








生物技术通报 ›› 2023, Vol. 39 ›› Issue (3): 152-162.doi: 10.13560/j.cnki.biotech.bull.1985.2022-0974
谢洋1,2( ), 邢雨蒙1,2, 周国彦1,2, 刘美妍1,2, 银珊珊1,2, 闫立英1,2
), 邢雨蒙1,2, 周国彦1,2, 刘美妍1,2, 银珊珊1,2, 闫立英1,2
收稿日期:2022-08-09
出版日期:2023-03-26
发布日期:2023-04-10
通讯作者:
谢洋作者简介:谢洋,女,博士,讲师,研究方向:蔬菜栽培生理及分子生物学;E-mail:xieyangly123@163.com
基金资助:
XIE Yang1,2( ), XING Yu-meng1,2, ZHOU Guo-yan1,2, LIU Mei-yan1,2, YIN Shan-shan1,2, YAN Li-ying1,2
), XING Yu-meng1,2, ZHOU Guo-yan1,2, LIU Mei-yan1,2, YIN Shan-shan1,2, YAN Li-ying1,2
Received:2022-08-09
Published:2023-03-26
Online:2023-04-10
摘要:
果形和糖含量分别是与瓜类蔬菜作物外观、营养及风味密切相关的重要品质性状。为探究黄瓜倍性差异导致果形及糖含量变化的分子机制,以果形和糖含量差异显著的黄瓜二倍体(长果/低糖)和四倍体(短果/高糖)为试材,分别取开花当天和花后9 d的果实进行转录组分析。结果表明,在二、四倍体花后9 d果实中共鉴定到1 874个差异表达基因(DEGs),其中上调基因818个和下调基因1 056个;GO分析显示,DEGs显著富集在“DNA复制”(GO:0006260)、“DNA代谢过程”(GO:0006259)、“蛋白质复合体”(GO:0032993)、“染色体”(GO:0005694)、“蛋白质二聚活性”(GO:0046983)等GO条目上;KEGG分析表明,DEGs富集在“植物激素信号转导”(csv04075)、“淀粉与蔗糖代谢”(csv00500)等代谢通路上;参与“植物激素信号转导”相关基因AUX1、DELLA、B-ARR、TCH4上调表达,GH3、GID1、PP2C、CYCD3下调表达,以及参与“淀粉与蔗糖代谢”相关基因SPS、SUSY上调表达,可能与果形指数减小和果实糖含量增加有关。荧光定量PCR(RT-qPCR)分析显示,除ARF4外,所选定DEGs的表达模式与RNA-Seq中表达趋势相一致,其中SUSY在二、四倍体黄瓜果实中差异表达极显著,推测该基因在调控黄瓜四倍体果形及糖含量形成过程中起到重要作用。
谢洋, 邢雨蒙, 周国彦, 刘美妍, 银珊珊, 闫立英. 黄瓜二倍体及其同源四倍体果实转录组分析[J]. 生物技术通报, 2023, 39(3): 152-162.
XIE Yang, XING Yu-meng, ZHOU Guo-yan, LIU Mei-yan, YIN Shan-shan, YAN Li-ying. Transcriptome Analysis of Diploid and Autotetraploid in Cucumber Fruit[J]. Biotechnology Bulletin, 2023, 39(3): 152-162.
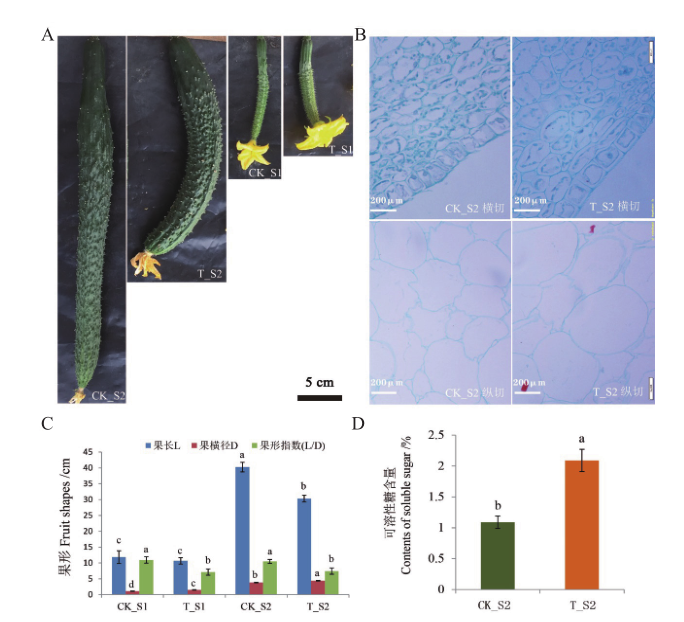
图1 黄瓜二倍体及其同源四倍体果形及可溶性糖含量分析 A:黄瓜二、四倍体不同发育时期果实形态学观察;B:黄瓜二、四倍体不同发育时期果实细胞学观察;C:黄瓜二、四倍体不同发育时期果实果形指标统计;D:黄瓜二、四倍体果实可溶性糖含量分析。不同小写字母表示在0.05水平差异显著
Fig. 1 Analysis of diploid and autotetraploid fruit shapes as well as their soluble sugar contents in cucumber A: Morphological observation of cucumber fruits at different developmental stages of diploid and tetraploid. B: Fruit cytology of cucumber diploid and tetraploid at different developmental stages. C: Statistics of fruit shape index of cucumber diploid and tetraploid at different developmental stages. D: Analysis of soluble sugar content in diploid and tetraploid fruits of cucumber. Different lower letters indicate significant difference at 0.05 level
| 样品名称 Sample name | Raw reads/Mb | Clean reads/Mb | Unique map/Mb | Q30/% | GC/% |
|---|---|---|---|---|---|
| CK_S1_1 | 45.32 | 43.78 | 40.87 | 92.94 | 40.88 |
| CK_S1_2 | 48.02 | 45.29 | 43.08 | 94.72 | 40.85 |
| CK_S2_1 | 44.23 | 43.35 | 40.80 | 93.02 | 42.14 |
| CK_S2_2 | 44.89 | 43.22 | 40.68 | 92.63 | 42.28 |
| T_S1_1 | 46.21 | 43.69 | 40.35 | 92.56 | 40.82 |
| T_S1_2 | 47.89 | 45.85 | 42.61 | 92.98 | 41.09 |
| T_S2_1 | 45.83 | 44.53 | 42.08 | 92.9 | 42.43 |
| T_S2_2 | 47.69 | 45.88 | 43.46 | 93.49 | 42.43 |
表1 转录组测序数据质量分析
Table 1 Quality analysis of transcriptome sequencing data
| 样品名称 Sample name | Raw reads/Mb | Clean reads/Mb | Unique map/Mb | Q30/% | GC/% |
|---|---|---|---|---|---|
| CK_S1_1 | 45.32 | 43.78 | 40.87 | 92.94 | 40.88 |
| CK_S1_2 | 48.02 | 45.29 | 43.08 | 94.72 | 40.85 |
| CK_S2_1 | 44.23 | 43.35 | 40.80 | 93.02 | 42.14 |
| CK_S2_2 | 44.89 | 43.22 | 40.68 | 92.63 | 42.28 |
| T_S1_1 | 46.21 | 43.69 | 40.35 | 92.56 | 40.82 |
| T_S1_2 | 47.89 | 45.85 | 42.61 | 92.98 | 41.09 |
| T_S2_1 | 45.83 | 44.53 | 42.08 | 92.9 | 42.43 |
| T_S2_2 | 47.69 | 45.88 | 43.46 | 93.49 | 42.43 |
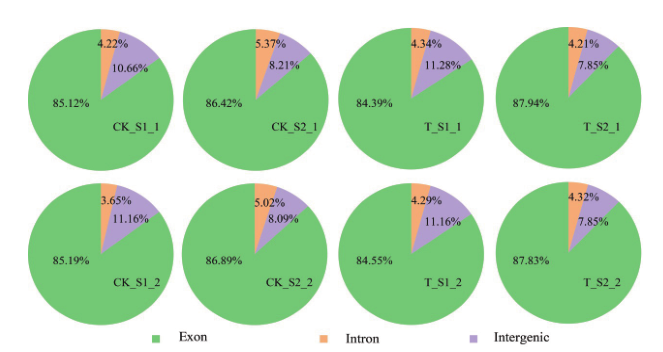
图2 Clean reads在基因组区域分布情况 黄瓜二、四倍体果实不同发育时期样本测序的clean reads在基因组区域分布情况
Fig. 2 Percents of clean reads mapped in genome regions Distribution of clean reads sequenced in different developmental stages of cucumber diploid and tetraploid fruits in genomic regions
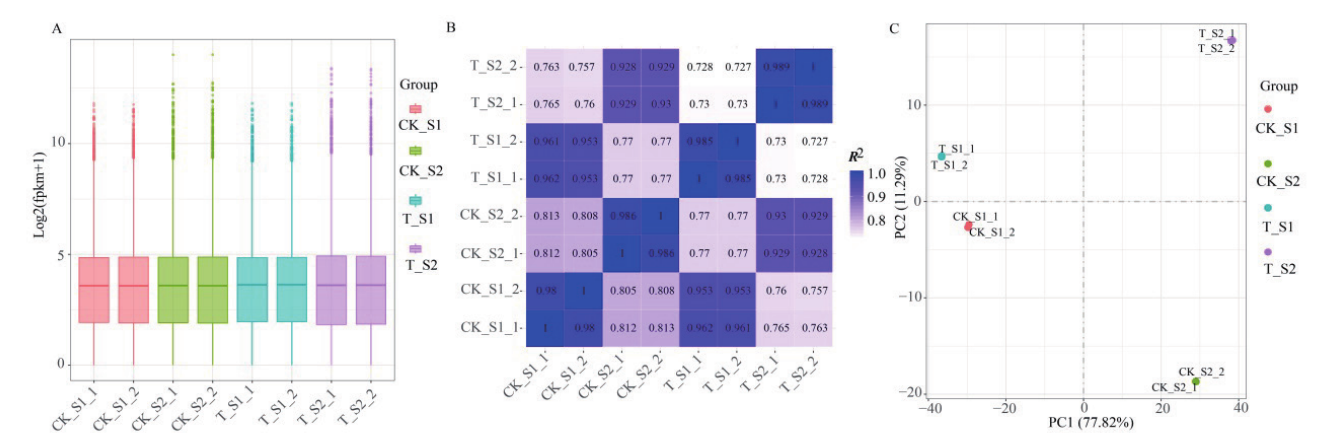
图3 样本基因表达量分布及相关性分析 A:黄瓜二、四倍体果实不同发育时期样本基因表达量分布盒形图;B:黄瓜二、四倍体果实不同发育时期样本皮尔逊相关性分析;C:黄瓜二、四倍体果实不同发育时期样本基因表达量PCA分析
Fig. 3 Gene expression distribution and correlation analysis of samples A: Box diagram of gene expression distribution in cucumber diploid and tetraploid fruits at different developmental stages. B: Pearson correlation analysis of cucumber diploid and tetraploid fruit samples at different developmental stages. C: PCA analysis of gene expression levels in diploid and tetraploid cucumber fruits at different developmental stages
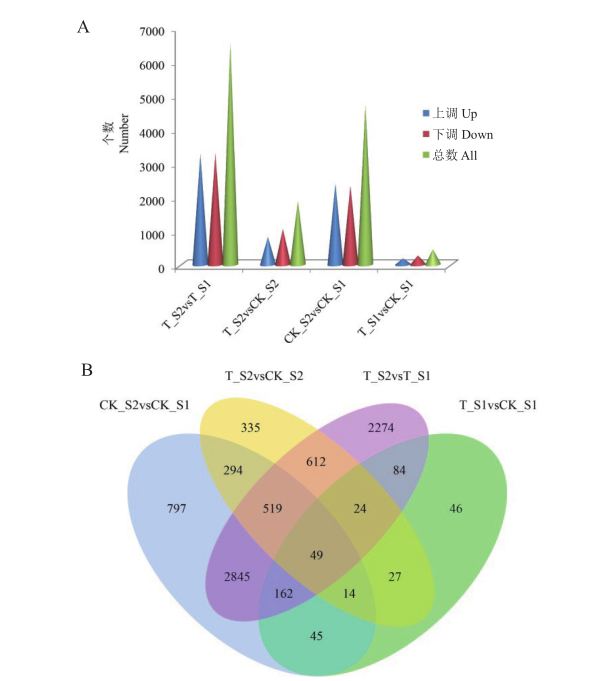
图4 差异基因数目统计及维恩图分析 A:黄瓜二、四倍体果实不同发育时期不同比较组差异基因统计柱状图;B:黄瓜二、四倍体果实不同发育时期不同比较组差异基因维恩图
Fig. 4 Analysis of DEGs statistical number and Venn diagram A: Bar chart of DEGs statistics of different comparison groups in different developmental stages of cucumber diploid and tetraploid fruits. B: Venn diagram of DEGs in different comparison groups at different developmental stages of cucumber diploid and tetraploid fruits
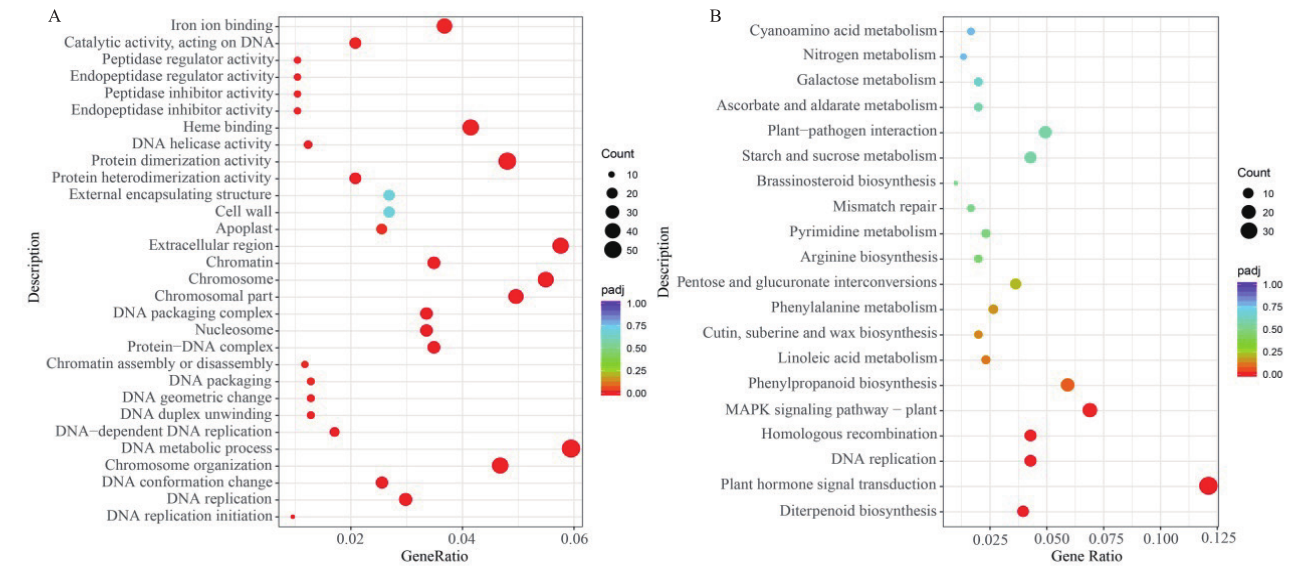
图5 GO与KEGG富集散点图 A:黄瓜二、四倍体果实(T_S2vsCK_S2)差异基因GO富集散点图;B:黄瓜二、四倍体果实(T_S2vsCK_S2)差异基因KEGG富集散点图
Fig. 5 Rich distribution point diagram for GO and KEGG A: Map of GO enrichment of DEGs in T_S2vsCK_S2 of cucumber fruit. B: Map of KEGG enrichment of DEGs in T_S2vsCK_S2 of cucumber fruit
| 基因名称Gene name | 基因 ID Gene ID | log2FC | 基因注释Gene annotation |
|---|---|---|---|
| AUX1 | Csa_3G731880 | 0.511028525 | Auxin influx carrier(AUX1 LAX family) |
| B-ARR | Csa_2G350290 | 1.199233905 | Uncharacterized protein LOC101203501 isoform X1 |
| DELLA | Csa_5G569350 | 0.641993478 | DELLA protein |
| SIMKK | Csa_3G651720 | 0.521551332 | Mitogen-activated protein kinase kinase 4/5 |
| EIN2 | Csa_6G445230 | 0.401357885 | Ethylene-insensitive protein 2 |
| EIN3 | Csa_6G051520 | 0.672607401 | Ethylene-insensitive protein 3 |
| BRI1 | Csa_6G486870 | 0.529640378 | Protein brassinosteroid insensitive 1 |
| BIN2 | Csa_6G157100 | 0.536467357 | Protein brassinosteroid insensitive 2 |
| BZR1/2 | Csa_2G361450 | 0.587660284 | Brassinosteroid resistant 1/2 |
| TCH4 | Csa_1G422470 | 1.185601828 | Xyloglucan:xyloglucosyl transferase TCH4 |
| TGA | Csa_6G152350 | 1.955263915 | Transcription factor TGA |
| GH3 | Csa_6G493310 | -0.840958686 | Auxin responsive GH3 gene family |
| GID1 | Csa_7G391240 | -1.403103089 | Gibberellin receptor GID1 |
| PP2C | Csa_1G574880 | -1.528004155 | Protein phosphatase 2C |
| ETR | Csa_2G070880 | -0.799320186 | Ethylene receptor |
| BAK1 | Csa_7G060160 | -0.875573337 | Brassinosteroid insensitive 1-associated receptor kinase 1 |
| BKI1 | Csa_2G359970 | -1.610933985 | BRI1 kinase inhibitor 1 |
| CYCD3 | Csa_6G491650 | -1.00709524 | Cyclin D3, plant |
| JAZ | Csa_1G042920 | -0.913564833 | Jasmonate ZIM domain-containing protein |
| AGLU1 | Csa_1G716250 | 0.650665663 | Alpha-glucosidase |
| SPS | Csa_2G401440 | 0.977042232 | Sucrose-phosphate synthase |
| SUSY | Csa_5G322500 | 2.088838655 | Sucrose synthase |
| FRK | Csa_3G345390 | -0.650140834 | Fuctokinase |
| AGPase | Csa_3G149890 | -1.393581098 | Glucose-1-phosphate adenylyltransferase |
| SS | Csa_6G497160 | -0.899393235 | Starch synthase |
| GTase | Csa_5G606600 | -0.684711983 | 4-alpha-glucanotransferase |
| GBE1 | Csa_7G213190 | -1.304743764 | 1,4-alpha-glucan branching enzyme |
| BMY | Csa_3G017020 | -3.271484788 | Beta-amylase |
| TRE | Csa_5G149880 | -1.411978481 | Alpha,alpha-trehalase |
表2 KEGG富集通路上关键候选基因
Table 2 Key candidate genes in KEGG enrichment pathway
| 基因名称Gene name | 基因 ID Gene ID | log2FC | 基因注释Gene annotation |
|---|---|---|---|
| AUX1 | Csa_3G731880 | 0.511028525 | Auxin influx carrier(AUX1 LAX family) |
| B-ARR | Csa_2G350290 | 1.199233905 | Uncharacterized protein LOC101203501 isoform X1 |
| DELLA | Csa_5G569350 | 0.641993478 | DELLA protein |
| SIMKK | Csa_3G651720 | 0.521551332 | Mitogen-activated protein kinase kinase 4/5 |
| EIN2 | Csa_6G445230 | 0.401357885 | Ethylene-insensitive protein 2 |
| EIN3 | Csa_6G051520 | 0.672607401 | Ethylene-insensitive protein 3 |
| BRI1 | Csa_6G486870 | 0.529640378 | Protein brassinosteroid insensitive 1 |
| BIN2 | Csa_6G157100 | 0.536467357 | Protein brassinosteroid insensitive 2 |
| BZR1/2 | Csa_2G361450 | 0.587660284 | Brassinosteroid resistant 1/2 |
| TCH4 | Csa_1G422470 | 1.185601828 | Xyloglucan:xyloglucosyl transferase TCH4 |
| TGA | Csa_6G152350 | 1.955263915 | Transcription factor TGA |
| GH3 | Csa_6G493310 | -0.840958686 | Auxin responsive GH3 gene family |
| GID1 | Csa_7G391240 | -1.403103089 | Gibberellin receptor GID1 |
| PP2C | Csa_1G574880 | -1.528004155 | Protein phosphatase 2C |
| ETR | Csa_2G070880 | -0.799320186 | Ethylene receptor |
| BAK1 | Csa_7G060160 | -0.875573337 | Brassinosteroid insensitive 1-associated receptor kinase 1 |
| BKI1 | Csa_2G359970 | -1.610933985 | BRI1 kinase inhibitor 1 |
| CYCD3 | Csa_6G491650 | -1.00709524 | Cyclin D3, plant |
| JAZ | Csa_1G042920 | -0.913564833 | Jasmonate ZIM domain-containing protein |
| AGLU1 | Csa_1G716250 | 0.650665663 | Alpha-glucosidase |
| SPS | Csa_2G401440 | 0.977042232 | Sucrose-phosphate synthase |
| SUSY | Csa_5G322500 | 2.088838655 | Sucrose synthase |
| FRK | Csa_3G345390 | -0.650140834 | Fuctokinase |
| AGPase | Csa_3G149890 | -1.393581098 | Glucose-1-phosphate adenylyltransferase |
| SS | Csa_6G497160 | -0.899393235 | Starch synthase |
| GTase | Csa_5G606600 | -0.684711983 | 4-alpha-glucanotransferase |
| GBE1 | Csa_7G213190 | -1.304743764 | 1,4-alpha-glucan branching enzyme |
| BMY | Csa_3G017020 | -3.271484788 | Beta-amylase |
| TRE | Csa_5G149880 | -1.411978481 | Alpha,alpha-trehalase |
| 基因名称Gene name | 注释Annotation | 引物序列Primer sequence(5'-3') |
|---|---|---|
| SUSY | 蔗糖合酶 Sucrose synthase | F:TGAGAACGATGAGCATATAG |
| R:TCCAACCACAACAAGATT | ||
| K-box | 转录因子K-box Transcription factor K-box | F:CAAGAACAGGAAGAGGAA |
| R:GAGGAGAAGACGATAAGAG | ||
| WUS | WUSCHEL相关homeobox WUSCHEL-related homeobox | F:GTGAGCCATTGATGACAT |
| R:GGAACCAGTTATAGACATTAGA | ||
| SUT | 蔗糖转运蛋白 Sucrose transporter | F:CTGAGACGGTTACTAAGAG |
| R:TTGAATATAAGGAGTGAGAAGA | ||
| GLU8 | 葡萄糖转运蛋白8 Glucose transporter protein 8 | F:CCTCACAACACTCAATCA |
| R:CACAGACCAACAATCAGA | ||
| DNAH | DNA解旋酶 DNA helicase | F:GCTCAAGTCGTTACCATT |
| R:CATCAACTGAACTCACCTT | ||
| ARF4 | 激素响应因子4 Auxin response factor 4 | F:GTCTACCATACTCGTGTT |
| R:CTAATGCCAGTGATTGTG | ||
| GIR | 赤霉素调控蛋白 Gibberellin-regulated protein | F:TCCTCTCTTCTCGTTCTT |
| R:GCTCCTCCACAATCAATA | ||
| LRK | 蛋白激酶家族蛋白 Protein kinase family protein | F:CGGAGATGACTGTTGTAA |
| R:TCCAATCAGCAATAATAACG | ||
| CsActin | 内参基因 Actin gene | F:ATTGTTCTCAGTGGTGGTTCTAC |
| R:CCTTTGAGATCCACATCTGCT |
表3 RT-qPCR引物序列信息
Table 3 Information of RT-qPCR primer sequences
| 基因名称Gene name | 注释Annotation | 引物序列Primer sequence(5'-3') |
|---|---|---|
| SUSY | 蔗糖合酶 Sucrose synthase | F:TGAGAACGATGAGCATATAG |
| R:TCCAACCACAACAAGATT | ||
| K-box | 转录因子K-box Transcription factor K-box | F:CAAGAACAGGAAGAGGAA |
| R:GAGGAGAAGACGATAAGAG | ||
| WUS | WUSCHEL相关homeobox WUSCHEL-related homeobox | F:GTGAGCCATTGATGACAT |
| R:GGAACCAGTTATAGACATTAGA | ||
| SUT | 蔗糖转运蛋白 Sucrose transporter | F:CTGAGACGGTTACTAAGAG |
| R:TTGAATATAAGGAGTGAGAAGA | ||
| GLU8 | 葡萄糖转运蛋白8 Glucose transporter protein 8 | F:CCTCACAACACTCAATCA |
| R:CACAGACCAACAATCAGA | ||
| DNAH | DNA解旋酶 DNA helicase | F:GCTCAAGTCGTTACCATT |
| R:CATCAACTGAACTCACCTT | ||
| ARF4 | 激素响应因子4 Auxin response factor 4 | F:GTCTACCATACTCGTGTT |
| R:CTAATGCCAGTGATTGTG | ||
| GIR | 赤霉素调控蛋白 Gibberellin-regulated protein | F:TCCTCTCTTCTCGTTCTT |
| R:GCTCCTCCACAATCAATA | ||
| LRK | 蛋白激酶家族蛋白 Protein kinase family protein | F:CGGAGATGACTGTTGTAA |
| R:TCCAATCAGCAATAATAACG | ||
| CsActin | 内参基因 Actin gene | F:ATTGTTCTCAGTGGTGGTTCTAC |
| R:CCTTTGAGATCCACATCTGCT |
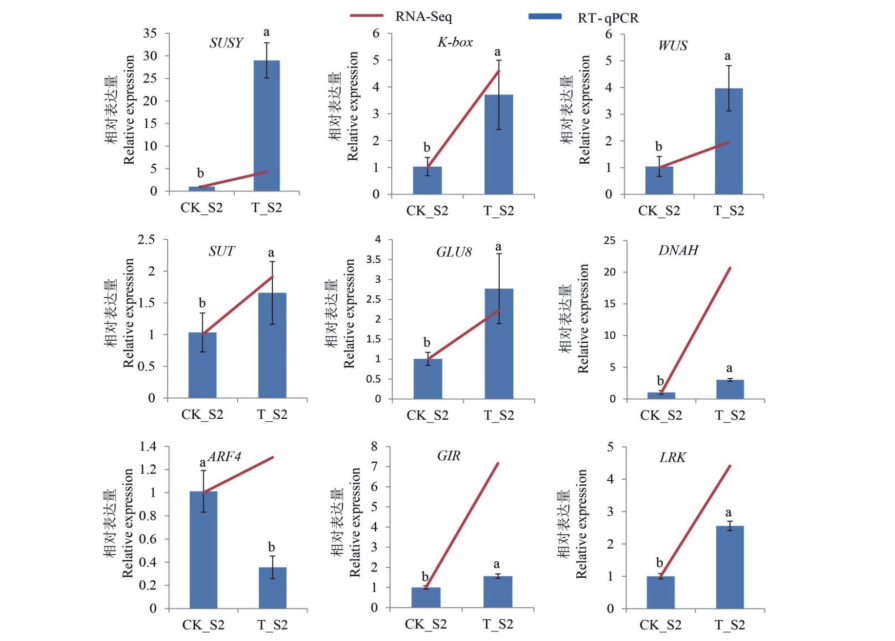
图6 候选基因RT-qPCR分析 不同小写字母代表同一基因在二、四倍体黄瓜果实中表达量存在显著差异(P<0.05)
Fig. 6 RT-qPCR analysis of candidate genes Different lowercase letters indicate significant differences in the expression levels of the same gene in diploid and tetraploid cucumber fruits at 0.05 level
| [1] | 潘玉朋, 孟焕文, 陈书霞, 等. 蔬菜作物果形研究进展[J]. 中国蔬菜, 2013(4): 6-13. |
| Pan YP, Meng HW, Chen SX, et al. Research progress on fruit shape of vegetable crops[J]. China Veg, 2013(4): 6-13. | |
| [2] | 黄三文. 黄瓜果实品质性状的基因组学研究[J]. 中国科技成果, 2013(21): 26-28. |
| Hang SW. Genomic studies on fruit quality traits in cucumber[J]. China Sci Technol Achiev, 2013(21): 26-28. | |
| [3] | 王一衡, 黄胜楠, 刘志勇, 等. 大白菜倍性变异引起花发育变化的microRNA调控机制研究[J]. 沈阳农业大学学报, 2018, 49(5): 529-536. |
| Wang YH, Huang SN, Liu ZY, et al. Mechanism of floral development variation resulted from ploidy changes regulated by microRNA in Chinese cabbage[J]. J Shenyang Agric Univ, 2018, 49(5): 529-536. | |
| [4] |
Yin L, Qu JJ, Zhou HW, et al. Comparison of leaf transcriptomes of cassava ‘Xinxuan 048’ diploid and autotetraploid plants[J]. Genes Genomics, 2018, 40(9): 927-935.
doi: 10.1007/s13258-018-0692-2 |
| [5] |
Cheng WW, Tang MJ, Xie Y, et al. Transcriptome-based gene expression profiling of diploid radish(Raphanus sativus L.)and the corresponding autotetraploid[J]. Mol Biol Rep, 2019, 46(1): 933-945.
doi: 10.1007/s11033-018-4549-1 |
| [6] |
Wang HL, Li YQ, Wang SB, et al. Comparative transcriptomic analyses of chlorogenic acid and luteolosides biosynthesis pathways at different flowering stages of diploid and tetraploid Lonicera japonica[J]. PeerJ, 2020, 8: e8690.
doi: 10.7717/peerj.8690 URL |
| [7] | 刘美妍, 周国彦, 李晓丽, 等. 秋水仙素对华北型密刺黄瓜的诱变效应[J]. 中国瓜菜, 2021, 34(11): 10-16. |
| Liu MY, Zhou GY, Li XL, et al. Mutagenesis effect of colchicine on North China cucumber with dense thorny[J]. China Cucurbits Veg, 2021, 34(11): 10-16. | |
| [8] |
Azimian J, Hervan EM, Azadi A, et al. Transcriptome analysis of a Triticum aestivum landrace(Roshan)in response to salt stress conditions[J]. Plant Genet Resour, 2021, 19(3): 261-274.
doi: 10.1017/S1479262121000319 URL |
| [9] |
Mortazavi A, Williams BA, McCue K, et al. Mapping and quantifying mammalian transcriptomes by RNA-seq[J]. Nat Methods, 2008, 5(7): 621-628.
doi: 10.1038/nmeth.1226 pmid: 18516045 |
| [10] |
Mereghetti L, Sitkiewicz I, M. Green N, et al. Principal component analysis(PCA)plot showing transcriptome differences between expression microarray data of GBS NEM316 strain incubated in human blood at 37℃[J]. PLoS One, 2009, 4(9): e7145.
doi: 10.1371/journal.pone.0007145 URL |
| [11] | Saetan W, Tian CX, Yu JW, et al. Comparative transcriptome analysis of gill tissue in response to hypoxia in silver Sillago(Sillago sihama)[J]. Animals(Basel), 2020, 10(4): 628. |
| [12] |
Xu YX, Chen JW, Yang ZH, et al. Identification of RNA expression profiles in thyroid cancer to construct a competing endogenous RNA(CeRNA)network of mRNAs, long noncoding RNAs(lncRNAs), and microRNAs(miRNAs)[J]. Med Sci Monit, 2019, 25: 1140-1154.
doi: 10.12659/MSM.912450 URL |
| [13] |
周国彦, 银珊珊, 高佳鑫, 等. 黄瓜AHP基因家族的鉴定及其非生物胁迫表达分析[J]. 生物技术通报, 2022, 38(6): 112-119.
doi: 10.13560/j.cnki.biotech.bull.1985.2021-1338 |
| Zhou GY, Yin SS, Gao JX, et al. Identification of AHP gene family in Cucumis sativus and its expression analysis under abiotic stress[J]. Biotechnol Bull, 2022, 38(6): 112-119. | |
| [14] |
Che G, Zhang XL. Molecular basis of cucumber fruit domestication[J]. Curr Opin Plant Biol, 2019, 47: 38-46.
doi: S1369-5266(18)30032-3 pmid: 30253288 |
| [15] |
Weng YQ, Colle M, Wang YH, et al. QTL mapping in multiple populations and development stages reveals dynamic quantitative trait loci for fruit size in cucumbers of different market classes[J]. Theor Appl Genet, 2015, 128(9): 1747-1763.
doi: 10.1007/s00122-015-2544-7 pmid: 26048092 |
| [16] |
Pan YP, Qu SP, Bo KL, et al. QTL mapping of domestication and diversifying selection related traits in round-fruited semi-wild Xishuangbanna cucumber(Cucumis sativus L. var. xishuangbannanesis)[J]. Theor Appl Genet, 2017, 130(7): 1531-1548.
doi: 10.1007/s00122-017-2908-2 URL |
| [17] |
Pan YP, Liang XJ, Gao ML, et al. Round fruit shape in WI7239 cucumber is controlled by two interacting quantitative trait loci with one putatively encoding a tomato SUN homolog[J]. Theor Appl Genet, 2017, 130(3): 573-586.
doi: 10.1007/s00122-016-2836-6 pmid: 27915454 |
| [18] |
Wang LJ, Sheng MY, Wen PC, et al. Morphological, physiological, cytological and phytochemical studies in diploid and colchicine-induced tetraploid plants of Fagopyrum tataricum(L.)Gaertn[J]. Bot Stud, 2017, 58(1): 2.
doi: 10.1186/s40529-016-0157-3 URL |
| [19] | 姜文芝. 不同倍性黄瓜植物学特征及生理生化的研究[D]. 泰安: 山东农业大学, 2017. |
| Jiang WZ. Study on the botanical characteristics and physiological and biochemical of different ploidy cucumber[D]. Tai'an: Shandong Agricultural University, 2017. | |
| [20] | 刘永月, 许建鹏, 李田田, 等. 无毛黄瓜同源四倍体诱导及鉴定[J]. 山东农业科学, 2016, 48(11): 21-25. |
| Liu YY, Xu JP, Li TT, et al. Induction and identification of autotetraploid of glabrous cucumber[J]. Shandong Agric Sci, 2016, 48(11): 21-25. | |
| [21] |
Martin SL, Husband BC. Whole genome duplication affects evolvability of flowering time in an autotetraploid plant[J]. PLoS One, 2012, 7(9): e44784.
doi: 10.1371/journal.pone.0044784 URL |
| [22] | 孙小武, 杨仁喜, 邓大成, 等. 无籽西瓜新品种‘雪峰新二号’的选育[J]. 中国瓜菜, 2018, 31(6): 20-22. |
| Sun XW, Yang RX, Deng DC, et al. Breeding of new seedless watermelon cultivar ‘Xuefeng Xin no. 2’[J]. China Cucurbits Veg, 2018, 31(6): 20-22. | |
| [23] | 高素燕, 焦定量, 商纪鹏, 等. 无籽西瓜新品种‘津蜜55’的选育[J]. 中国瓜菜, 2018, 31(6): 23-25. |
| Gao SY, Jiao DL, Shang JP, et al. Breeding of new seedless watermelon variety ‘Jinmi No.55’[J]. China Cucurbits Veg, 2018, 31(6): 23-25. | |
| [24] |
Adams KL, Cronn R, Percifield R, et al. Genes duplicated by polyploidy show unequal contributions to the transcriptome and organ-specific reciprocal silencing[J]. Proc Natl Acad Sci USA, 2003, 100(8): 4649-4654.
doi: 10.1073/pnas.0630618100 pmid: 12665616 |
| [25] |
Adams KL, Wendel JF. Novel patterns of gene expression in polyploid plants[J]. Trends Genet, 2005, 21(10): 539-543.
pmid: 16098633 |
| [26] |
Alabadí D, Blázquez MA, Carbonell J, et al. Instructive roles for hormones in plant development[J]. Int J Dev Biol, 2009, 53(8/9/10): 1597-1608.
doi: 10.1387/ijdb.072423da URL |
| [27] |
Nemhauser JL, Hong FX, Chory J. Different plant hormones regulate similar processes through largely nonoverlapping transcriptional responses[J]. Cell, 2006, 126(3): 467-475.
doi: 10.1016/j.cell.2006.05.050 pmid: 16901781 |
| [28] |
Anfang MR, Shani E. Transport mechanisms of plant hormones[J]. Curr Opin Plant Biol, 2021, 63: 102055.
doi: 10.1016/j.pbi.2021.102055 URL |
| [29] |
Liu JG, Guo SG, He HJ, et al. Dynamic characteristics of sugar accumulation and related enzyme activities in sweet and non-sweet watermelon fruits[J]. Acta Physiol Plant, 2013, 35(11): 3213-3222.
doi: 10.1007/s11738-013-1356-0 URL |
| [30] |
Fan JW, Wang HY, Li X, et al. Down-regulating cucumber sucrose synthase 4(CsSUS4)suppresses the growth and development of flowers and fruits[J]. Plant Cell Physiol, 2019, 60(4): 752-764.
doi: 10.1093/pcp/pcy239 URL |
| [31] |
Baroja-Fernández E, Muñoz FJ, Montero M, et al. Enhancing sucrose synthase activity in transgenic potato(Solanum tuberosum L.)tubers results in increased levels of starch, ADPglucose and UDPglucose and total yield[J]. Plant Cell Physiol, 2009, 50(9): 1651-1662.
doi: 10.1093/pcp/pcp108 pmid: 19608713 |
| [32] |
Xie Y, Ying JL, Xu L, et al. Genome-wide sRNA and mRNA transcriptomic profiling insights into dynamic regulation of taproot thickening in radish(Raphanus sativus L.)[J]. BMC Plant Biol, 2020, 20(1): 373.
doi: 10.1186/s12870-020-02585-z |
| [33] |
D’Aoust MA, Yelle S, Nguyen-Quoc B. Antisense inhibition of tomato fruit sucrose synthase decreases fruit setting and the sucrose unloading capacity of young fruit[J]. Plant Cell, 1999, 11(12): 2407-2418.
doi: 10.1105/tpc.11.12.2407 pmid: 10590167 |
| [34] |
Jiang QY, Hou J, Hao CY, et al. The wheat(T. aestivum)sucrose synthase 2 gene(TaSus2)active in endosperm development is associated with yield traits[J]. Funct Integr Genomics, 2011, 11(1): 49-61.
doi: 10.1007/s10142-010-0188-x URL |
| [1] | 林红妍, 郭晓蕊, 刘迪, 李慧, 陆海. 转录组分析转录因子AtbHLH68调控细胞壁发育的分子机制[J]. 生物技术通报, 2023, 39(9): 105-116. |
| [2] | 娄慧, 朱金成, 杨洋, 张薇. 抗、感品种棉花根系分泌物对尖孢镰刀菌生长及基因表达的影响[J]. 生物技术通报, 2023, 39(9): 156-167. |
| [3] | 苗永美, 苗翠苹, 于庆才. 枯草芽孢杆菌BBs-27发酵液性质及脂肽对黄色镰刀菌的抑菌作用[J]. 生物技术通报, 2023, 39(9): 255-267. |
| [4] | 付钰, 贾瑞瑞, 何荷, 王良桂, 杨秀莲. 两种砧木楸树嫁接苗生长差异及转录组比较分析[J]. 生物技术通报, 2023, 39(8): 251-261. |
| [5] | 褚睿, 李昭轩, 张学青, 杨东亚, 曹行行, 张雪艳. 黄瓜枯萎病拮抗芽孢杆菌的筛选、鉴定及其生防潜力[J]. 生物技术通报, 2023, 39(8): 262-271. |
| [6] | 赵金玲, 安磊, 任晓亮. 单细胞转录组测序技术及其在秀丽隐杆线虫中的应用[J]. 生物技术通报, 2023, 39(6): 158-170. |
| [7] | 孔德真, 段震宇, 王刚, 张鑫, 席琳乔. 盐、碱胁迫下高丹草苗期生理特征及转录组学分析[J]. 生物技术通报, 2023, 39(6): 199-207. |
| [8] | 杨洋, 朱金成, 娄慧, 韩泽刚, 张薇. 海岛棉与枯萎病菌的互作转录组分析[J]. 生物技术通报, 2023, 39(6): 259-273. |
| [9] | 刘辉, 卢扬, 叶夕苗, 周帅, 李俊, 唐健波, 陈恩发. 外源硫诱导苦荞镉胁迫响应的比较转录组学分析[J]. 生物技术通报, 2023, 39(5): 177-191. |
| [10] | 王琪, 胡哲, 富薇, 李光哲, 郝林. 伯克霍尔德氏菌GD17对黄瓜幼苗耐干旱的调节[J]. 生物技术通报, 2023, 39(3): 163-175. |
| [11] | 扈丽丽, 林柏荣, 王宏洪, 陈建松, 廖金铃, 卓侃. 最短尾短体线虫转录组及潜在效应蛋白分析[J]. 生物技术通报, 2023, 39(3): 254-266. |
| [12] | 杨东亚, 祁瑞雪, 李昭轩, 林薇, 马慧, 张雪艳. 黄瓜茄病镰刀菌拮抗芽孢杆菌的筛选、鉴定及促生效果[J]. 生物技术通报, 2023, 39(2): 211-220. |
| [13] | 孙言秋, 谢采芸, 汤岳琴. 耐高温酿酒酵母的构建与高温耐受机制解析[J]. 生物技术通报, 2023, 39(11): 226-237. |
| [14] | 徐俊, 叶雨晴, 牛雅静, 黄河, 张蒙蒙. 菊花根状茎发育的转录组分析[J]. 生物技术通报, 2023, 39(10): 231-245. |
| [15] | 罗皓天, 王龙, 王禹茜, 王月, 李佳祯, 杨梦珂, 张杰, 邓欣, 王红艳. 青狗尾草RNAi途径相关基因的全基因组鉴定和表达分析[J]. 生物技术通报, 2023, 39(1): 175-186. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||