








• 研究报告 • 下一篇
王家彬1( ), 胡玥2, 陈嘉杰1, 王蒙2, 何秀云1, 李之勇2, 李健2, 王美娜2(
), 胡玥2, 陈嘉杰1, 王蒙2, 何秀云1, 李之勇2, 李健2, 王美娜2( )
)
收稿日期:2025-08-27
出版日期:2026-03-09
通讯作者:
王美娜,女,博士,高级工程师,研究方向 :兰科植物宏基因组学、分类与系统发育学、药用植物学;E-mail: snow-wmn2005@163.com作者简介:王家彬,男,高级工程师,研究方向 :林业保护;E-mail: 44376283@qq.com基金资助:
WANG Jia-bin1( ), HU Yue2, CHEN Jia-jie1, WANG Meng2, HE Xiu-yun1, LI Zhi-yong2, LI Jian2, WANG Mei-na2(
), HU Yue2, CHEN Jia-jie1, WANG Meng2, HE Xiu-yun1, LI Zhi-yong2, LI Jian2, WANG Mei-na2( )
)
Received:2025-08-27
Published:2026-03-09
摘要:
目的 解析紫纹兜兰(Paphiopedilum purpuratum)不同部位的功能分化机制,通过多组学策略系统揭示其代谢物分布特征、基因表达模式及“代谢‒基因”协同调控网络,为理解该物种环境适应性提供分子基础。 方法 采用非靶向代谢组学(超高效液相色谱‒串联质谱,UHPLC-MS/MS)分析紫纹兜兰叶片(SL)、花(SF)、根(SR)3个不同部位代谢物差异,通过转录组测序分析差异表达基因,并构建其代谢物‒基因共表达网络。 结果 代谢组共鉴定2 164个代谢物(正离子模式1 471个,负离子模式693个),组间差异显著,SL vs. SF(正/负离子模式:499/193种)、SL vs. SR(534/243种)、SF vs. SR(433/187种)。KEGG富集分析显示差异代谢物主要富集在苯丙烷类物质生物合成、类黄酮生物合成、亚油酸等核心代谢通路上。转录组学分析揭示了强烈的组织特异性基因表达:叶片中光合基因(如sqdB、DVR)显著上调,花朵中芳香/色素合成基因(如FAH、crtZ)高表达,根部胁迫响应基因(如CCR、GPAT)上调。共表达网络鉴定出关键调控因子,如INO80B(调控黄酮合成)、CCT1(调控萜类代谢)和CCR(参与根部防御)。 结论 紫纹兜兰通过其叶片的光合与抗逆协同、花朵的传粉吸引以及根部的胁迫防御这3个组织特异性的“代谢‒基因”协同调控网络,共同构成了其生态适应性的分子机制。
王家彬, 胡玥, 陈嘉杰, 王蒙, 何秀云, 李之勇, 李健, 王美娜. 紫纹兜兰不同部位的代谢组和转录组联合分析[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0927.
WANG Jia-bin, HU Yue, CHEN Jia-jie, WANG Meng, HE Xiu-yun, LI Zhi-yong, LI Jian, WANG Mei-na. Combined Transcriptomic and Metabolomic Analysis in Different Parts of Paphiopedilum purpuratum[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-0927.
| 类型 Type | SL vs. SF | SL vs. SR | SF vs. SR | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 上调 Up | 下调 Down | 上调 Up | 下调 Down | 上调 Up | 下调 Down | |||||||
| Positive | Negative | Positive | Negative | Positive | Negative | Positive | Negative | Positive | Negative | Positive | Negative | |
| 脂质 Lipids | 86 | 27 | 21 | 18 | 87 | 27 | 20 | 47 | 30 | 14 | 36 | 37 |
| 苯丙类 Phenylpropanoids | 24 | 14 | 23 | 17 | 26 | 10 | 38 | 20 | 27 | 14 | 38 | 17 |
| 有机氧 Organic oxygen compounds | 23 | 7 | 11 | 18 | 15 | 7 | 10 | 14 | 5 | 15 | 6 | 10 |
| 有机酸 Organic acids | 19 | 1 | 27 | 11 | 10 | 4 | 48 | 16 | 13 | 6 | 28 | 7 |
| 有机杂环 Heterocyclic compounds | 25 | 2 | 15 | 9 | 19 | 2 | 25 | 5 | 11 | 3 | 31 | 2 |
| 核苷类 Nucleosides | 5 | 1 | 6 | 8 | 1 | 2 | 8 | 7 | 3 | 5 | 8 | 0 |
| 苯类 Benzenoids | 27 | 2 | 14 | 6 | 21 | 2 | 18 | 11 | 9 | 2 | 22 | 9 |
| 有机氮 Organic nitrogen compounds | 1 | 1 | 10 | 1 | 0 | 1 | 5 | 1 | 4 | 0 | 3 | 0 |
| 生物碱 Alkaloids | 6 | 1 | 1 | 0 | 4 | 2 | 2 | 1 | 2 | 1 | 3 | 0 |
| 木脂素 Lignans | 2 | 1 | 1 | 0 | 1 | 0 | 1 | 4 | 0 | 0 | 3 | 4 |
| 其他类 Others | 101 | 21 | 51 | 27 | 81 | 35 | 94 | 25 | 49 | 25 | 102 | 16 |
| 总计 Total | 319 | 78 | 180 | 115 | 265 | 92 | 269 | 151 | 153 | 85 | 280 | 102 |
表1 代谢物在各部位差异表达情况
Table 1 Differential expressions of metabolites in different tissues
| 类型 Type | SL vs. SF | SL vs. SR | SF vs. SR | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 上调 Up | 下调 Down | 上调 Up | 下调 Down | 上调 Up | 下调 Down | |||||||
| Positive | Negative | Positive | Negative | Positive | Negative | Positive | Negative | Positive | Negative | Positive | Negative | |
| 脂质 Lipids | 86 | 27 | 21 | 18 | 87 | 27 | 20 | 47 | 30 | 14 | 36 | 37 |
| 苯丙类 Phenylpropanoids | 24 | 14 | 23 | 17 | 26 | 10 | 38 | 20 | 27 | 14 | 38 | 17 |
| 有机氧 Organic oxygen compounds | 23 | 7 | 11 | 18 | 15 | 7 | 10 | 14 | 5 | 15 | 6 | 10 |
| 有机酸 Organic acids | 19 | 1 | 27 | 11 | 10 | 4 | 48 | 16 | 13 | 6 | 28 | 7 |
| 有机杂环 Heterocyclic compounds | 25 | 2 | 15 | 9 | 19 | 2 | 25 | 5 | 11 | 3 | 31 | 2 |
| 核苷类 Nucleosides | 5 | 1 | 6 | 8 | 1 | 2 | 8 | 7 | 3 | 5 | 8 | 0 |
| 苯类 Benzenoids | 27 | 2 | 14 | 6 | 21 | 2 | 18 | 11 | 9 | 2 | 22 | 9 |
| 有机氮 Organic nitrogen compounds | 1 | 1 | 10 | 1 | 0 | 1 | 5 | 1 | 4 | 0 | 3 | 0 |
| 生物碱 Alkaloids | 6 | 1 | 1 | 0 | 4 | 2 | 2 | 1 | 2 | 1 | 3 | 0 |
| 木脂素 Lignans | 2 | 1 | 1 | 0 | 1 | 0 | 1 | 4 | 0 | 0 | 3 | 4 |
| 其他类 Others | 101 | 21 | 51 | 27 | 81 | 35 | 94 | 25 | 49 | 25 | 102 | 16 |
| 总计 Total | 319 | 78 | 180 | 115 | 265 | 92 | 269 | 151 | 153 | 85 | 280 | 102 |
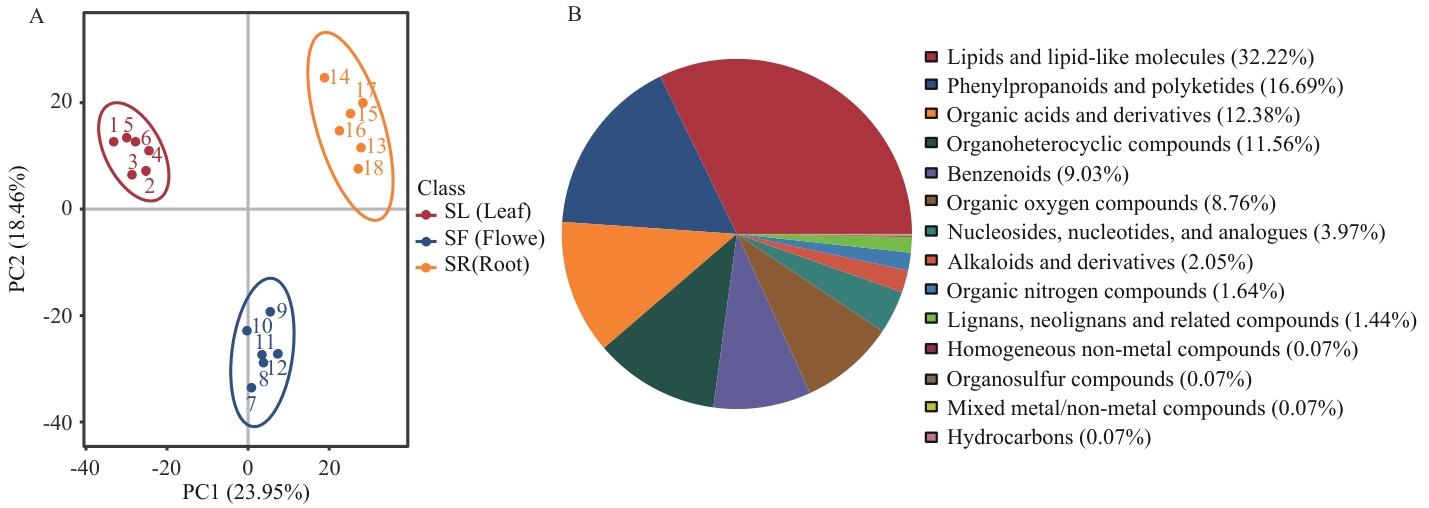
图1 紫纹兜兰代谢物分类情况A:总样本PCA分析图;B:代谢物分类饼状图。SL:叶;SF:花;SR:根。下同
Fig. 1 Classification of metabolites of P. purpuratumA: PCA analysis diagram of the total sample. B: Pie chart for metabolite classification. SL: Leaf. SF: Flower. SR: Root. The same below
代谢物类别 Metabolite category | 代谢物名称 Metabolite name | SL vs. SF | SL vs. SR | SF vs. SR | |||
|---|---|---|---|---|---|---|---|
| Log2FC | VIP | Log2FC | VIP | Log2FC | VIP | ||
| 脂质和类脂分子 Lipids and lipid-like molecules | 姜糖脂B Gingerglycolipid B | 1.95 | 1.52 | 6.92 | 1.19 | 4.96 | 1.07 |
| 重楼皂苷 VI Polyphyllin VI | 2.51 | 1.36 | 6.36 | 1.63 | 3.85 | 1.34 | |
| 柴胡皂苷C Saikosaponin C | 5.72 | 1.66 | 6.06 | 1.05 | / | / | |
| 单酰甘油(18∶3) MAG (18∶3) | / | / | 6.01 | 1.31 | 5.17 | 1.71 | |
| 海葱苷A Proscillaridin A | 5.07 | 1.68 | 6.01 | 1.47 | / | / | |
| 知母皂苷A3 Timosaponin A3 | / | / | 5.95 | 1.39 | 5.26 | 1.50 | |
| 脱氧皮质酮21-葡萄糖苷 Deoxycorticosterone 21-glucoside | 2.52 | 1.76 | 5.76 | 1.86 | 3.23 | 1.25 | |
| 羟基积雪草苷 Madecassoside | 5.31 | 1.41 | 2.78 | 1.50 | / | / | |
| 远志酸 Polygalic acid | 3.18 | 1.60 | 5.20 | 1.79 | / | / | |
| 香叶基丙酮 Geranylacetone | 4.55 | 1.66 | 5.13 | 1.07 | / | / | |
| 二十二碳六烯酸 Docosahexaenoic acid | 4.93 | 1.00 | 3.80 | 1.17 | / | / | |
| 20-羟基蜕皮酮 20-Hydroxyecdysone | 4.05 | 1.43 | 4.88 | 1.14 | / | / | |
| 二氢玫瑰苷 Dihydroroseoside | 4.58 | 1.84 | 4.67 | 1.12 | / | / | |
| 茯苓酸B Poricoic acid B | / | / | 4.67 | 1.01 | 3.84 | 1.52/ | |
| 草乌甲素A Bulleyaconi cine A | / | / | -3.66 | 1.18 | -4.85 | 2.10 | |
| 拉帕醇 Lapachol | -2.75 | 1.07 | -3.81 | 2.68 | / | / | |
| 丹参酚C Danshenol C | / | / | -4.06 | 1.54 | -4.14 | 1.74 | |
| 桦木酮酸 Betulonic acid | 2.34 | 1.17 | / | / | -4.32 | 2.24 | |
| 乙酰哈帕苷 Acetylharpagide | / | / | -3.59 | 1.47 | -4.41 | 1.06 | |
| 毒毛旋花苷 Ouabain | / | / | -4.54 | 1.38 | -4.34 | 1.66 | |
| 竹节参皂苷 IVa Chikusetsusponin IVa | 3.69 | 1.67 | -1.98 | 1.69 | -4.80 | 1.10 | |
| 8-异前列腺素A2 8-iso Prostaglandin A2 | / | / | -4.86 | 1.73 | -5.02 | 1.72 | |
| 马鞭草苷 Verbenalin | 1.16 | 1.04 | -5.09 | 1.15 | -6.25 | 2.02 | |
| 亚油酸 Linoleic acid | / | / | -4.58 | 1.53 | -5.61 | 2.07 | |
| 玫瑰苷 Roseoside | / | / | 4.67 | 1.12 | -6.34 | 1.30 | |
| (±)-脱落酸 (±)-Abscisic acid | / | / | -6.28 | 1.49 | -6.46 | 1.47 | |
| 苯丙类和聚酮类 Phenylpropanoids and polyketides | 槲皮素-3β-D-葡萄糖苷 Quercetin-3β-D-glucoside | -7.65 | 1.84 | / | / | 8.26 | 1.31 |
| 山奈酚 Kaempferol | / | / | 7.27 | 1.63 | 6.60 | 1.29 | |
| 荭草素 Orientin | -1.97 | 1.13 | 3.84 | 1.13 | 5.81 | 1.84 | |
| 矢车菊素-3-O-葡萄糖苷 Kuromanin | / | / | 5.24 | 1.33 | 5.15 | 1.74 | |
| 柚皮素 Naringenin | -3.70 | 1.45 | / | / | 2.65 | 1.06 | |
| 咖啡酸乙酯 Ethyl caffeate | / | / | -4.13 | 1.93 | -3.10 | 1.47 | |
| 落新妇苷 Astilbin | -1.65 | 1.30 | -4.57 | 1.48 | / | / | |
| 异茴芹素 Isopimpinellin | -4.71 | 2.03 | / | / | 4.46 | 1.00 | |
| 曲克芦丁 Troxerutin | -5.37 | 1.48 | -5.72 | 1.61 | / | / | |
| 车前苷 Plantagoside | -6.19 | 1.28 | / | / | 3.64 | 1.32 | |
| 毛蕊花糖苷 Verbascoside | / | / | -6.41 | 1.32 | -4.20 | 1.54 | |
| 槲皮素3-O-槐糖苷 Quercetin 3-O-sophoroside | -9.93 | 1.20 | / | / | 9.64 | 1.91 | |
表2 紫纹兜兰不同部位主要显著差异代谢物信息
Table 2 Information on the main significant differential metabolites in different parts of the P. purpuratum
代谢物类别 Metabolite category | 代谢物名称 Metabolite name | SL vs. SF | SL vs. SR | SF vs. SR | |||
|---|---|---|---|---|---|---|---|
| Log2FC | VIP | Log2FC | VIP | Log2FC | VIP | ||
| 脂质和类脂分子 Lipids and lipid-like molecules | 姜糖脂B Gingerglycolipid B | 1.95 | 1.52 | 6.92 | 1.19 | 4.96 | 1.07 |
| 重楼皂苷 VI Polyphyllin VI | 2.51 | 1.36 | 6.36 | 1.63 | 3.85 | 1.34 | |
| 柴胡皂苷C Saikosaponin C | 5.72 | 1.66 | 6.06 | 1.05 | / | / | |
| 单酰甘油(18∶3) MAG (18∶3) | / | / | 6.01 | 1.31 | 5.17 | 1.71 | |
| 海葱苷A Proscillaridin A | 5.07 | 1.68 | 6.01 | 1.47 | / | / | |
| 知母皂苷A3 Timosaponin A3 | / | / | 5.95 | 1.39 | 5.26 | 1.50 | |
| 脱氧皮质酮21-葡萄糖苷 Deoxycorticosterone 21-glucoside | 2.52 | 1.76 | 5.76 | 1.86 | 3.23 | 1.25 | |
| 羟基积雪草苷 Madecassoside | 5.31 | 1.41 | 2.78 | 1.50 | / | / | |
| 远志酸 Polygalic acid | 3.18 | 1.60 | 5.20 | 1.79 | / | / | |
| 香叶基丙酮 Geranylacetone | 4.55 | 1.66 | 5.13 | 1.07 | / | / | |
| 二十二碳六烯酸 Docosahexaenoic acid | 4.93 | 1.00 | 3.80 | 1.17 | / | / | |
| 20-羟基蜕皮酮 20-Hydroxyecdysone | 4.05 | 1.43 | 4.88 | 1.14 | / | / | |
| 二氢玫瑰苷 Dihydroroseoside | 4.58 | 1.84 | 4.67 | 1.12 | / | / | |
| 茯苓酸B Poricoic acid B | / | / | 4.67 | 1.01 | 3.84 | 1.52/ | |
| 草乌甲素A Bulleyaconi cine A | / | / | -3.66 | 1.18 | -4.85 | 2.10 | |
| 拉帕醇 Lapachol | -2.75 | 1.07 | -3.81 | 2.68 | / | / | |
| 丹参酚C Danshenol C | / | / | -4.06 | 1.54 | -4.14 | 1.74 | |
| 桦木酮酸 Betulonic acid | 2.34 | 1.17 | / | / | -4.32 | 2.24 | |
| 乙酰哈帕苷 Acetylharpagide | / | / | -3.59 | 1.47 | -4.41 | 1.06 | |
| 毒毛旋花苷 Ouabain | / | / | -4.54 | 1.38 | -4.34 | 1.66 | |
| 竹节参皂苷 IVa Chikusetsusponin IVa | 3.69 | 1.67 | -1.98 | 1.69 | -4.80 | 1.10 | |
| 8-异前列腺素A2 8-iso Prostaglandin A2 | / | / | -4.86 | 1.73 | -5.02 | 1.72 | |
| 马鞭草苷 Verbenalin | 1.16 | 1.04 | -5.09 | 1.15 | -6.25 | 2.02 | |
| 亚油酸 Linoleic acid | / | / | -4.58 | 1.53 | -5.61 | 2.07 | |
| 玫瑰苷 Roseoside | / | / | 4.67 | 1.12 | -6.34 | 1.30 | |
| (±)-脱落酸 (±)-Abscisic acid | / | / | -6.28 | 1.49 | -6.46 | 1.47 | |
| 苯丙类和聚酮类 Phenylpropanoids and polyketides | 槲皮素-3β-D-葡萄糖苷 Quercetin-3β-D-glucoside | -7.65 | 1.84 | / | / | 8.26 | 1.31 |
| 山奈酚 Kaempferol | / | / | 7.27 | 1.63 | 6.60 | 1.29 | |
| 荭草素 Orientin | -1.97 | 1.13 | 3.84 | 1.13 | 5.81 | 1.84 | |
| 矢车菊素-3-O-葡萄糖苷 Kuromanin | / | / | 5.24 | 1.33 | 5.15 | 1.74 | |
| 柚皮素 Naringenin | -3.70 | 1.45 | / | / | 2.65 | 1.06 | |
| 咖啡酸乙酯 Ethyl caffeate | / | / | -4.13 | 1.93 | -3.10 | 1.47 | |
| 落新妇苷 Astilbin | -1.65 | 1.30 | -4.57 | 1.48 | / | / | |
| 异茴芹素 Isopimpinellin | -4.71 | 2.03 | / | / | 4.46 | 1.00 | |
| 曲克芦丁 Troxerutin | -5.37 | 1.48 | -5.72 | 1.61 | / | / | |
| 车前苷 Plantagoside | -6.19 | 1.28 | / | / | 3.64 | 1.32 | |
| 毛蕊花糖苷 Verbascoside | / | / | -6.41 | 1.32 | -4.20 | 1.54 | |
| 槲皮素3-O-槐糖苷 Quercetin 3-O-sophoroside | -9.93 | 1.20 | / | / | 9.64 | 1.91 | |
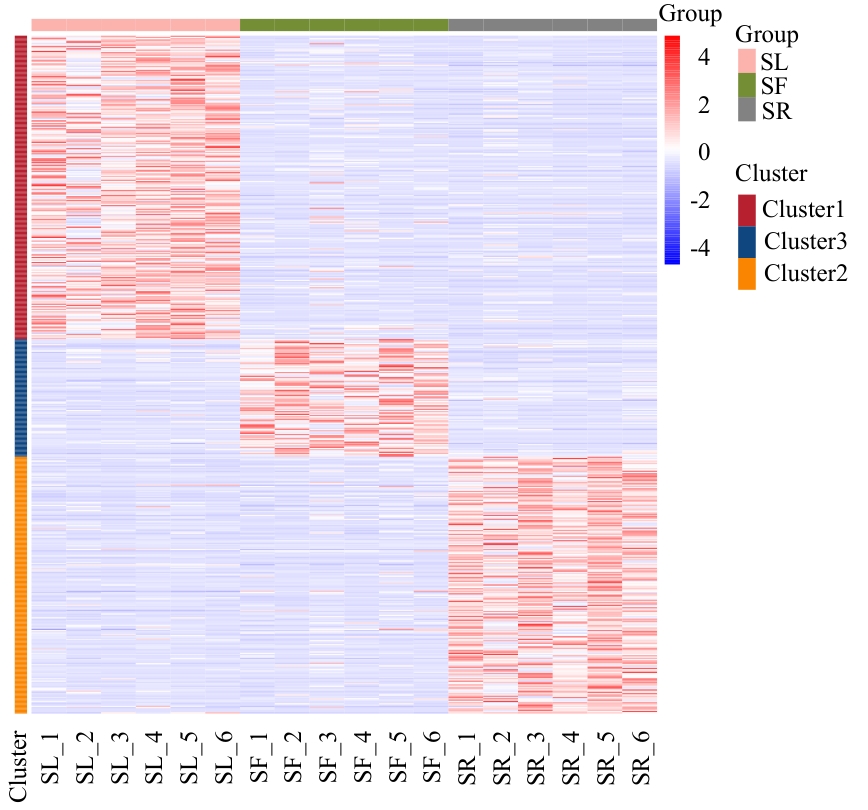
图2 差异代谢物聚类热图红色代表上调,蓝色代表下调。下同
Fig. 2 Clustering heatmap of differential metabolitesRed indicates up-regulation, and blue indicates down-regulation. The same below
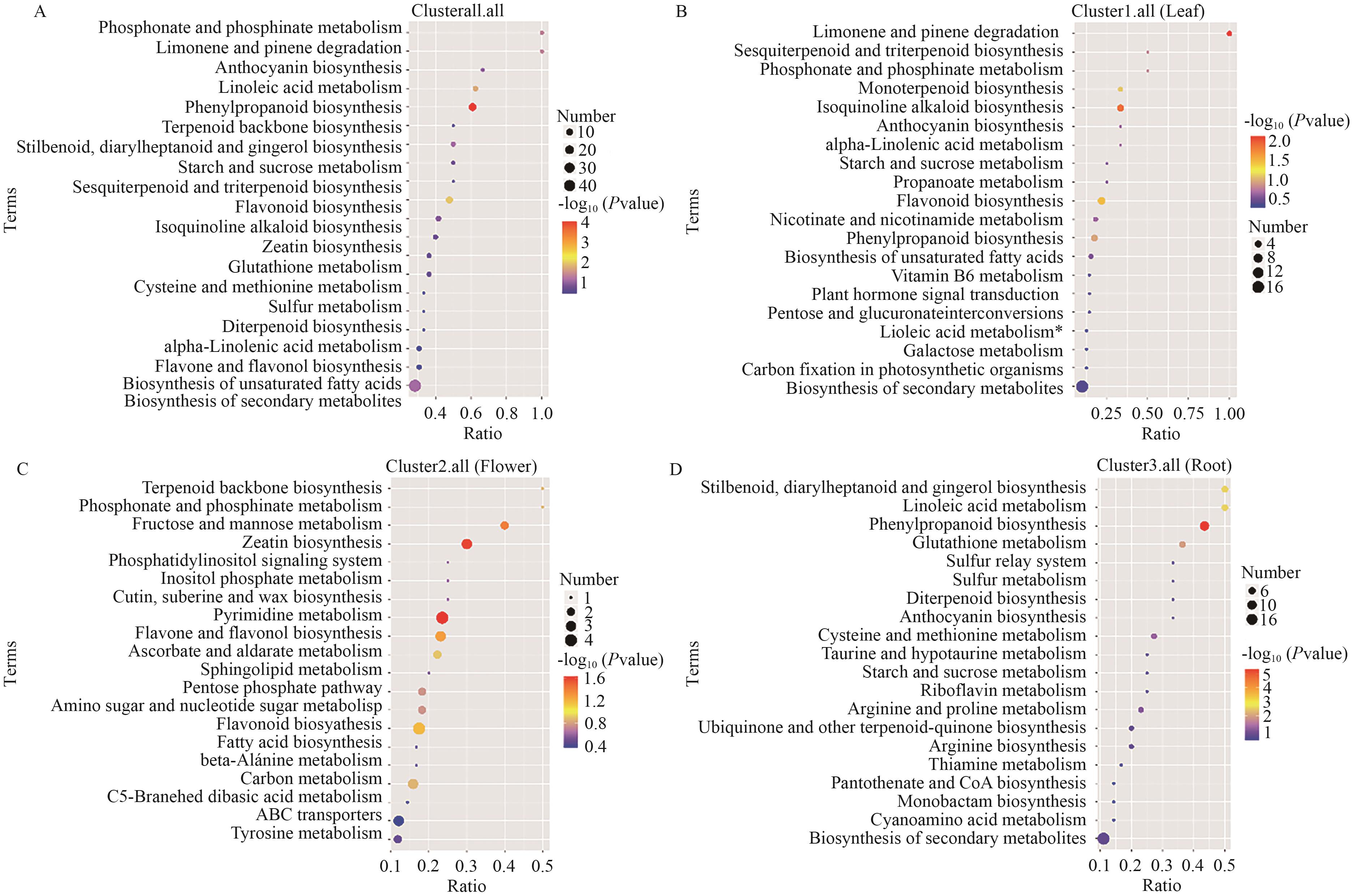
图3 非靶向KEGG富集气泡图差异代谢物显著富集的生物合成通路,A:所有部位;B:叶片;C:花朵;D:根部
Fig. 3 Non-targeted KEGG enrichment bubble chartBiosynthetic pathways significantly enriched with differential metabolites. A: All parts. B: Leaves. C: Flowers. D: Roots
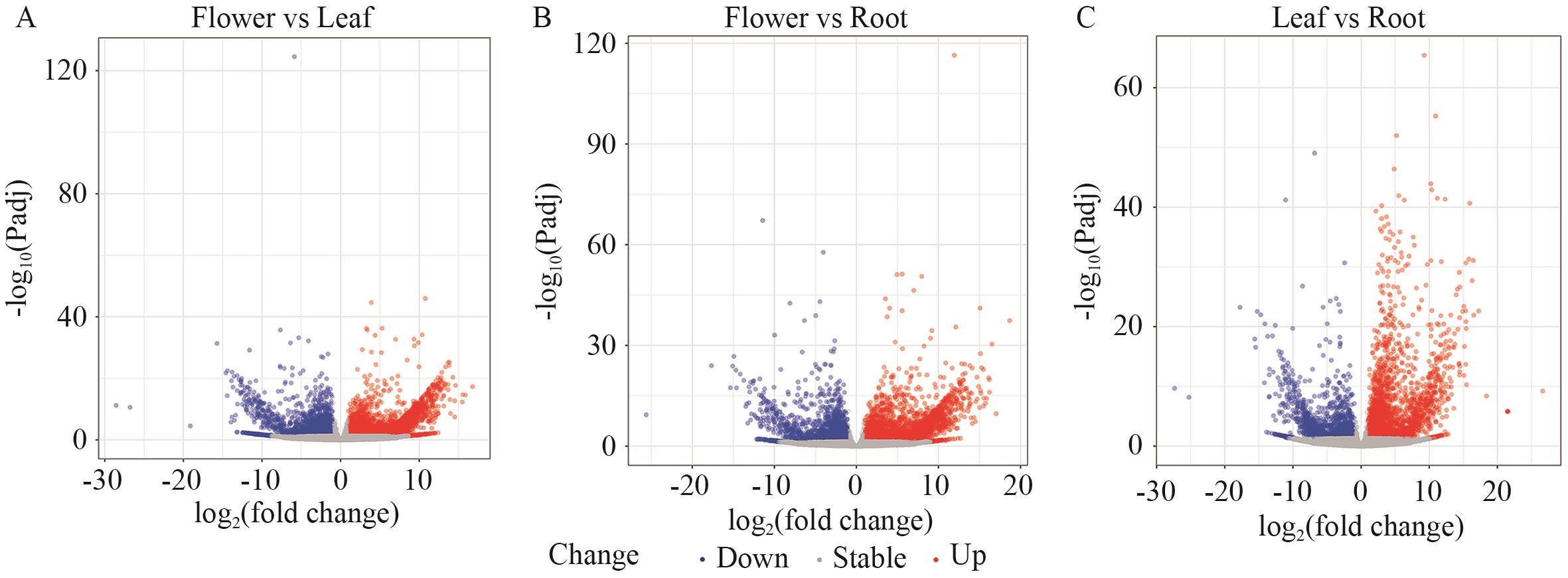
图6 紫纹兜兰不同部位差异表达基因火山图A: SF vs SL; B: SF vs SR; C: SL vs SR
Fig. 6 Volcano plot of differentially expressed genes (DEGs) in different parts of P. purpuratum
| 比较组 Comparison Group | 调控方向 Up/Down | 基因ID Gene ID | Log2FC | FDR | 功能注释 Functional annotation |
|---|---|---|---|---|---|
| SL vs. SF | 上调 Up | GLDC | 15.72 | 4.40e-32 | 碳代谢 Carbon metabolism |
| 上调 Up | LOX1_5 | 14.59 | 1.74e-22 | 脂质代谢和亚油酸代谢 Lipid metabolism and linoleic acid metabolism | |
| 上调 Up | chlH | 14.42 | 4.33e-15 | 叶绿素合成 Chlorophyll synthesis | |
| 上调 Up | cynT | 13.94 | 1.61e-6 | 氮代谢 Nitrogen metabolism | |
| 上调 Up | PMA1/PMA2 | 13.51 | 4.38e-7 | 能量代谢 Energy metabolism | |
| 下调 Down | GPAT | -10.00 | 1.14e-10 | 脂质代谢 Lipid metabolism | |
| 下调 Down | P5CS | -9.98 | 1.37e-11 | 氨基酸代谢 Amino acid metabolism | |
| 下调 Down | pdhC | -9.95 | 1.23e-7 | 碳代谢 Carbon metabolism | |
| 下调 Down | ALDH | -9.92 | 2.47e-9 | 新陈代谢 Metabolism | |
| 下调 Down | PIP5K | -9.85 | 5.33e-12 | 信号转导 Signal transduction | |
| SL vs. SR | 上调 Up | nadB | 9.99 | 3.65e-12 | 新陈代谢 Metabolism |
| 上调 Up | ALDO | 9.30 | 1.34e-07 | 新陈代谢 Metabolism | |
| 下调 Down | E1.11.1.7 | -14.72 | 1.04e-22 | 次生代谢物的生物合成 Biosynthesis of secondary metabolites | |
| 下调 Down | CISZOG | -14.18 | 2.63e-11 | 萜类和多酮类化合物的代谢 Terpenoid and polyketide metabolism | |
| 下调 Down | PAL | -13.46 | 4.69e-9 | 新陈代谢 Metabolism | |
| 下调 Down | TGA | -11.80 | 1.37e-7 | 植物激素信号转导 Plant hormone signal transduction | |
| SF vs. SR | 上调 Up | psbS | 10.53 | 1.04e-05 | 新陈代谢 Metabolism |
| 上调 Up | FAH | 8.10 | 1.61e-06 | 脂质代谢 Lipid metabolism | |
| 上调 Up | crtZ | 6.94 | 3.17e-03 | 次生代谢物的生物合成 Biosynthesis of secondary metabolites | |
| 下调 Down | ROMT | -15.07 | 1.49e-24 | 次生代谢物的生物合成 Biosynthesis of secondary metabolites | |
| 下调 Down | HHT1 | -14.69 | 2.42e-23 | 脂质代谢 Lipid metabolism | |
| 下调 Down | asnB | -13.24 | 1.58e-07 | 氨基酸代谢 Amino acid metabolism | |
| 下调 Down | SGT1 | -12.89 | 1.74e-08 | 聚糖的生物合成与代谢 Glycan biosynthesis and metabolism | |
| 下调 Down | LOX1_5 | -8.82 | 5.84e-09 | 脂质代谢和亚油酸代谢 Lipid metabolism and linoleic acid metabolism |
表3 紫纹兜兰不同部位主要显著差异基因信息
Table 3 Main significant differential gene information in different parts of P. purpuratum
| 比较组 Comparison Group | 调控方向 Up/Down | 基因ID Gene ID | Log2FC | FDR | 功能注释 Functional annotation |
|---|---|---|---|---|---|
| SL vs. SF | 上调 Up | GLDC | 15.72 | 4.40e-32 | 碳代谢 Carbon metabolism |
| 上调 Up | LOX1_5 | 14.59 | 1.74e-22 | 脂质代谢和亚油酸代谢 Lipid metabolism and linoleic acid metabolism | |
| 上调 Up | chlH | 14.42 | 4.33e-15 | 叶绿素合成 Chlorophyll synthesis | |
| 上调 Up | cynT | 13.94 | 1.61e-6 | 氮代谢 Nitrogen metabolism | |
| 上调 Up | PMA1/PMA2 | 13.51 | 4.38e-7 | 能量代谢 Energy metabolism | |
| 下调 Down | GPAT | -10.00 | 1.14e-10 | 脂质代谢 Lipid metabolism | |
| 下调 Down | P5CS | -9.98 | 1.37e-11 | 氨基酸代谢 Amino acid metabolism | |
| 下调 Down | pdhC | -9.95 | 1.23e-7 | 碳代谢 Carbon metabolism | |
| 下调 Down | ALDH | -9.92 | 2.47e-9 | 新陈代谢 Metabolism | |
| 下调 Down | PIP5K | -9.85 | 5.33e-12 | 信号转导 Signal transduction | |
| SL vs. SR | 上调 Up | nadB | 9.99 | 3.65e-12 | 新陈代谢 Metabolism |
| 上调 Up | ALDO | 9.30 | 1.34e-07 | 新陈代谢 Metabolism | |
| 下调 Down | E1.11.1.7 | -14.72 | 1.04e-22 | 次生代谢物的生物合成 Biosynthesis of secondary metabolites | |
| 下调 Down | CISZOG | -14.18 | 2.63e-11 | 萜类和多酮类化合物的代谢 Terpenoid and polyketide metabolism | |
| 下调 Down | PAL | -13.46 | 4.69e-9 | 新陈代谢 Metabolism | |
| 下调 Down | TGA | -11.80 | 1.37e-7 | 植物激素信号转导 Plant hormone signal transduction | |
| SF vs. SR | 上调 Up | psbS | 10.53 | 1.04e-05 | 新陈代谢 Metabolism |
| 上调 Up | FAH | 8.10 | 1.61e-06 | 脂质代谢 Lipid metabolism | |
| 上调 Up | crtZ | 6.94 | 3.17e-03 | 次生代谢物的生物合成 Biosynthesis of secondary metabolites | |
| 下调 Down | ROMT | -15.07 | 1.49e-24 | 次生代谢物的生物合成 Biosynthesis of secondary metabolites | |
| 下调 Down | HHT1 | -14.69 | 2.42e-23 | 脂质代谢 Lipid metabolism | |
| 下调 Down | asnB | -13.24 | 1.58e-07 | 氨基酸代谢 Amino acid metabolism | |
| 下调 Down | SGT1 | -12.89 | 1.74e-08 | 聚糖的生物合成与代谢 Glycan biosynthesis and metabolism | |
| 下调 Down | LOX1_5 | -8.82 | 5.84e-09 | 脂质代谢和亚油酸代谢 Lipid metabolism and linoleic acid metabolism |
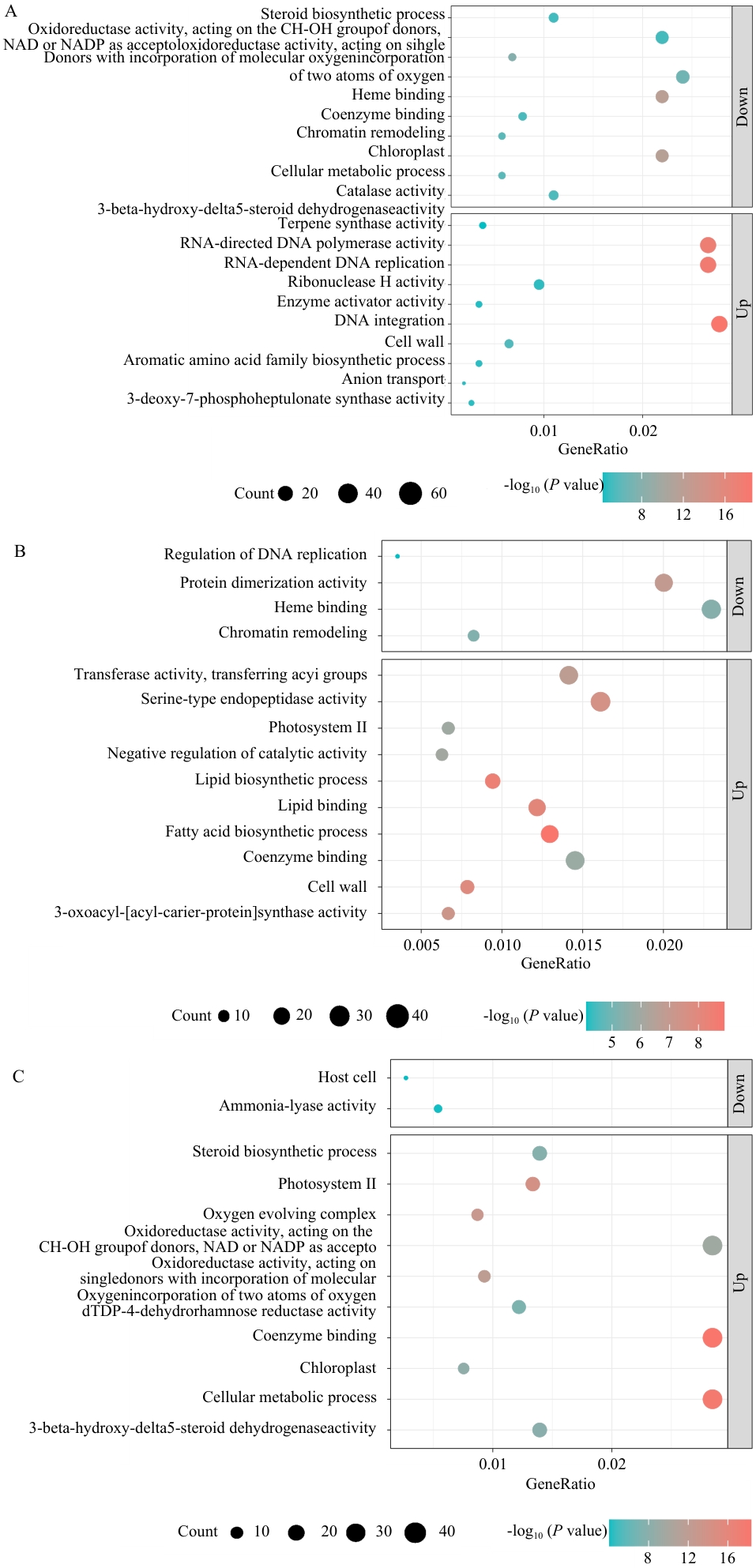
图7 紫纹兜兰不同部位差异表达基因GO功能富集分析A: SF vs. SL; B: SF vs. SR; C: SL vs. SR
Fig. 7 GO functional enrichment analysis of DEGs in different parts of P. purpuratum
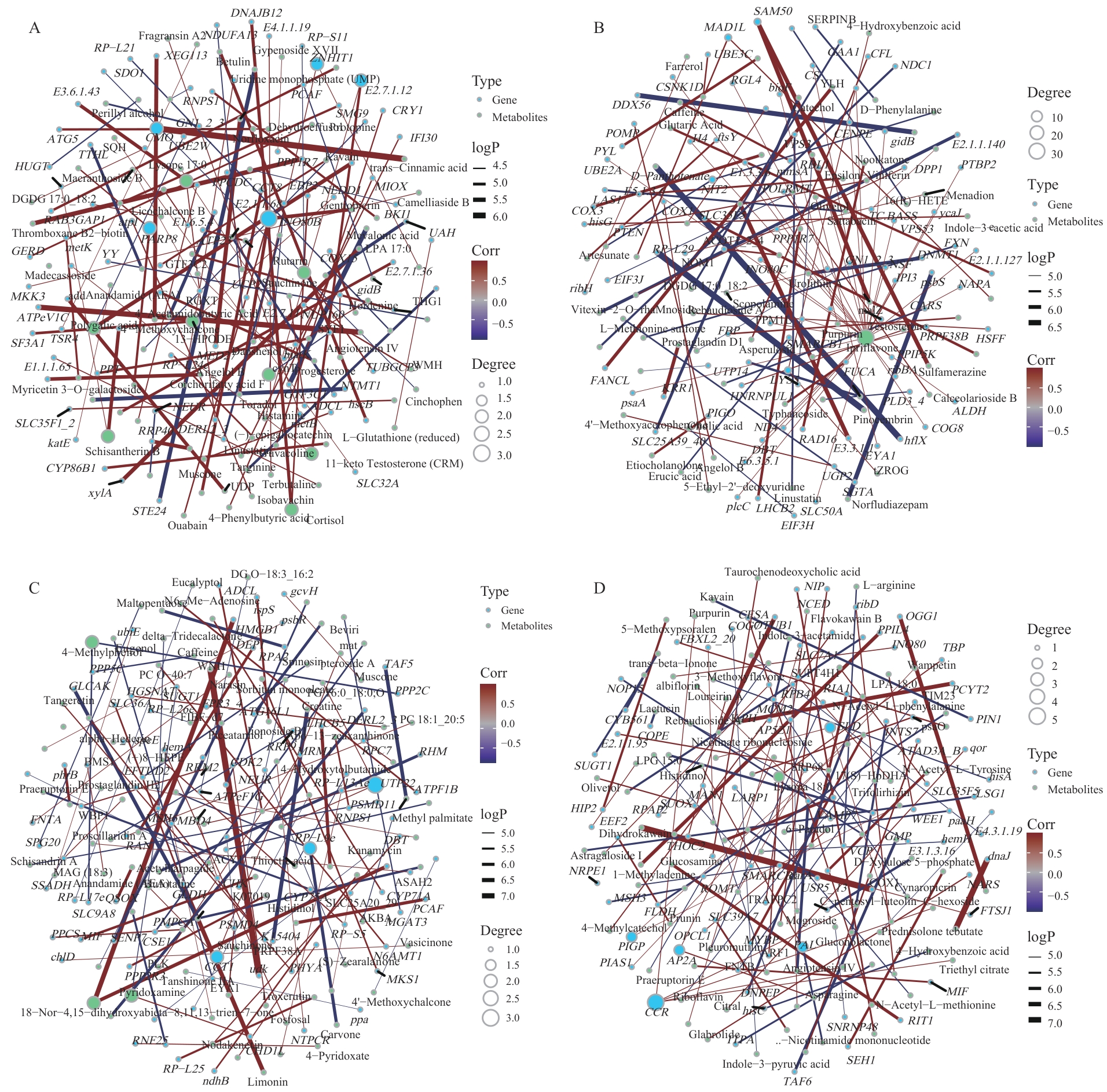
图9 紫纹兜兰不同部位代谢组和转录组的相关性分析A:所有部位;B:叶片;C:花朵;D:根部
Fig. 9 Correlation analysis between metabolome and transcriptome in different parts of P. purpuratumA: All parts. B: Leaves. C: Flowers. D: Roots
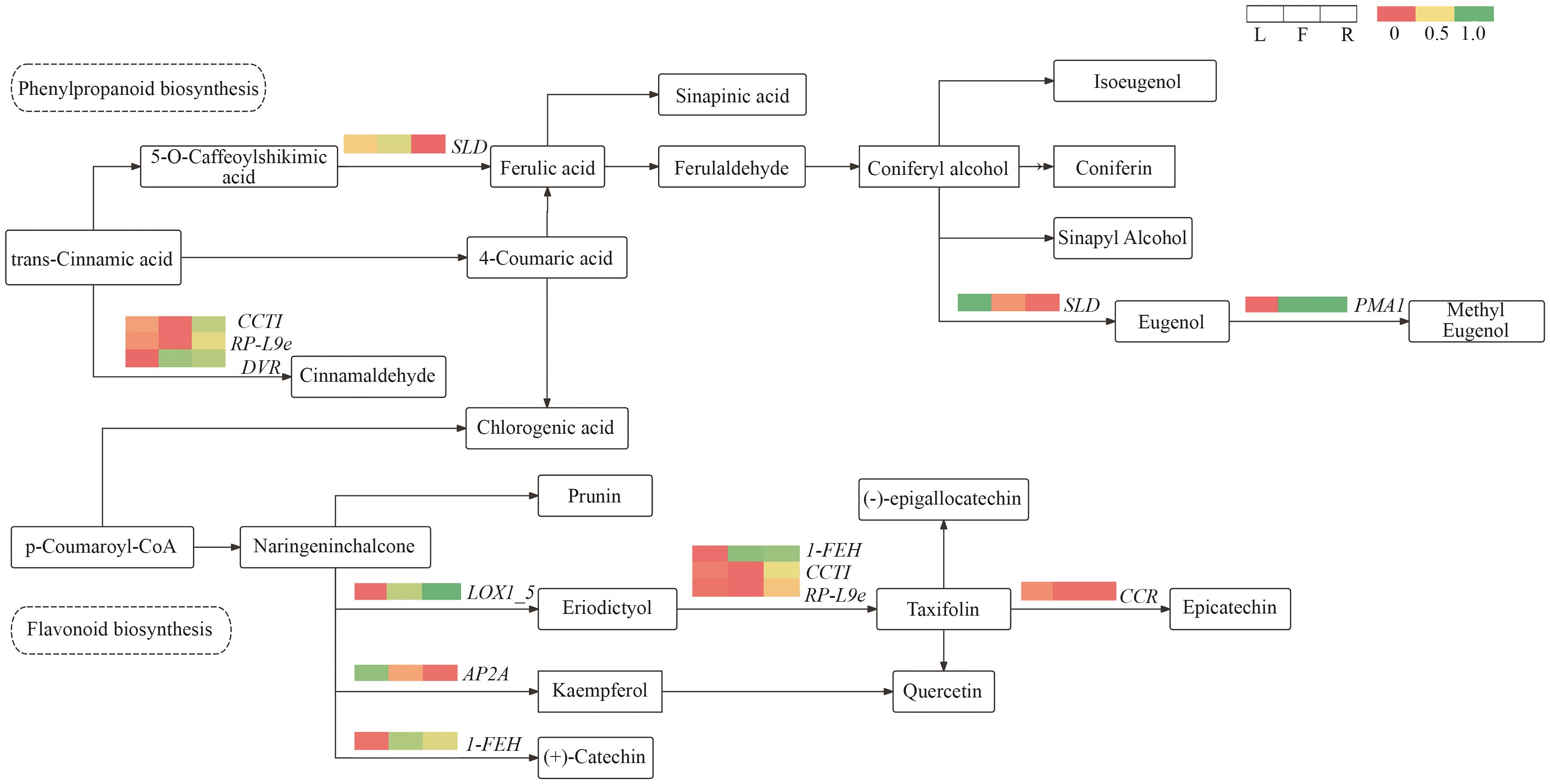
图10 共表达网络关键节点中代谢物和基因变化情况热图代表差异表达基因在不同部位中的表达显著性,红色代表高水平,绿色代表低水平
Fig. 10 Changes in metabolites and genes at hubs of the co-expression networkThe heatmap indicates the expressions of differentially expressed genes across different tissues. Red color indicates high expression level, and green color indicates low expression level
| [1] | 罗毅波, 贾建生, 王春玲. 初论中国兜兰属植物的保护策略及其潜在资源优势 [J]. 生物多样性, 2003, 11(6): 491-498. |
| Luo YB, Jia JS, Wang CL. Conservation strategy and potential advantages of the Chinese Paphiopedilum [J]. Chin Biodivers, 2003, 11(6): 491-498. | |
| [2] | 刘仲健, 陈心启, 陈利君, 等. 中国兜兰属植物 [M]. 北京: 科学出版社, 2009. |
| Liu ZJ, Chen XQ, Chen LJ, et al. The genus Paphiopedilum in China [M]. Beijing: Science Press, 2009. | |
| [3] | Sun JJ, Li QL, Xu H, et al. Analysis of metabolomic changes in xylem and phloem sap of cucumber under phosphorus stresses [J]. Metabolites, 2022, 12(4): 361. |
| [4] | Zhou ZW, Luo MD, Chen X, et al. Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics [J]. Nat Commun, 2020, 11(1): 4334. |
| [5] | Guo AH, Yang Y, Wu J, et al. Lipidomic and transcriptomic profiles of glycerophospholipid metabolism during Hemerocallis citrina Baroni flowering [J]. BMC Plant Biol, 2023, 23(1): 50. |
| [6] | Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics [J]. Nat Rev Genet, 2009, 10(1): 57-63. |
| [7] | Zhao RX, Yan S, Hu YD, et al. Metabolic and transcriptomic profile revealing the differential accumulating mechanism in different parts of Dendrobium nobile [J]. Int J Mol Sci, 2024, 25(10): 5356. |
| [8] | Qiu YJ, Cai CC, Mo X, et al. Transcriptome and metabolome analysis reveals the effect of flavonoids on flower color variation in Dendrobium nobile Lindl [J]. Front Plant Sci, 2023, 14: 1220507. |
| [9] | Ahmad S, Lu CQ, Gao J, et al. Integrated proteomic, transcriptomic, and metabolomic profiling reveals that the gibberellin-abscisic acid hub runs flower development in the Chinese orchid Cymbidium sinense [J]. Hortic Res, 2024, 11(5): uhae073. |
| [10] | Zhang GJ, Hu Y, Huang MZ, et al. Comprehensive phylogenetic analyses of Orchidaceae using nuclear genes and evolutionary insights into epiphytism [J]. J Integr Plant Biol, 2023, 65(5): 1204-1225. |
| [11] | 王蒙, 王婷, 夏增强, 等. 基于转录组数据揭示4种兜兰的全基因组复制历史 [J]. 植物学报, 2021, 56(6): 699-714. |
| Wang M, Wang T, Xia ZQ, et al. Revealing the new whole-genome duplication event of four Paphiopedilum species based on transcriptome data [J]. Chin Bull Bot, 2021, 56(6): 699-714. | |
| [12] | Liang YY, Hao J, Wang JY, et al. Statistical genomics analysis of simple sequence repeats from the Paphiopedilum malipoense transcriptome reveals control knob motifs modulating gene expression [J]. Adv Sci, 2024, 11(24): 2304848. |
| [13] | Ye YQ, Chang YT, Ma YJ, et al. Comparative study on the mechanism of yellow petal formation in Paphiopedilum armeniacum: an integrated transcriptomic and metabolomic analysis of three Paphiopedilum species [J]. BMC Genomics, 2025, 26(1): 560. |
| [14] | Wen B, Mei ZL, Zeng CW, et al. metaX: a flexible and comprehensive software for processing metabolomics data [J]. BMC Bioinformatics, 2017, 18(1): 183. |
| [15] | Haspel JA, Chettimada S, Shaik RS, et al. Circadian rhythm reprogramming during lung inflammation [J]. Nat Commun, 2014, 5: 4753. |
| [16] | Sreekumar A, Poisson LM, Rajendiran TM, et al. Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression [J]. Nature, 2009, 457(7231): 910-914. |
| [17] | Chen SF, Zhou YQ, Chen YR, et al. Fastp: an ultra-fast all-in-one FASTQ preprocessor [J]. Bioinformatics, 2018, 34(17): i884-i890. |
| [18] | Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data [J]. Bioinformatics, 2014, 30(15): 2114-2120. |
| [19] | Patro R, Duggal G, Love MI, et al. Salmon provides fast and bias-aware quantification of transcript expression [J]. Nat Meth, 2017, 14(4): 417-419. |
| [20] | Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2 [J]. Genome Biol, 2014, 15(12): 550. |
| [21] | Yu GC, Wang LG, Han YY, et al. clusterProfiler: an R package for comparing biological themes among gene clusters [J]. OMICS, 2012, 16(5): 284-287. |
| [22] | Fan JW, Shen YT, Chen C, et al. A large-scale integrated transcriptomic atlas for soybean organ development [J]. Mol Plant, 2025, 18(4): 669-689. |
| [23] | 孙晓琛, 栗锦鹏, 原静静, 等. 基于转录组测序分析干旱胁迫对党参不同组织基因表达的调控 [J]. 中草药, 2022, 53(14): 4465-4475. |
| Sun XC, Li JP, Yuan JJ, et al. Sequencing and analysis of transcriptome to reveal regulation gene expression in different tissues of Codonopsis pilosula under drought stress [J]. Chin Tradit Herb Drugs, 2022, 53(14): 4465-4475. | |
| [24] | Zhao XC, Wei JP, He L, et al. Identification of fatty acid desaturases in maize and their differential responses to low and high temperature [J]. Genes, 2019, 10(6): 445. |
| [25] | Qiao Q, Wu C, Cheng TT, et al. Comparative analysis of the metabolome and transcriptome between the green and yellow-green regions of variegated leaves in a mutant variety of the tree species Pteroceltis tatarinowii [J]. Int J Mol Sci, 2022, 23(9): 4950. |
| [26] | Bao YR, Nie TK, Wang DD, et al. Anthocyanin regulatory networks in Solanum tuberosum L. leaves elucidated via integrated metabolomics, transcriptomics, and StAN1 overexpression [J]. BMC Plant Biol, 2022, 22(1): 228. |
| [27] | Ghasemzadeh A, Jaafar HZE, Rahmat A. Synthesis of phenolics and flavonoids in ginger (Zingiber officinale Roscoe) and their effects on photosynthesis rate [J]. Int J Mol Sci, 2010, 11(11): 4539-4555. |
| [28] | Hong JY, Gunasekara C, He C, et al. Identification of biological pathway and process regulators using sparse partial least squares and triple-gene mutual interaction [J]. Sci Rep, 2021, 11(1): 13174. |
| [29] | He XJ, Zhao XC, Gao LP, et al. Isolation and characterization of key genes that promote flavonoid accumulation in purple-leaf tea (Camellia sinensis L.) [J]. Sci Rep, 2018, 8(1): 130. |
| [30] | Luo JR, Duan JJ, Huo D, et al. Transcriptomic analysis reveals transcription factors related to leaf anthocyanin biosynthesis in Paeonia qiui [J]. Molecules, 2017, 22(12): 2186. |
| [31] | Xiang P, Zhu QF, Tukhvatshin M, et al. Light control of catechin accumulation is mediated by photosynthetic capacity in tea plant (Camellia sinensis) [J]. BMC Plant Biol, 2021, 21(1): 478. |
| [32] | 陈伟, 顿春垚, 李双龙, 等. 不同油茶品种间儿茶素含量差异及油茶炭疽病关系 [J]. 山东农业大学学报: 自然科学版, 2024, 55(5): 733-739. |
| Chen W, Dun CY, Li SL, et al. Differences in catechin content among different Camellia oleifera varieties and connection [J]. J Shandong Agric Univ Nat Sci Ed, 2024, 55(5): 733-739. | |
| [33] | Sohlenkamp C, Geiger O. Bacterial membrane lipids: diversity in structures and pathways [J]. FEMS Microbiol Rev, 2016, 40(1): 133-159. |
| [34] | Du K, Jiang SX, Chen H, et al. Spatiotemporal miRNA and transcriptomic network dynamically regulate the developmental and senescence processes of poplar leaves [J]. Hortic Res, 2023, 10(10): uhad186. |
| [35] | Dun HF, Hung TH, Green S, et al. Comparative transcriptomic responses of European and Japanese larches to infection by Phytophthora ramorum [J]. BMC Plant Biol, 2022, 22(1): 480. |
| [36] | Wang J, Cao K, Wang LR, et al. Two MYB and three bHLH family genes participate in anthocyanin accumulation in the flesh of peach fruit treated with glucose, sucrose, sorbitol, and fructose in vitro [J]. Plants, 2022, 11(4): 507. |
| [37] | Peng YY, Thrimawithana AH, Cooney JM, et al. The proanthocyanin-related transcription factors MYBC1 and WRKY44 regulate branch points in the kiwifruit anthocyanin pathway [J]. Sci Rep, 2020, 10(1): 14161. |
| [38] | Tu MX, Fang JH, Zhao RK, et al. CRISPR/Cas9-mediated mutagenesis of VvbZIP36 promotes anthocyanin accumulation in grapevine (Vitis vinifera) [J]. Hortic Res, 2022, 9: uhac022. |
| [39] | Mbanjo EGN, Kretzschmar T, Jones H, et al. The genetic basis and nutritional benefits of pigmented rice grain [J]. Front Genet, 2020, 11: 229. |
| [40] | Hong H, Seo H, Park W, et al. Sequence, structure and function-based classification of the broadly conserved FAH superfamily reveals two distinct fumarylpyruvate hydrolase subfamilies [J]. Environ Microbiol, 2020, 22(1): 270-285. |
| [41] | Athmika, Ghate SD, Arun AB, et al. Genome analysis of a halophilic bacterium Halomonas malpeensis YU-PRIM-29(T) reveals its exopolysaccharide and pigment producing capabilities [J]. Sci Rep, 2021, 11(1): 1749. |
| [42] | Iwasaka H, Koyanagi R, Satoh R, et al. A possible trifunctional β-carotene synthase gene identified in the draft genome of Aurantiochytrium sp. strain KH105 [J]. Genes, 2018, 9(4): 200. |
| [43] | Chuang YC, Lee MC, Chang YL, et al. Diurnal regulation of the floral scent emission by light and circadian rhythm in the Phalaenopsis orchids [J]. Bot Stud, 2017, 58(1): 50. |
| [44] | Jin JJ, Kim MJ, Dhandapani S, et al. The floral transcriptome of ylang ylang (Cananga odorata var. fruticosa) uncovers biosynthetic pathways for volatile organic compounds and a multifunctional and novel sesquiterpene synthase [J]. J Exp Bot, 2015, 66(13): 3959-3975. |
| [45] | Krug C, Cordeiro GD, Schäffler I, et al. Nocturnal bee pollinators are attracted to guarana flowers by their scents [J]. Front Plant Sci, 2018, 9: 1072. |
| [46] | Wu D, Yu L, Nair MG, et al. Cyclooxygenase enzyme inhibitory compounds with antioxidant activities from Piper methysticum (kava kava) roots [J]. Phytomedicine, 2002, 9(1): 41-47. |
| [47] | Gakière B, Hao JF, de Bont L, et al. NAD+ biosynthesis and signaling in plants [J]. Crit Rev Plant Sci, 2018, 37(4): 259-307. |
| [48] | Zhu FD, Fu X, Ye HC, et al. Antibacterial activities of coumarin-3-carboxylic acid against Acidovorax citrulli [J]. Front Microbiol, 2023, 14: 1207125. |
| [49] | 陈荷莹, 刘毅, 刘会珍, 等. 落新妇苷的稳定性及生物活性研究进展 [J]. 世界中医药, 2023, 18(11): 1609-1614. |
| Chen HY, Liu Y, Liu HZ, et al. Research progress on the stability and biological activity of astilbin [J]. World Chin Med, 2023, 18(11): 1609-1614. | |
| [50] | Cao Y, Du PH, Zhang JR, et al. Dopamine alleviates cadmium stress in apple trees by recruiting beneficial microorganisms to enhance the physiological resilience revealed by high-throughput sequencing and soil metabolomics [J]. Hortic Res, 2023, 10(7): uhad112. |
| [51] | Singh AK, Dhanapal S, Yadav BS. The dynamic responses of plant physiology and metabolism during environmental stress progression [J]. Mol Biol Rep, 2020, 47(2): 1459-1470. |
| [52] | Li WL, Lee J, Yu S, et al. Characterization and analysis of the transcriptome response to drought in Larix kaempferi using PacBio full-length cDNA sequencing integrated with de novo RNA-seq reads [J]. Planta, 2021, 253(2): 28. |
| [53] | Zou S, Lu YC, Ma HY, et al. Microalgal glycerol-3-phosphate acyltransferase role in galactolipids and high-value storage lipid biosynthesis [J]. Plant Physiol, 2023, 192(1): 426-441. |
| [54] | Li R, Zhao Y, Sun Z, et al. Genome-wide identification of switchgrass laccases involved in lignin biosynthesis and heavy-metal responses [J]. Int J Mol Sci, 2022, 23(12): 6530. |
| [55] | Weremczuk-Jeżyna I, Hnatuszko-Konka K, Lebelt L, et al. The protective function and modification of secondary metabolite accumulation in response to light stress in Dracocephalum forrestii shoots [J]. Int J Mol Sci, 2021, 22(15): 7965. |
| [56] | Wang MN, Manzoor MA, Wang XY, et al. Comparative genomic analysis of SAUR gene family, cloning and functional characterization of two genes (PbrSAUR13 and PbrSAUR52) in Pyrus bretschneideri [J]. Int J Mol Sci, 2022, 23(13): 7054. |
| [57] | Zhang LS, Zheng LT, Wu JW, et al. OsCCRL1 is essential for phenylpropanoid metabolism in rice anthers [J]. Rice, 2023, 16(1): 10. |
| [58] | Badri DV, De-la-Peña C, Lei ZT, et al. Root secreted metabolites and proteins are involved in the early events of plant-plant recognition prior to competition [J]. PLoS One, 2012, 7(10): e46640. |
| [59] | Puig J, Pauluzzi G, Guiderdoni E, et al. Regulation of shoot and root development through mutual signaling [J]. Mol Plant, 2012, 5(5): 974-983. |
| [1] | 王玉昆, 原远, 王斌, 朱云娜, 任晓强, 任飞, 叶红. 转录组和脂质代谢组联合分析不同紫苏α-亚麻酸合成调控差异[J]. 生物技术通报, 2026, 42(4): 129-140. |
| [2] | 刘青媛, 吴洪启, 陈秀娥, 陈剑, 姜远泽, 何燕子, 喻奇伟, 刘仁祥. 转录因子NtMYB96a调控烟草耐旱性的功能研究[J]. 生物技术通报, 2026, 42(4): 239-250. |
| [3] | 杨婷, 杨宗桃, 艾静, 王禹童, 李燕烨, 邓军, 刘家勇, 赵勇, 张跃彬. 不同基因型甘蔗表型特征及根部转录组学分析[J]. 生物技术通报, 2026, 42(4): 190-201. |
| [4] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [5] | 樊荣辉, 罗远华, 陈艺荃, 方能炎, 陈燕, 钟淮钦, 叶秀仙. 文心兰‘金辉’和‘香水文心’花香形成比较分析[J]. 生物技术通报, 2026, 42(1): 105-113. |
| [6] | 刘语诗, 李镇, 邹宇琛, 汤维维, 李彬. 药用植物空间代谢组学研究进展[J]. 生物技术通报, 2025, 41(9): 22-31. |
| [7] | 刘建国, 刘格儿, 郭颖欣, 王斌, 王玉昆, 卢金凤, 黄文庭, 朱云娜. 转录组和代谢组联合解析‘桂柚1号’和‘沙田柚’果实品质差异[J]. 生物技术通报, 2025, 41(9): 168-181. |
| [8] | 刘泽洲, 段乃彬, 岳丽昕, 王清华, 姚行浩, 高莉敏, 孔素萍. 大蒜叶片蜡质成分分析及蜡质缺失基因Ggl-1筛选[J]. 生物技术通报, 2025, 41(9): 219-231. |
| [9] | 闫梦阳, 梁晓阳, 戴君昂, 张妍, 关团, 张辉, 刘良波, 孙志华. 阿莫西林降解菌的筛选及降解机制研究[J]. 生物技术通报, 2025, 41(9): 314-325. |
| [10] | 张雅祺, 王芹芹, 沈夏, 李旭苗, 高敏, 李军, 李辰, 王慧. 食管鳞状细胞癌早期进展风险的代谢物预警模型[J]. 生物技术通报, 2025, 41(9): 335-344. |
| [11] | 柴军发, 洪波, 贾彦霞. 转录组和代谢组联合分析三株蜡蚧轮枝菌菌株毒力差异[J]. 生物技术通报, 2025, 41(8): 311-321. |
| [12] | 白雨果, 李婉迪, 梁建萍, 石志勇, 卢庚龙, 刘红军, 牛景萍. 哈茨木霉T9131对黄芪幼苗的促生机理[J]. 生物技术通报, 2025, 41(8): 175-185. |
| [13] | 蒋天威, 马培杰, 李亚娇, 陈才俊, 刘晓霞, 王小利. 二穗短柄草对光周期的代谢响应分析[J]. 生物技术通报, 2025, 41(7): 237-247. |
| [14] | 段敏杰, 李怡斐, 王春萍, 黄任中, 黄启中, 张世才. 辣椒果实颜色性状与SSR分子标记的关联分析及指纹图谱构建[J]. 生物技术通报, 2025, 41(7): 81-94. |
| [15] | 张越, 毕钰, 慕雪男, 郑子薇, 王志刚, 徐伟慧. 小麦赤霉病拮抗菌JB7的生防特性[J]. 生物技术通报, 2025, 41(7): 261-271. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||