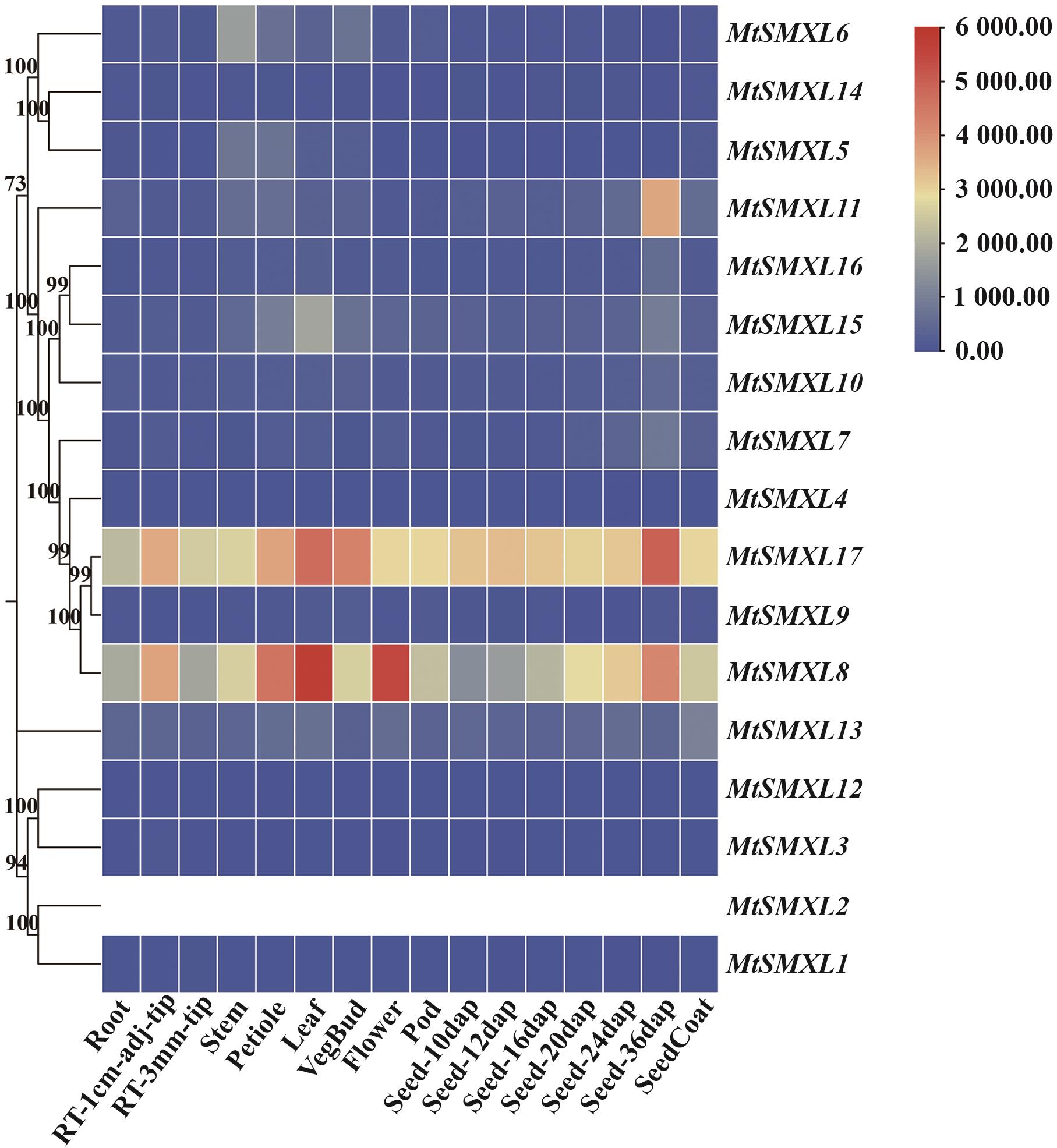
Fig. 8 Expression analysis of SMXL family genes indifferent tissues of M. truncatulaThe upper right corner shows the expression range of the MtSMXL gene in different tissues. Darker red indicates higher expression, and darker blue indicates lower expression. In the figure, adj refers to distance, and dap refers to days after pollination
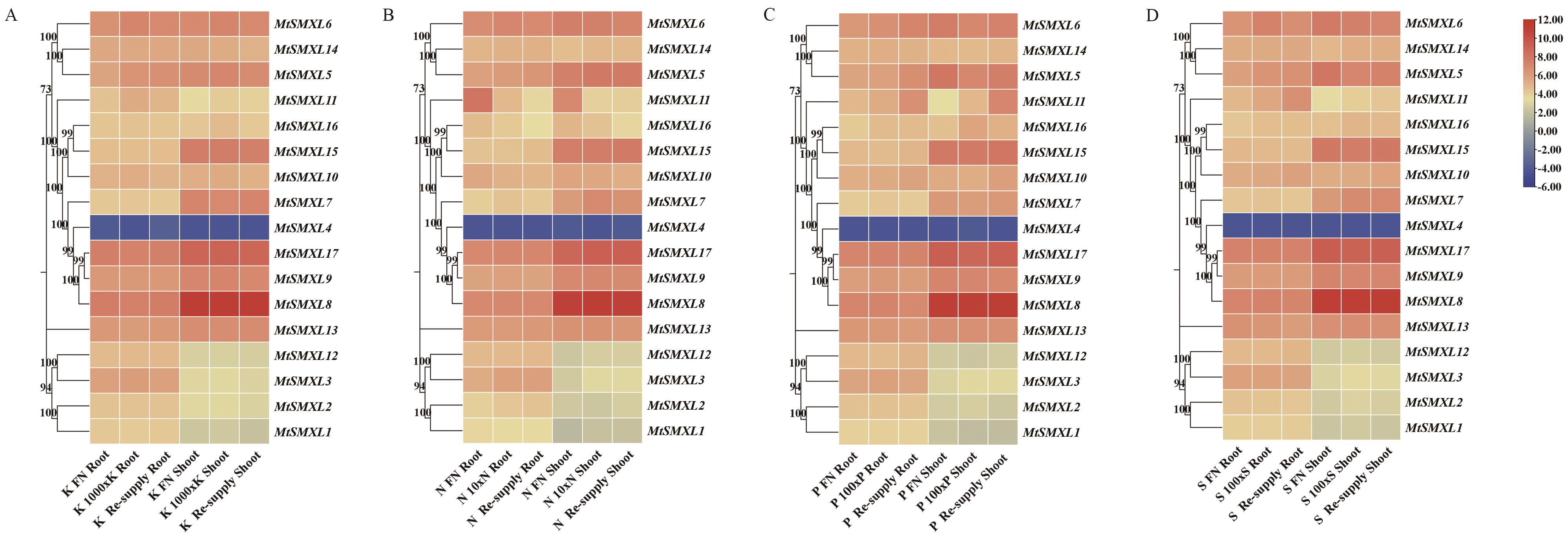
Fig. 9 Expression analysis of SMXL family genes of M. truncatula under conditions of N, P, K and S deficiencyA: Heatmap of the expressions of MtSMXLs genes in response to low potassium treatment in the roots and shoots. B: Heatmap of the expressions of MtSMXLs genes in response to low nitrogen treatment in the roots and shoots. C: Heatmap of the expressions of MtSMXLs genes in response to low phosphorus treatment in the roots and shoots. D: Heatmap of the expressions of MtSMXLs gene in response to low sulfur treatment in the roots and shoots
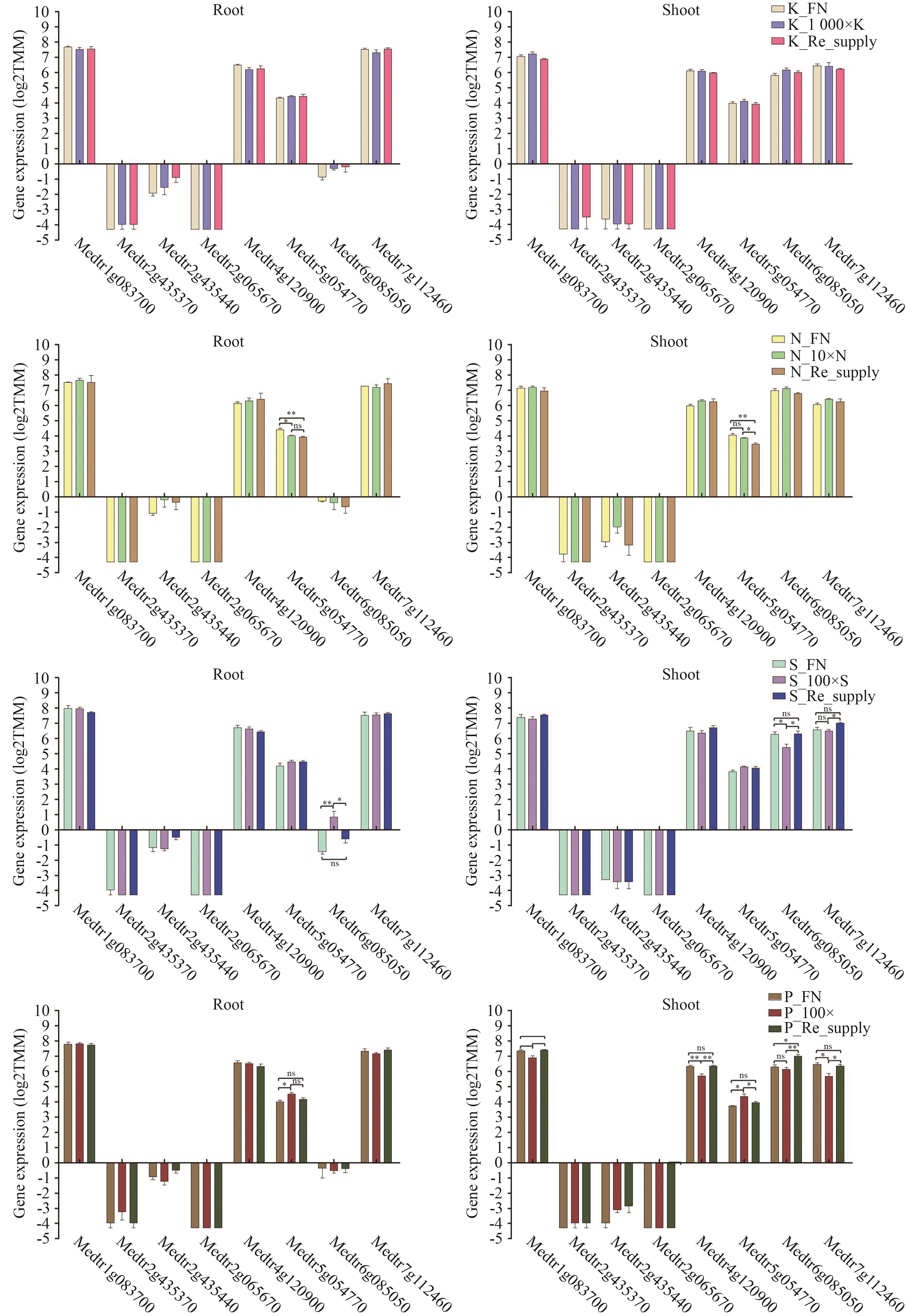
Fig. 10 Gene expression analysis of candidate genes encoding interacting proteins under N, P, K and S treatmentsBased on GraphPad Prism 5.0 analysis, *P<0.05, **P<0.005
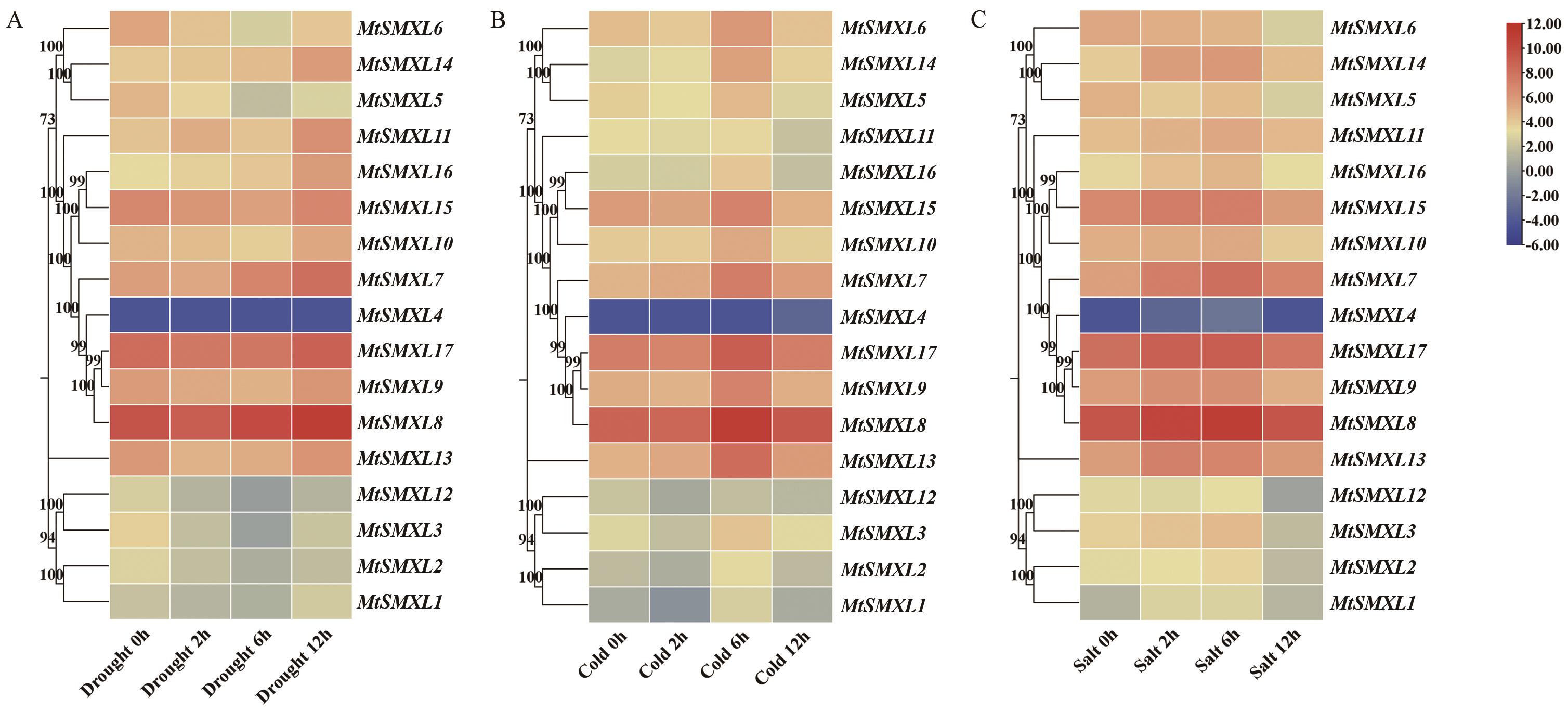
Fig. 12 Analysis of the gene expression patterns of SMXL family genes under drought, cold and salt treatmentA: Heatmap of the expression levels of MtSMXLs gene under drought treatment in seedlings. B: Heatmap of the expression levels of MtSMXLs gene under cold treatment in seedlings. C: Heatmap of the expression levels of MtSMXLs gene under salt treatment in seedlings
| [1] | Ma B, Zhu JB, Huang XZ. Diversification of plant SUPPRESSOR OF MAX2 1 (SMAX1)-like genes and genome-wide identification and characterization of cotton SMXL gene family [J]. BMC Plant Biol, 2023, 23(1): 419. |
| [2] | 卢昱帆, 闫怡秀, 秦颖杭, 等. 玉米SMXL基因家族的全基因组鉴定及在非生物胁迫中的潜在功能分析 [J/OL]. 分子植物育种, 2024: 1-19. |
| Lu YF, Yan YX, Qin YH, et al. Genome-wide identification and potential function analysis of maize SMXL gene family under abiotic stress [J/OL]. Mol Plant Breed, 2024: 1-19. | |
| [3] | Bhoi A, Yadu B, Chandra J, et al. Contribution of strigolactone in plant physiology, hormonal interaction and abiotic stresses [J]. Planta, 2021, 254(2): 28. |
| [4] | Wang L, Wang B, Yu H, et al. Transcriptional regulation of strigolactone signalling in Arabidopsis [J]. Nature, 2020, 583(7815): 277-281. |
| [5] | 王雪菱, 王如月, 李际红, 等. 独脚金内酯抑制因子D53/SMXLs基因的研究进展 [J]. 农学学报, 2023, 13(10): 37-43. |
| Wang XL, Wang RY, Li JH, et al. The strigolactones inhibitory factor D53/SMXLs: a review [J]. J Agric, 2023, 13(10): 37-43. | |
| [6] | Arite T, Umehara M, Ishikawa S, et al. d14, a strigolactone-insensitive mutant of rice, shows an accelerated outgrowth of tillers [J]. Plant Cell Physiol, 2009, 50(8): 1416-1424. |
| [7] | Jiang L, Liu X, Xiong GS, et al. DWARF 53 acts as a repressor of strigolactone signalling in rice [J]. Nature, 2013, 504(7480): 401-405. |
| [8] | Stanga JP, Smith SM, Briggs WR, et al. SUPPRESSOR OF MORE AXILLARY GROWTH2 1 controls seed germination and seedling development in Arabidopsis [J]. Plant Physiol, 2013, 163(1): 318-330. |
| [9] | Yao RF, Ming ZH, Yan LM, et al. DWARF14 is a non-canonical hormone receptor for strigolactone [J]. Nature, 2016, 536(7617): 469-473. |
| [10] | Wang L, Wang B, Jiang L, et al. Strigolactone signaling in Arabidopsis regulates shoot development by targeting D53-like SMXL repressor proteins for ubiquitination and degradation [J]. Plant Cell, 2015, 27(11): 3128-3142. |
| [11] | Wallner ES, López-Salmerón V, Belevich I, et al. Strigolactone- and karrikin-independent SMXL proteins are central regulators of phloem formation [J]. Curr Biol, 2017, 27(8): 1241-1247. |
| [12] | Zhou F, Lin QB, Zhu LH, et al. Correction: Corrigendum: D14-SCFD3-dependent degradation of D53 regulates strigolactone signalling [J]. Nature, 2016, 532(7599): 402. |
| [13] | Wu YY, Hou BH, Lee WC, et al. DCL2- and RDR6-dependent transitive silencing of SMXL4 and SMXL5 in Arabidopsis dcl4 mutants causes defective phloem transport and carbohydrate over-accumulation [J]. Plant J, 2017, 90(6): 1064-1078. |
| [14] | Stanga JP, Morffy N, Nelson DC. Functional redundancy in the control of seedling growth by the karrikin signaling pathway [J]. Planta, 2016, 243(6): 1397-1406. |
| [15] | Villaécija-Aguilar JA, Hamon-Josse M, Carbonnel S, et al. SMAX1/SMXL2 regulate root and root hair development downstream of KAI2-mediated signalling in Arabidopsis [J]. PLoS Genet, 2019, 15(8): e1008327. |
| [16] | Wang ZJ, Jiang ZN, Wan HP, et al. Genome-wide identification of the SMXL gene family in common wheat and expression analysis of TaSMXLs under abiotic stress [J]. Agronomy, 2025, 15(3): 656. |
| [17] | 贾婷婷, 竺丽萍, 肖光辉, 等. 棉花SMXL基因家族的全基因组分析及表达分析 [J]. 中国科学: 生命科学, 2022, 52(12): 1868-1882. |
| Jia TT, Zhu LP, Xiao GH, et al. Genome-wide identification and expression analysis of the SMXL gene family in cotton [J]. Sci Sin Vitae, 2022, 52(12): 1868-1882. | |
| [18] | Zhang H, Wang L, Gao Y, et al. Genome-wide identification of SMXL gene family in soybean and expression analysis of GmSMXLs under shade stress [J]. Plants, 2022, 11(18): 2410. |
| [19] | Fu XJ, Wang J, Shangguan TW, et al. SMXLs regulate seed germination under salinity and drought stress in soybean [J]. Plant Growth Regul, 2022, 96(3): 397-408. |
| [20] | 陈莹, 张春渝, 许小琼, 等. 4种无患子科植物的SMXL家族全基因组鉴定、进化及DlSMXLs在龙眼体胚发生过程中的表达分析 [J]. 东南园艺, 2024, 12(1): 17-29. |
| Chen Y, Zhang CY, Xu XQ, et al. Genome-wide identification and evolutionary analysis of the SMXL gene family in four Sapindaceae species and expression analysis of DlSMXLs during somatic embryogenesis in Longan [J]. Southeast Hortic, 2024, 12(1): 17-29. | |
| [21] | Li R, An JP, You CX, et al. Genome-wide analysis and identification of the SMXL gene family in apple (Malus×domestica) [J]. Tree Genet Genomes, 2018, 14(4): 61. |
| [22] | 孙卫健. 苹果独脚金内酯信号途径阻遏蛋白同源基因MdSMXL8.2的功能鉴定 [D]. 泰安: 山东农业大学, 2020. |
| Sun WJ. Functional identification of apple strigolactone signaling pathway repressor homologous gene MdSMXL8.2 [D]. Tai’an: Shandong Agricultural University, 2020. | |
| [23] | Basso MF, Contaldi F, Lo Celso F, et al. Identification and expression profile of the SMAX/SMXL family genes in chickpea and lentil provide important players of biotechnological interest involved in plant branching [J]. Planta, 2023, 259(1): 1. |
| [24] | Yuan S, Zhang WL, Zhang YX. Characterization of SUPPRESSOR OF MAX2 1-LIKE (SMXL) genes in ‘Duli’ (Pyrus betulifolia L.) and expression analysis of PbSMXLs in response to plant growth regulators and salt stress [J]. Agronomy, 2024, 14(12): 2778. |
| [25] | Fang PP, Li MX, Guo QW, et al. Genome-wide analysis of the SMXL gene family in common bean and identification of karrikin-responsive PvSMXL2 as a negative regulator of PEG-induced drought stress [J]. Gene, 2023, 887: 147741. |
| [26] | Sun MT, Wang DY, Liu CS, et al. Genome-wide identification and analysis of the SUPPRESSOR of MAX2 1-LIKE gene family and its interaction with DWARF14 in poplar [J]. BMC Plant Biol, 2023, 23(1): 105. |
| [27] | Moturu TR, Thula S, Singh RK, et al. Molecular evolution and diversification of the SMXL gene family [J]. J Exp Bot, 2018, 69(9): 2367-2378. |
| [28] | Yang TJ, Kim JS, Kwon SJ, et al. Sequence-level analysis of the diploidization process in the triplicated FLOWERING LOCUS C region of Brassica rapa [J]. Plant Cell, 2006, 18(6): 1339-1347. |
| [29] | de Bang TC, Lundquist PK, Dai XB, et al. Genome-wide identification of Medicago Peptides involved in macronutrient responses and nodulation [J]. Plant Physiol, 2017, 175(4): 1669-1689. |
| [30] | Hu B, Wu H, Huang WF, et al. SWEET gene family in Medicago truncatula: genome-wide identification, expression and substrate specificity analysis [J]. Plants, 2019, 8(9): 338. |
| [31] | Qiao X, Li QH, Yin H, et al. Gene duplication and evolution in recurring polyploidization-diploidization cycles in plants [J]. Genome Biol, 2019, 20(1): 38. |
| [32] | 贺红利, 朴京培, 孙嘉囡, 等. 蒺藜苜蓿CIPK基因家族全基因组鉴定及表达分析 [J]. 中国草地学报, 2021, 43(9): 1-13. |
| He HL, Piao JP, Sun JN, et al. Genome-wide identification and expression analysis of CIPK gene family in Medicago truncatula [J]. Chin J Grassland, 2021, 43(9): 1-13. | |
| [33] | Wolfe KH, Gouy M, Yang YW, et al. Date of the monocot-dicot divergence estimated from chloroplast DNA sequence data [J]. Proc Natl Acad Sci U S A, 1989, 86(16): 6201-6205. |
| [34] | Zhao YY, Zhang R, Jiang KW, et al. Nuclear phylotranscriptomics and phylogenomics support numerous polyploidization events and hypotheses for the evolution of rhizobial nitrogen-fixing symbiosis in Fabaceae [J]. Mol Plant, 2021, 14(5): 748-773. |
| [35] | Zhang L, Wu RL, Mur LAJ, et al. Assembly of high-quality genomes of the locoweed Oxytropis ochrocephala and its endophyte Alternaria oxytropis provides new evidence for their symbiotic relationship and swainsonine biosynthesis [J]. Mol Ecol Resour, 2023, 23(1): 253-272. |
| [36] | 张孝廉, 张吉顺, 雷波, 等. 植物MLO蛋白研究进展 [J]. 植物生理学报, 2018, 54(7): 1159-1171. |
| Zhang XL, Zhang JS, Lei B, et al. Research progress of plant MLO protein [J]. Plant Physiol Commun, 2018, 54(7): 1159-1171. | |
| [37] | Kim MY, Kang YJ, Lee T, et al. Divergence of flowering-related genes in three legume species [J]. Plant Genome, 2013, 6(3): plantgenome2013.03.0008. |
| [38] | Liang YY, Ward S, Li P, et al. SMAX1-LIKE7 signals from the nucleus to regulate shoot development in Arabidopsis via partially EAR motif-independent mechanisms [J]. Plant Cell, 2016, 28(7): 1581-1601. |
| [39] | Fang JJ, Guo TT, Xie ZW, et al. The URL1-ROC5-TPL2 transcriptional repressor complex represses the ACL1 gene to modulate leaf rolling in rice [J]. Plant Physiol, 2021, 185(4): 1722-1744. |
| [1] | GAO Fei, ZHANG Yu-xi, MU Di, CHEN Zheng, CHEN Hong-yan. Isolation, Identification, and Whole-genome Sequencing Analysis of Murine-Derived Limosilactobacillus reuteri [J]. Biotechnology Bulletin, 2026, 42(7): 1-13. |
| [2] | ZHANG Chi-hao, LIU Jin-nan, CHAO Yue-hui. Cloning and Functional Analysis of a bZIP Transcription Factor MtbZIP29 from Medicago truncatula [J]. Biotechnology Bulletin, 2026, 42(1): 241-250. |
| [3] | LIU Chang-ming, ZHANG Xin-yue, YANG Xin-meng, LIU Yang, LI Yue-han, WANG Ke-qing. Biological Characteristics and Genomic Analysis of a Novel Pathogen Strain Raoultella ornithinolytica Causing Soft Rot Disease in Konjac [J]. Biotechnology Bulletin, 2026, 42(1): 294-304. |
| [4] | LI Ya-tao, ZHANG Zhi-peng, ZHAO Meng-yao, LYU Zhen, GAN Tian, WEI Hao, WU Shu-feng, MA Yu-chao. Whole Genome Analysis of Bradyrhizobium sp. Bd1 and the Negative Regulating Function of TetR3 during Cell Growth and Nodulation [J]. Biotechnology Bulletin, 2025, 41(9): 289-301. |
| [5] | LYU Zhen, GAN Tian, HUO Si-yu, ZHAO Chen-di, ZHAO Meng-yao, LI Ya-tao, MA Yu-chao, GENG Yu-qing. Identification of Surfactin-producing Bacillus Velezensis C5A-1 and Evaluation of the Plant Growth-promoting Effects of Its Surfactin [J]. Biotechnology Bulletin, 2025, 41(9): 265-276. |
| [6] | CHENG Ting-ting, LIU Jun, WANG Li-li, LIAN Cong-long, WEI Wen-jun, GUO Hui, WU Yao-lin, YANG Jing-fan, LAN Jin-xu, CHEN Sui-qing. Genome-wide Identification of the Chalcone Isomerase Gene Family in Eucommia ulmoides and Analysis of Their Expression Patterns [J]. Biotechnology Bulletin, 2025, 41(9): 242-255. |
| [7] | WEI Yao, ZHANG Jing-jing, CUI Yun-xiao, LIU Yu, LIU Hai-rui. Phylogenetic Analysis of Lonicera Sect. Nintooa Based on Chloroplast Genomes Data [J]. Biotechnology Bulletin, 2025, 41(8): 276-288. |
| [8] | LI Kai-yue, DENG Xiao-xia, YIN Yuan, DU Ya-tong, XU Yuan-jing, WANG Jing-hong, YU Song, LIN Ji-xiang. Identification of LEA Gene Family and Analysis on Its Response to Aluminum Stress in Ricinus communis L. [J]. Biotechnology Bulletin, 2025, 41(7): 128-138. |
| [9] | GONG Yu-han, CHEN Lan, SHANGFANG Hui-zi, HAO Ling-ying, LIU Shuo-qian. Identification and Expression Profile Analysis of the TRB Gene Family in Tea Plant [J]. Biotechnology Bulletin, 2025, 41(7): 214-225. |
| [10] | AN Chang, XU Wen-bo, LU Lin, LI Deng-lin, YAO Yi-xin, LIN Yan-xiang, YANG Cheng-zi, QIN Yuan, ZHENG Ping. Comparative Analysis of Glehnia littoralis from Different Geographic Regions Based on the Characteristics of Chloroplast Genome [J]. Biotechnology Bulletin, 2025, 41(6): 229-242. |
| [11] | HUO Guan-zhong, ZHANG Xin-ru, TIAN Shi-jun, LI Jun. Current Progress and Applications of CRISPR/Cas12a Gene Editing Technology in Plants [J]. Biotechnology Bulletin, 2025, 41(6): 1-11. |
| [12] | LI Chen-ying, KONG Da-shuai, LI Ruo-nan, ZHANG Yu-bo, YAN ping, LI Kui, KONG Si-yuan. Cutting-edge Omics Technology Innovations Empower Livestock and Poultry Biological Breeding [J]. Biotechnology Bulletin, 2025, 41(6): 71-86. |
| [13] | ZHOU Yi, LIU Yong-bo. Research Progress in the Mechanisms and Functions of Gene Loss in Genome Evolution [J]. Biotechnology Bulletin, 2025, 41(6): 38-48. |
| [14] | ZHANG Hui, LU Wen-cai, WANG Dong, LIU Qian, MA Lian-jie. Identification of Bacillus cereus YT2-1C with High Indoleacetic Acid Yield and Its Growth-promoting Effect [J]. Biotechnology Bulletin, 2025, 41(5): 300-309. |
| [15] | WU Ze-yin, HUANG Chen-yang, ZHAO Meng-ran, ZHANG Li-jiao, YAO Fang-jie. Genome-specific Analysis of Pleurotus cornucopiae CCMSSC 04611 with Short Stipe [J]. Biotechnology Bulletin, 2025, 41(5): 320-332. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||