








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 239-249.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0844
Previous Articles Next Articles
ZHANG Chi-hao1( ), LIU Jin-nan1, DONG Jia-le2, CHAO Yue-hui1(
), LIU Jin-nan1, DONG Jia-le2, CHAO Yue-hui1( )
)
Received:2025-08-04
Online:2026-02-26
Published:2026-03-17
Contact:
CHAO Yue-hui
E-mail:zzzch0127@126.com;chaoyuehui@bjfu.edu.cn
ZHANG Chi-hao, LIU Jin-nan, DONG Jia-le, CHAO Yue-hui. Construction of Alfalfa Yeast Expression Library and Screening and Identification of Salt-tolerance Genes[J]. Biotechnology Bulletin, 2026, 42(2): 239-249.
| 引物名称 Primer name | 序列 Sequence (5′‒3′) | 用途 Usage |
|---|---|---|
| CDS III/3' PCR Primer | GACGATAAGGTACCTAAGGATCCACCGGG | 逆转录 Reverse transcription |
| SMART IV Oligonucleotide | GTGCTGGATATCTGCAGAATTCCATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTVN | |
| P1-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAGGG | ds cDNA扩增 ds cDNA amplification |
| P2-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAGGG | |
| P3-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAAGGG | |
| P4-R | ACTCGAGCGGCCGCCACTGTGCTGGATATCTGCAGAATTCCA | |
| pYES2-NTB-F | CAGCTGTAATACGACTCACTATAGG | 文库菌落PCR鉴定 PCR identification of library colonies |
| pYES2-NTB-R | AGGGTTAGGGATAGGCTTACCTTCG | |
| NtActin-F | AGGTGGAGACATGGGTGGTG | 烟草内参基因表达分析 Analysis of tobacco internal reference gene expression |
| NtActin-R | TCATTAGGCACACAGATCTCTG | |
| MsG5270-qRT-F | CGGCTTGCTAACTACCGTTCT | 基因表达分析 Gene expression analysis |
| MsG5270-qRT-R | TTGATTTGATCTCGTTGAGGC | |
| MsG0180-qRT-F | GCTGATGAAATCGTCCCA | |
| MsG0180-qRT-R | CAACCCTGTTCCTCCCAC | |
| MsG9320-qRT-F | CTCTTGGCTATGGTTCTTC | |
| MsG9320-qRT-R | TTGGCATCGTTTACTTCA | |
| MsG9120-qRT-F | TCCAATGTTCGTCCCACT | |
| MsG9120-qRT-R | CTCTTTCCCATCTTCGTG | |
| MsG5270-F | TCACCATTTACGAACGATACATGTTACCAAACCCAACAAC | 克隆及表达载体构建 Construction of cloning and expression vectors |
| MsG5270-R | ATGGTGATGCATTCCCGGGTTTATCGTTGATTTGATCTCG | |
| MsG9320-F | TCACCATTTACGAACGATACATGGATTCGAGAAAAGCAAT | |
| MsG9320-R | ATGGTGATGCATTCCCGGGTTCAATTCTGAGTGTTGTCCT | |
| MsG9120-F | TCACCATTTACGAACGATACATGGAAGTTCTTTCTGCTAC | |
| MsG9120-R | ATGGTGATGCATTCCCGGGTCTACCTGGGCTGATGAATAG | |
| MsG0180-F | TCACCATTTACGAACGATACATGGGTGTTTTTACTTTCAA | |
| MsG0180-R | ATGGTGATGCATTCCCGGGTCTAGTAATTAGGGTTTGCTA | |
| 3302Y-F | TGACGCACAATCCCACTATCCTT | 植物表达载体PCR鉴定 PCR identification of plant expression vector |
| NOS181-R | CGTATTAAATGTATAATTGCGGGAC |
Table 1 Primers and sequences
| 引物名称 Primer name | 序列 Sequence (5′‒3′) | 用途 Usage |
|---|---|---|
| CDS III/3' PCR Primer | GACGATAAGGTACCTAAGGATCCACCGGG | 逆转录 Reverse transcription |
| SMART IV Oligonucleotide | GTGCTGGATATCTGCAGAATTCCATTTTTTTTTTTTTTTTTTTTTTTTTTTTTTVN | |
| P1-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAGGG | ds cDNA扩增 ds cDNA amplification |
| P2-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAGGG | |
| P3-F | GATCTGTACGACGATGACGATAAGGTACCTAAGGATCCAAAGGG | |
| P4-R | ACTCGAGCGGCCGCCACTGTGCTGGATATCTGCAGAATTCCA | |
| pYES2-NTB-F | CAGCTGTAATACGACTCACTATAGG | 文库菌落PCR鉴定 PCR identification of library colonies |
| pYES2-NTB-R | AGGGTTAGGGATAGGCTTACCTTCG | |
| NtActin-F | AGGTGGAGACATGGGTGGTG | 烟草内参基因表达分析 Analysis of tobacco internal reference gene expression |
| NtActin-R | TCATTAGGCACACAGATCTCTG | |
| MsG5270-qRT-F | CGGCTTGCTAACTACCGTTCT | 基因表达分析 Gene expression analysis |
| MsG5270-qRT-R | TTGATTTGATCTCGTTGAGGC | |
| MsG0180-qRT-F | GCTGATGAAATCGTCCCA | |
| MsG0180-qRT-R | CAACCCTGTTCCTCCCAC | |
| MsG9320-qRT-F | CTCTTGGCTATGGTTCTTC | |
| MsG9320-qRT-R | TTGGCATCGTTTACTTCA | |
| MsG9120-qRT-F | TCCAATGTTCGTCCCACT | |
| MsG9120-qRT-R | CTCTTTCCCATCTTCGTG | |
| MsG5270-F | TCACCATTTACGAACGATACATGTTACCAAACCCAACAAC | 克隆及表达载体构建 Construction of cloning and expression vectors |
| MsG5270-R | ATGGTGATGCATTCCCGGGTTTATCGTTGATTTGATCTCG | |
| MsG9320-F | TCACCATTTACGAACGATACATGGATTCGAGAAAAGCAAT | |
| MsG9320-R | ATGGTGATGCATTCCCGGGTTCAATTCTGAGTGTTGTCCT | |
| MsG9120-F | TCACCATTTACGAACGATACATGGAAGTTCTTTCTGCTAC | |
| MsG9120-R | ATGGTGATGCATTCCCGGGTCTACCTGGGCTGATGAATAG | |
| MsG0180-F | TCACCATTTACGAACGATACATGGGTGTTTTTACTTTCAA | |
| MsG0180-R | ATGGTGATGCATTCCCGGGTCTAGTAATTAGGGTTTGCTA | |
| 3302Y-F | TGACGCACAATCCCACTATCCTT | 植物表达载体PCR鉴定 PCR identification of plant expression vector |
| NOS181-R | CGTATTAAATGTATAATTGCGGGAC |
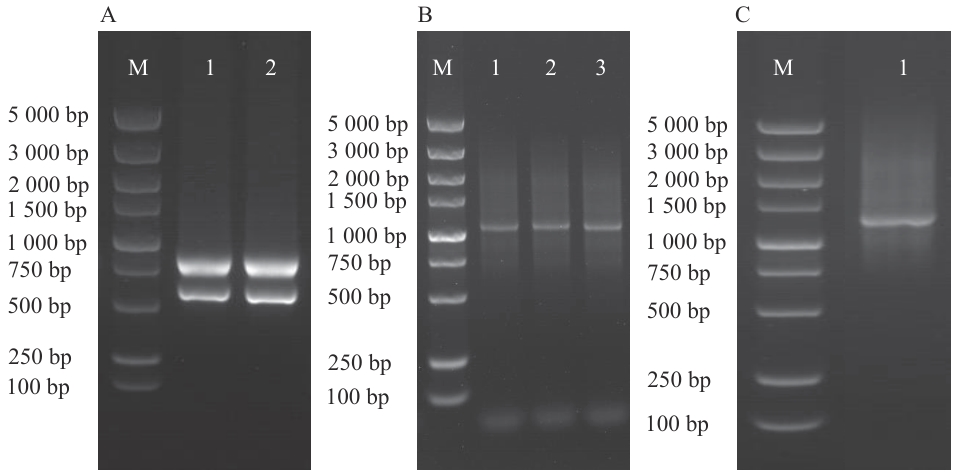
Fig. 1 RNA extraction and identification of synthesized ds cDNAA: Sample RNA extraction (M: Marker, 1‒2: alfalfa (Medicago sativa) samples). B: Amplified results of synthesized ds cDNA (M: Marker; 1: P1-F/P4-R amplified ds cDNA; 2: P2-F/P4-R amplified ds cDNA; 3: P3-F/P4-R amplified ds cDNA). C: Purified and removed small fragments (M: Marker; 1: results after the three ds cDNAs were mixed and purified)
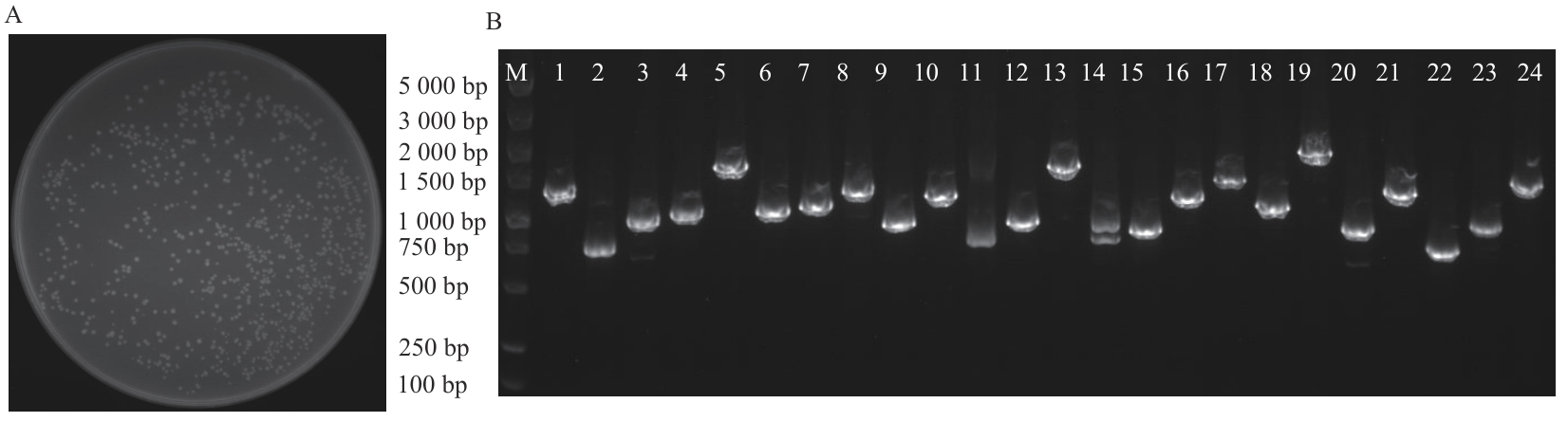
Fig. 2 Count and library identification of Escherichia coli colonyA: Library capacity and clone count. B: PCR identification of library quality (M: DNA marker; 1‒24: 24 different cloned colonies)
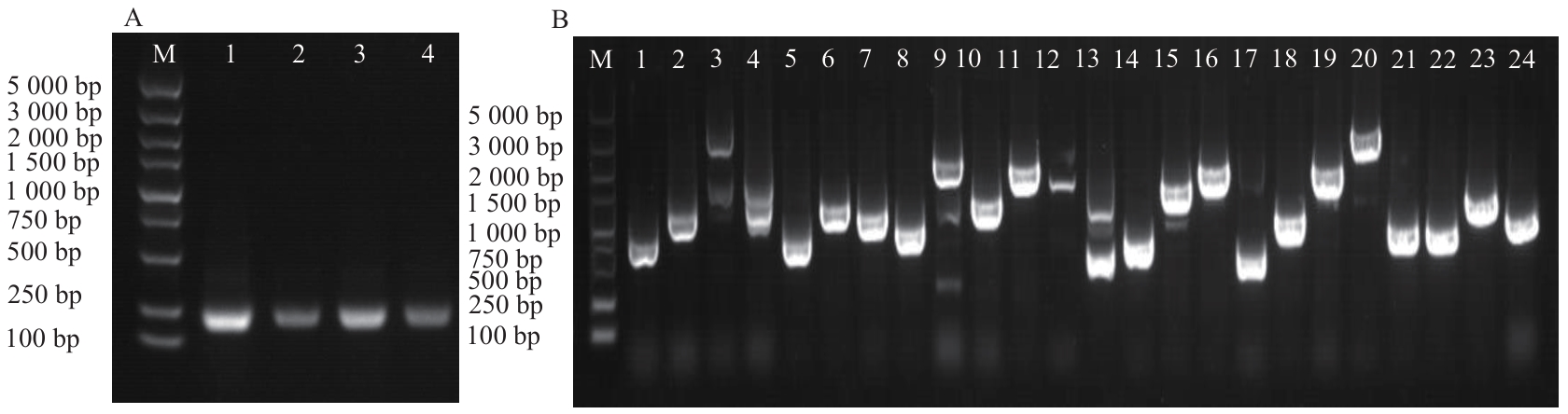
Fig. 3 PCR identification of yeast transformed with empty vector (A) and library cultures (B)In Fig. A, M: marker, 1-4: yeast transformation empty vector. In Fig. B, M: DNA marker, 1-24: yeast library colonies
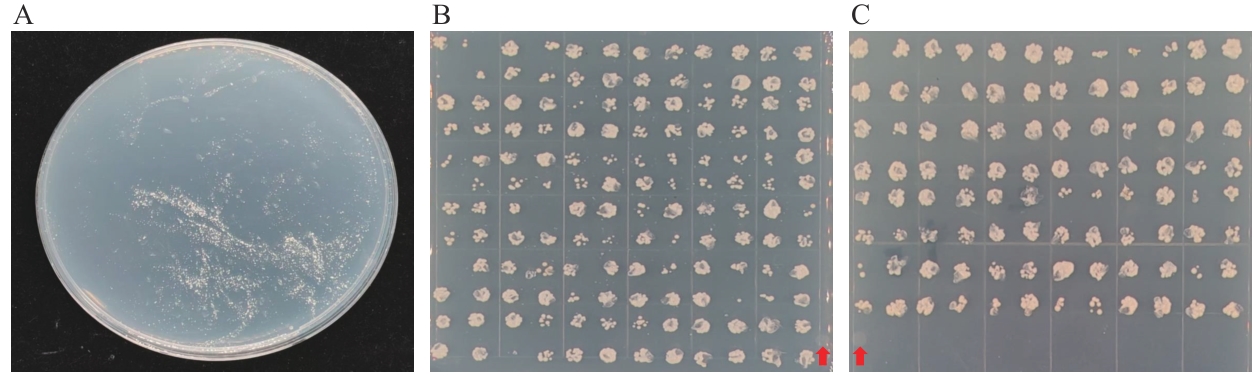

Fig. 5 Positive clone plate and results of rotation verificationA: The result of applying the library bacterial solution on 1.5 mol/L NaCl solid culture medium. B: Verification results of monoclonal colony rotation (144). C: Verification results of monoclonal colony rotation (96). : Control group
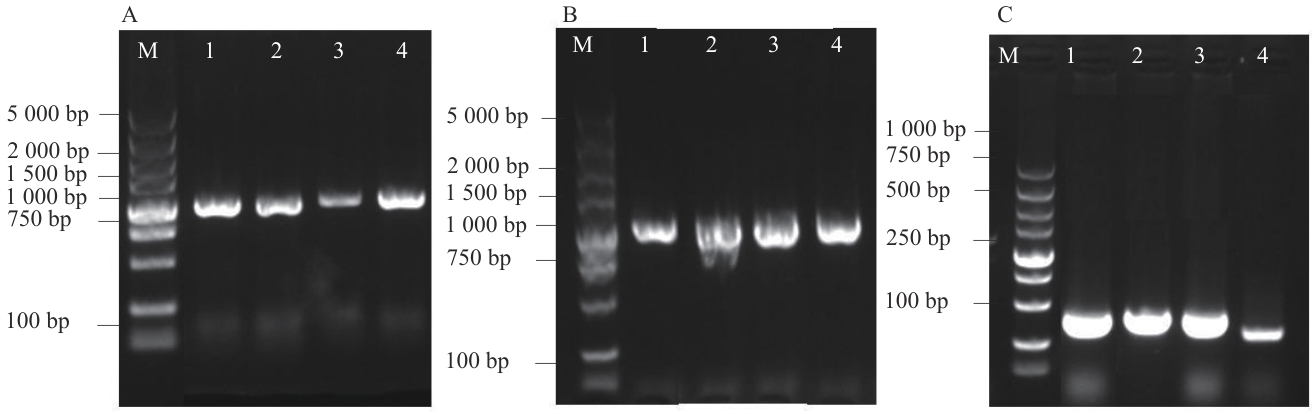
Fig. 8 Detection vector colony PCRA: Vector construction colony PCR (1: pBⅠ-MsG5270; 2: pBⅠ-MsG0180; 3: pBⅠ-MsG9320; 4: pBⅠ-MsG9120). B: Agrobacterium transformed colony PCR (1: pBⅠ-MsG5270; 2: pBⅠ- MsG0180; 3: pBⅠ-MsG9320; 4: pBⅠ-MsG9120). C: cDNA PCR identification (1: pBⅠ-MsG5270; 2: pBⅠ-MsG0180; 3: pBⅠ-MsG9320; 4: pBⅠ-MsG9120)
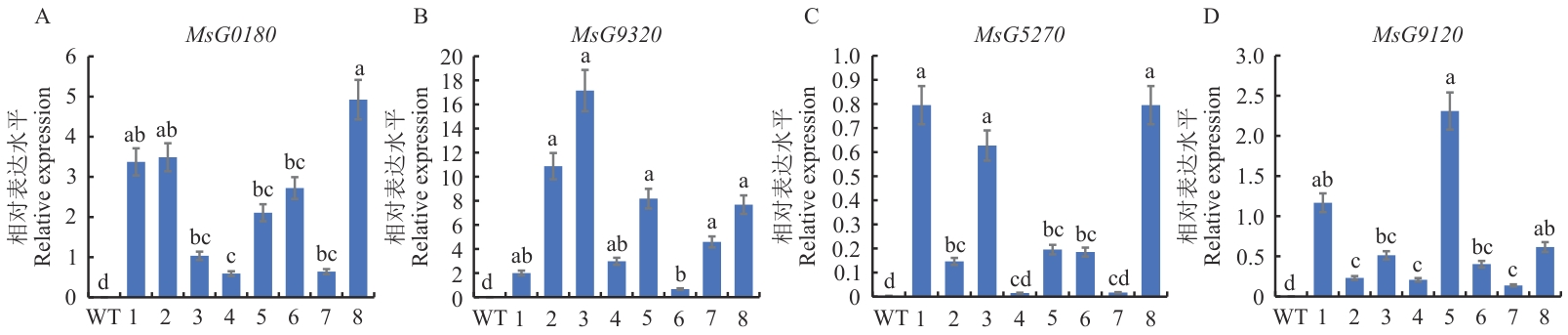
Fig. 9 Expression analysis of MsG0180, MsG9320, MsG5270, and MsG9120Different letters indicate significant differences at P<0.05 level. The data are in means ±SE (n=3), WT: Wild type; 1‒8: transgenic plants
| [1] | Gong ZZ. Plant abiotic stress: New insights into the factors that activate and modulate plant responses [J]. J Integr Plant Biol, 2021, 63(3): 429-430. |
| [2] | Joshi S, Nath J, Singh AK, et al. Ion transporters and their regulatory signal transduction mechanisms for salinity tolerance in plants [J]. Physiol Plant, 2022, 174(3): e13702. |
| [3] | Liang XY, Li JF, Yang YQ, et al. Designing salt stress-resilient crops: Current progress and future challenges [J]. J Integr Plant Biol, 2024, 66(3): 303-329. |
| [4] | Ali Aazami M, Rasouli F, Ebrahimzadeh A. Oxidative damage, antioxidant mechanism and gene expression in tomato responding to salinity stress under in vitro conditions and application of iron and zinc oxide nanoparticles on callus induction and plant regeneration [J]. BMC Plant Biol, 2021, 21(1): 597. |
| [5] | van Zelm E, Zhang YX, Testerink C. Salt tolerance mechanisms of plants [J]. Annu Rev Plant Biol, 2020, 71: 403-433. |
| [6] | Soltabayeva A, Ongaltay A, Omondi JO, et al. Morphological, physiological and molecular markers for salt-stressed plants [J]. Plants, 2021, 10(2): 243. |
| [7] | Prăvălie R, Patriche C, Borrelli P, et al. Arable lands under the pressure of multiple land degradation processes. A global perspective [J]. Environ Res, 2021, 194: 110697. |
| [8] | Popova LG, Khramov DE, Nedelyaeva OI, et al. Yeast heterologous expression systems for the study of plant membrane proteins [J]. Int J Mol Sci, 2023, 24(13): 10768. |
| [9] | Du LY, Huang XL, Ding L, et al. TaERF87 and TaAKS1 synergistically regulate TaP5CS1/TaP5CR1-mediated proline biosynthesis to enhance drought tolerance in wheat [J]. New Phytol, 2023, 237(1): 232-250. |
| [10] | Kumari S, Singh P, Singla-Pareek SL, et al. Heterologous expression of a salinity and developmentally regulated rice cyclophilin gene (OsCyp2) in E. coli and S. cerevisiae confers tolerance towards multiple abiotic stresses [J]. Mol Biotechnol, 2009, 42(2): 195-204. |
| [11] | Kumar R, Mustafiz A, Sahoo KK, et al. Functional screening of cDNA library from a salt tolerant rice genotype Pokkali identifies mannose-1-phosphate guanyl transferase gene (OsMPG1) as a key member of salinity stress response [J]. Plant Mol Biol, 2012, 79(6): 555-568. |
| [12] | Lee JH, Van Montagu M, Verbruggen N. A highly conserved kinase is an essential component for stress tolerance in yeast and plant cells [J]. Proc Natl Acad Sci USA, 1999, 96(10): 5873-5877. |
| [13] | Abulfaraj AA, Alshareef SA. Concordant gene expression and alternative splicing regulation under abiotic stresses in Arabidopsis [J]. Genes, 2024, 15(6): 675. |
| [14] | Yamanaka T, Miyama M, Tada Y. Transcriptome profiling of the mangrove plant Bruguiera gymnorhiza and identification of salt tolerance genes by Agrobacterium functional screening [J]. Biosci Biotechnol Biochem, 2009, 73(2): 304-310. |
| [15] | Song A, Lu J, Jiang J, et al. Isolation and characterisation of Chrysanthemum crassum SOS1, encoding a putative plasma membrane Na+/H+ antiporter [J]. Plant Biol, 2012, 14(5): 706-713. |
| [16] | Kappachery S, Yu JW, Baniekal-Hiremath G, et al. Rapid identification of potential drought tolerance genes from Solanum tuberosum by using a yeast functional screening method [J]. Comptes Rendus Biol, 2013, 336(11/12): 530-545. |
| [17] | Eswaran N, Parameswaran S, Sathram B, et al. Yeast functional screen to identify genetic determinants capable of conferring abiotic stress tolerance in Jatropha curcas . [J]. BMC Biotechnol, 2010, 10: 23. |
| [18] | Obata T, Kitamoto HK, NakamUra A, et al. Rice shaker potassium channel OsKAT1 confers tolerance to salinity stress on yeast and rice cells [J]. Plant Physiol, 2007, 144(4): 1978-1985. |
| [19] | Burezq H. Reproductive ecology of forage alfalfa (Medicago sativa L.): recent advances [M]//Plant Reproductive Ecology-Recent Advances. London: IntechOpen, 2022. |
| [20] | 王馨磊, 朱思琪, 田晓娜, 等. 紫花苜蓿RALF基因家族的全基因组鉴定及盐胁迫下表达分析 [J]. 江苏农业学报, 2025, 41(7): 1260-1269. |
| Wang XL, Zhu SQ, Tian XN, et al. Genome-wide identification of the RALF gene family in Medicago sativa and expression analysis under salt stress [J]. Jiangsu J Agric Sci, 2025, 41(7): 1260-1269. | |
| [21] | 杭嘉慧. 紫花苜蓿褪黑素合成基因MsSNAT1和MsASMT1调控耐盐性机制研究 [D]. 银川: 宁夏大学, 2025. |
| Hang JH. Regulation mechanism of melatonin synthesis genes MsSNAT1 and MsASMT1 in salt tolerance of alfalfa [D]. Yinchuan: Ningxia University, 2025. | |
| [22] | 沙吾列·沙比汗, 赛里克·都曼, 张丽萍, 等. 20份苜蓿种质材料苗期耐盐性综合评价 [J]. 现代农业科技, 2016(8): 247-250, 262. |
| Shawulie·S, Sailike·D, Zhang LP, et al. A comprehensive evaluation on salt tolerance of 20 alfalfa germplasm materials in seedling stage [J]. XianDai NongYe KeJi, 2016(8): 247-250, 262. | |
| [23] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method [J]. Methods, 2001, 25(4): 402-408. |
| [24] | Li YK, Jiang FL, Niu LF, et al. Synergistic regulation at physiological, transcriptional and metabolic levels in tomato plants subjected to a combination of salt and heat stress [J]. Plant J, 2024, 117(6): 1656-1675. |
| [25] | Afzal M, El Sayed Hindawi S, Alghamdi SS, et al. Potential breeding strategies for improving salt tolerance in crop plants [J]. J Plant Growth Regul, 2023, 42(6): 3365-3387. |
| [26] | Ashraf M, Munns R. Evolution of approaches to increase the salt tolerance of crops [J]. Crit Rev Plant Sci, 2022, 41(2): 128-160. |
| [27] | Rasheed A, Raza A, Jie HD, et al. Molecular tools and their applications in developing salt-tolerant soybean (Glycine max L.) cultivars [J]. Bioengineering, 2022, 9(10): 495. |
| [28] | Fan XR, Jiang HZ, Meng LJ, et al. Gene mapping, cloning and association analysis for salt tolerance in rice [J]. Int J Mol Sci, 2021, 22(21): 11674. |
| [29] | Kausar R, Komatsu S. Proteomic approaches to uncover salt stress response mechanisms in crops [J]. Int J Mol Sci, 2022, 24(1): 518. |
| [30] | Atta K, Mondal S, Gorai S, et al. Impacts of salinity stress on crop plants: improving salt tolerance through genetic and molecular dissection [J]. Front Plant Sci, 2023, 14: 1241736. |
| [31] | Fidler J, Graska J, Gietler M, et al. PYR/PYL/RCAR receptors play a vital role in the abscisic-acid-dependent responses of plants to external or internal stimuli [J]. Cells, 2022, 11(8): 1352. |
| [32] | Fan WQ, Zhao MY, Li SX, et al. Contrasting transcriptional responses of PYR1/PYL/RCAR ABA receptors to ABA or dehydration stress between maize seedling leaves and roots [J]. BMC Plant Biol, 2016, 16: 99. |
| [33] | Zhao Y, Chan ZL, Gao JH, et al. ABA receptor PYL9 promotes drought resistance and leaf senescence [J]. Proc Natl Acad Sci USA, 2016, 113(7): 1949-1954. |
| [34] | Krishnaswamy SS, Srivastava S, Mohammadi M, et al. Transcriptional profiling of pea ABR17 mediated changes in gene expression in Arabidopsis thaliana . [J]. BMC Plant Biol, 2008, 8: 91. |
| [35] | Luo M, Liu JH, Mohapatra S, et al. Characterization of a gene family encoding abscisic acid- and environmental stress-inducible proteins of alfalfa [J]. J Biol Chem, 1992, 267(22): 15367-15374. |
| [36] | Wang XS, Huo ZC, Ma L, et al. The salt and ABA inducible transcription factor gene, SlAITR3, negatively regulates tomato salt tolerance [J]. Plant Physiol Biochem, 2025, 222: 109735. |
| [37] | 刘颖慧, 王天宇, 黎裕. 拟南芥脱水诱导早期应答基因研究进展 [J]. 中国农业大学学报, 2009, 14(3): 7-11. |
| Liu YH, Wang TY, Li Y. Review of early responsive genes expressed during dehydration in Arabidopsis sp [J]. J China Agric Univ, 2009, 14(3): 7-11. | |
| [38] | Kariola T, Brader G, Helenius E, et al. EARLY RESPONSIVE TO DEHYDRATION 15, a negative regulator of abscisic acid responses in Arabidopsis [J]. Plant Physiol, 2006, 142(4): 1559-1573. |
| [1] | LIU Qing-yuan, WU Hong-qi, CHEN Xiu-e, CHEN Jian, JIANG Yuan-ze, HE Yan-zi, YU Qi-wei, LIU Ren-xiang. Function of Transcription Factor NtMYB96a in Regulating the Tolerance of Tobacco to Drought [J]. Biotechnology Bulletin, 2026, 42(4): 239-250. |
| [2] | LIU Meng-ru, XIA Lin, LIU Rui-xia, YAN Xin-ke, YANG Jun, ZHANG Jing-hua, WU Ming-zhu. Cloning and Functional Analysis of Tobacco Polyphenol Oxidase NtPPO1-2 Gene [J]. Biotechnology Bulletin, 2026, 42(2): 218-227. |
| [3] | ZHANG Chi-hao, LIU Jin-nan, CHAO Yue-hui. Cloning and Functional Analysis of a bZIP Transcription Factor MtbZIP29 from Medicago truncatula [J]. Biotechnology Bulletin, 2026, 42(1): 241-250. |
| [4] | ZHAI Ying, JI Jun-jie, CHEN Jiong-xin, YU Hai-wei, LI Shan-shan, ZHAO Yan, MA Tian-yi. Heterologous Overexpression of Soybean GmNF-YB24 Improves the Resistance of Transgenic Tobacco to Drought [J]. Biotechnology Bulletin, 2025, 41(8): 137-145. |
| [5] | HUANG Xu-sheng, ZHOU Ya-li, CHAI Xu-dong, WEN Jing, WANG Ji-ping, JIA Xiao-yun, LI Run-zhi. Cloning of Plastidial PfLPAT1B Gene from Perilla frutescens and Its Functional Analysis in Oil Biosynthesis [J]. Biotechnology Bulletin, 2025, 41(7): 226-236. |
| [6] | DONG Xu-kun, CHE Yong-mei, WANG Ming-shuo, LUO Zheng-gang, GUAN En-sen, ZHAO Fang-gui, YE Qing, LIU Xin. Effects of Co-treatment of Nano-silica and Bacillus cereus SS1 on the Growth of Tobacco [J]. Biotechnology Bulletin, 2025, 41(7): 292-298. |
| [7] | WANG Wei-wei, ZHAO Zhen-jie, WANG Zhi, ZOU Jing-wei, LUO Zheng-hui, ZHANG Yu-jie, NIU Li-ya, YU Liang, YANG Xue-ju. Research Progress in Salt-tolerant Genes Related to Physiological Response of Wheat to Salt Stress [J]. Biotechnology Bulletin, 2025, 41(5): 14-22. |
| [8] | LU Yong-jie, XIA Hai-qian, LI Yong-ling, ZHANG Wen-jian, YU Jing, ZHAO Hui-na, WANG Bing, XU Ben-bo, LEI Bo. Cloning and Expression Analysis of AP2/ERF Transcription Factor NtESR2 in Nicotiana tabacum [J]. Biotechnology Bulletin, 2025, 41(4): 266-277. |
| [9] | JI Ying-tong, GAO Jia-ning, QIAN Meng-ying, PAN Ning, ZHANG Jun, CUI Hong, YAN Xiao-xiao. Function Study of NtMYC2 Gene on Tobacco Trichome Development [J]. Biotechnology Bulletin, 2025, 41(4): 278-288. |
| [10] | ZHAO Chun-duo, LI Yu-e, LIU You-jie, WANG Xin-hang, ZHAO Wei, HUANG Yong-cheng, LI Hu-lin, JI Wen-xiu. Effects of Rotating Cropping and Continuous Cropping on Soil Nutrients, Enzyme Activities and Microbial Community Structure of Rhizosphere Soil in Tobacco [J]. Biotechnology Bulletin, 2025, 41(4): 312-322. |
| [11] | MA Yao-wu, ZHANG Qi-yu, YANG Miao, JIANG Cheng, ZHANG Zhen-yu, ZHANG Yi-lin, LI Meng-sha, XU Jia-yang, ZHANG Bin, CUI Guang-zhou, JIANG Ying. Screening, Indentification and Promotion Performance Investigation of Tobacco Growth-promoting Rhizobacteria [J]. Biotechnology Bulletin, 2025, 41(3): 271-281. |
| [12] | HAN Meng-qiao, WU Jiang, LI Li-hua, WANG Zhao-yi, DENG Xi, WEI Feng-jie, REN Min, SUN Yang-yang, LI Fu-xin. Establishment and Optimization of a Tobacco Chromosome Preparation System [J]. Biotechnology Bulletin, 2025, 41(3): 44-50. |
| [13] | XIANG Bo-ka, ZHOU Zuan-zuan, FENG Jia-hui, XIA Chen, LI Qi, CHEN Chun. Isolation and Identification of a Fungus from Moldy Tobacco Leaf and Study on Its Mold-causing Factors [J]. Biotechnology Bulletin, 2025, 41(2): 321-330. |
| [14] | FENG Xiao-hu, ZHANG Wen-mei, XU Jing, XIONG Shu-bin, WANG Li-bing, SONG Wen-jing. Screening and Biocontrol Efficiency of Phage Cocktail Targeted on a Group of Tobacco Bacterial Wilt [J]. Biotechnology Bulletin, 2025, 41(11): 311-318. |
| [15] | WANG Bi-cheng, JING Hai-qing, WAN Kun, ZHANG Ying-ying, DING Jia-hao, LI Run-zhi, XUE Jin-ai, ZHANG Hai-ping. Identification of Soybean BCAT Gene Family and Functional Analysis of GmBCAT3 in Soybean Responses to Drought Stress [J]. Biotechnology Bulletin, 2025, 41(10): 196-209. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||