








• 微生物组学专题 • 下一篇
胡阔均1( ), 黄晓辉2, 黄谊3, 张宇宇3, 邓征宇3, 郭军3, 曾银1, 尹华群1, 周向平3, 孟德龙1(
), 黄晓辉2, 黄谊3, 张宇宇3, 邓征宇3, 郭军3, 曾银1, 尹华群1, 周向平3, 孟德龙1( )
)
收稿日期:2025-11-10
出版日期:2026-03-02
通讯作者:
孟德龙,男,博士,副教授,研究方向 :微生物学;E-mail: delong.meng@csu.edu.cn作者简介:胡阔均,男,硕士,研究方向 :微生物学;E-mail: 245611015@csu.edu.cn
基金资助:
HU Kuo-jun1( ), HUANG Xiao-hui2, HUANG Yi3, ZHANG Yu-yu3, DENG Zheng-yu3, GUO Jun3, ZENG Yin1, YIN Hua-qun1, ZHOU Xiang-ping3, MENG De-long1(
), HUANG Xiao-hui2, HUANG Yi3, ZHANG Yu-yu3, DENG Zheng-yu3, GUO Jun3, ZENG Yin1, YIN Hua-qun1, ZHOU Xiang-ping3, MENG De-long1( )
)
Received:2025-11-10
Published:2026-03-02
摘要:
目的 针对烟草连作导致的土壤理化性质恶化及青枯雷尔氏菌(Ralstonia solanacearum)引起的青枯病加剧问题,探讨大球盖菇(Stropharia rugosoannulata)基质对烟草根际土壤微生态及病害发生的调控机制。 方法 采用盆栽试验,以云烟87为供试品种,利用连作患病土壤设置对照组(CK)与添加0.5 kg大球盖菇基质的处理组(BK),移栽45 d后调查烟草病情指数,测定根际土壤理化性质,并利用宏基因组测序技术,分析微生物群落结构与其代谢功能通路的差异。 结果 大球盖菇基质处理显著改变了土壤理化性质,与对照组相比,处理组土壤速效磷、有机质含量显著提高(P<0.01),氮含量显著增加(P<0.05),而速效钾含量显著降低(P<0.01),并有效抑制了青枯病的发生(P<0.05)。微生物群落分析显示,处理组根际土壤微生物Simpson多样性指数和Pielou均匀度指数显著降低。宏基因组功能分析表明,处理组在碳水化合物代谢通路上的表达显著上调(P<0.05),具体表现为果糖与甘露糖代谢、半胱氨酸与甲硫氨酸代谢、以及丙氨酸、天冬氨酸和谷氨酸代谢通路的基因丰度显著增加,包括海藻酸盐合成的关键基因(manB、alg44、algG)及氨基酸代谢基因(asdA、asnA)。 结论 大球盖菇基质通过提高土壤有机质和氮磷含量、降低速效钾含量来优化土壤理化性质;通过定向选择作用富集有益菌群,重塑根际微生态结构;同时激活果糖、甘露糖及氨基酸等关键代谢通路,抑制病原菌生物膜形成并增强植株免疫与生长,从而显著降低烟草青枯病的发病率。
胡阔均, 黄晓辉, 黄谊, 张宇宇, 邓征宇, 郭军, 曾银, 尹华群, 周向平, 孟德龙. 大球盖菇基质对烟草青枯病和土壤微生物功能的影响[J]. 生物技术通报, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1210.
HU Kuo-jun, HUANG Xiao-hui, HUANG Yi, ZHANG Yu-yu, DENG Zheng-yu, GUO Jun, ZENG Yin, YIN Hua-qun, ZHOU Xiang-ping, MENG De-long. Effects of Stropharia rugosoannulata Substrate on Tobacco Bacterial Wilt and Soil Microbial Function[J]. Biotechnology Bulletin, doi: 10.13560/j.cnki.biotech.bull.1985.2025-1210.
土壤理化性质 Soil physicochemical properties | CK | BK | 显著性 Significance |
|---|---|---|---|
| 速效钾 AK/(mg/kg) | 655.00±108.70 | 281.25±34.73 | ** |
| 速效磷 AP/(mg/kg) | 308.81±28.00 | 438.73±43.32 | ** |
| 有机质 OM/(mg/kg ) | 142.60±4.73 | 155.67±2.19 | ** |
| 氮 N/(mg/kg) | 661.95±65.68 | 810.9±45.79 | * |
| 含水量 Water/% | 51.87±2.89 | 50.56±1.75 | |
| 酸碱度 pH | 7.15±0.19 | 7.43±0.1 | |
| 总钾 TK/(mg/kg) | 13.36±0.49 | 14.04±0.42 | |
| 总磷 TP/(mg/kg) | 2.44±0.18 | 2.4±0.14 | |
| 病情指数 Disease index | 61.1±9.65 | 30.6±2.25 | * |
表1 对照组与大球盖菇基质处理组土壤理化性质和病情指数
Table 1 Soil physicochemical properties and disease index in control and S. rugosoannulata substrate-treated groups
土壤理化性质 Soil physicochemical properties | CK | BK | 显著性 Significance |
|---|---|---|---|
| 速效钾 AK/(mg/kg) | 655.00±108.70 | 281.25±34.73 | ** |
| 速效磷 AP/(mg/kg) | 308.81±28.00 | 438.73±43.32 | ** |
| 有机质 OM/(mg/kg ) | 142.60±4.73 | 155.67±2.19 | ** |
| 氮 N/(mg/kg) | 661.95±65.68 | 810.9±45.79 | * |
| 含水量 Water/% | 51.87±2.89 | 50.56±1.75 | |
| 酸碱度 pH | 7.15±0.19 | 7.43±0.1 | |
| 总钾 TK/(mg/kg) | 13.36±0.49 | 14.04±0.42 | |
| 总磷 TP/(mg/kg) | 2.44±0.18 | 2.4±0.14 | |
| 病情指数 Disease index | 61.1±9.65 | 30.6±2.25 | * |
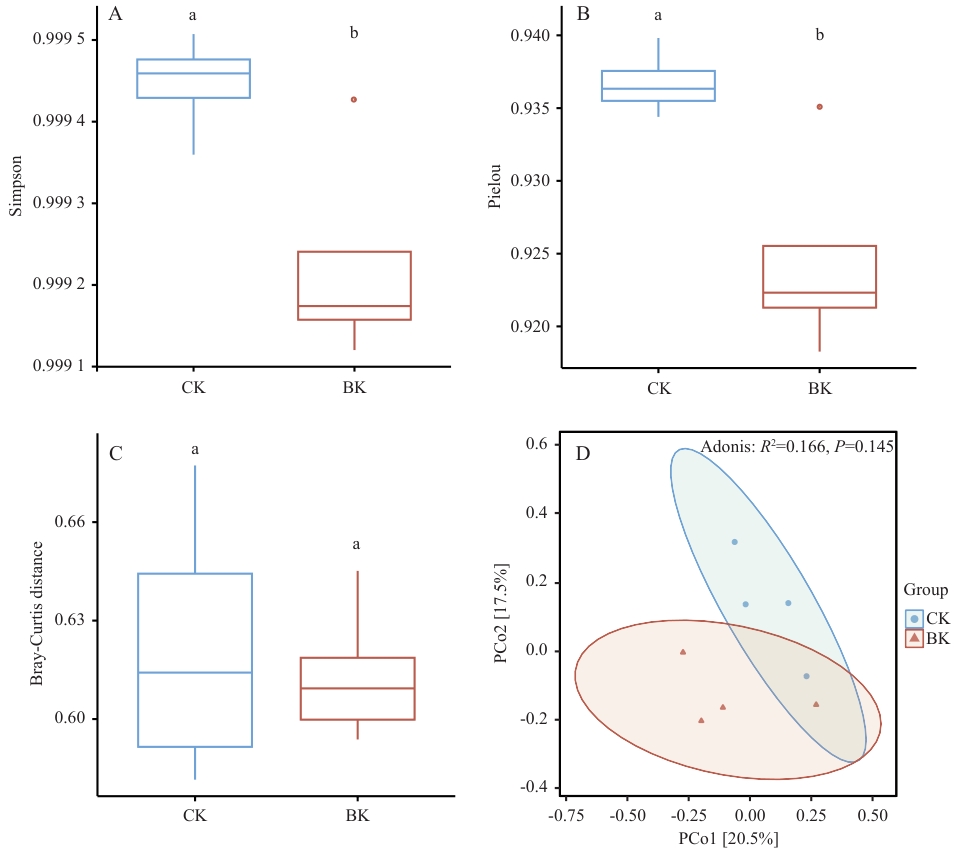
图1 对照组与大球盖菇基质处理组多样性指数A:对照组与大球盖菇基质处理组根际微生物群落Simpson多样性指数;B:对照组与大球盖菇基质处理组根际微生物群落Pielou均匀度指数;C:对照组与大球盖菇基质处理组根际微生物群落Bray-Curtis距离;D:对照组与大球盖菇基质处理组根际微生物群落主坐标分析(PCoA)图。图中不同小写字母(a、b)表示处理间差异显著(P<0.05),相同字母表示差异不显著。下同
Fig. 1 Diversity indices of the control group and S. rugosoannulata substrate treatment groupA: Simpson diversity index of the rhizosphere microbial community in the control and S. rugosoannulata substrate-treated groups. B: Pielou evenness index of the rhizosphere microbial community in the control and S. rugosoannulata substrate-treated groups. C: Bray-Curtis distance of the rhizosphere microbial community in the control and S. rugosoannulata substrate-treated groups. D: Principal coordinate analysis (PCoA) plot of the rhizosphere microbial community in the control and S. rugosoannulata substrate-treated groups. Different lowercase letters (a, b) indicate significant differences between treatments (P<0.05); the same letter indicates no significant difference. The same below
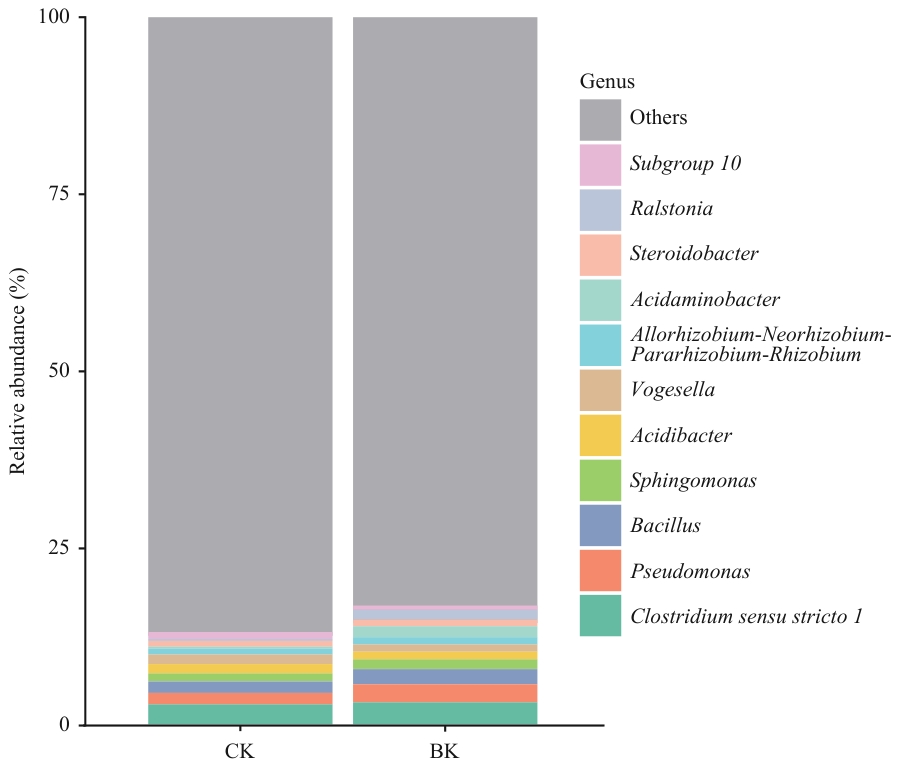
图2 对照组与大球盖菇基质处理组根际土壤微生物属水平相对丰度堆积图
Fig. 2 Stacked bar plot of relative abundance at the genus level of rhizosphere soil microbial communities in control group and S. rugosoannulata substrate treatment group
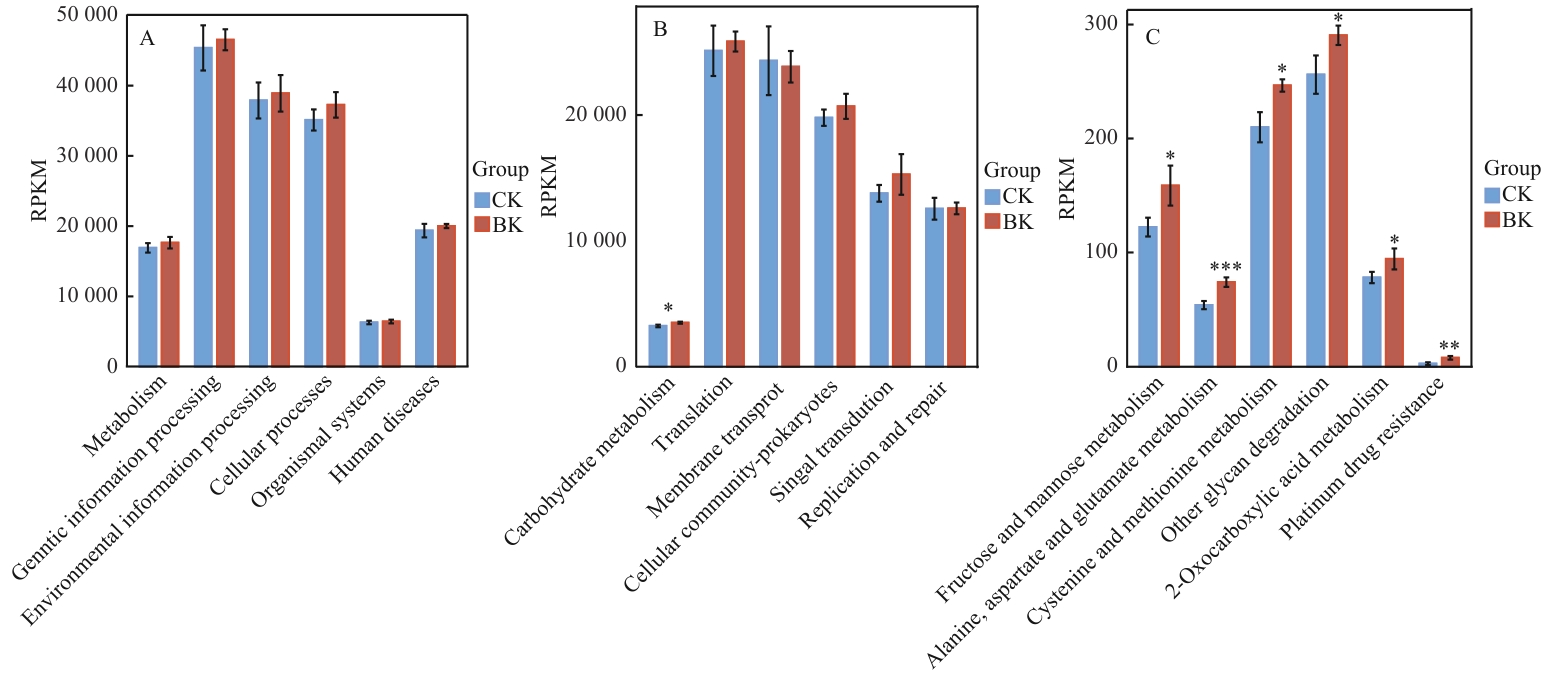
图3 对照组与大球盖菇基质处理组KEGG通路丰度汇总及通路差异分析A:对照组与大球盖菇基质处理组KEGG一级通路RPKM(reads per kilobase per million mapped reads)汇总;B:对照组与大球盖菇基质处理组KEGG二级通路RPKM汇总及差异性;C:对照组与大球盖菇基质处理组KEGG三级通路RPKM汇总及差异性
Fig. 3 KEGG pathway abundance summary and pathway differential analysis between the control group and S. rugosoannulata substrate treatment groupA: KEGG Level 1 pathway RPKM summary of control group and S. rugosoannulata substrate treatment group. B: KEGG Level 2 pathway RPKM summary and differences of control group and S. rugosoannulata substrate treatment group. C: KEGG Level 3 pathway RPKM summary and differences of control group and S. rugosoannulata substrate treatment group
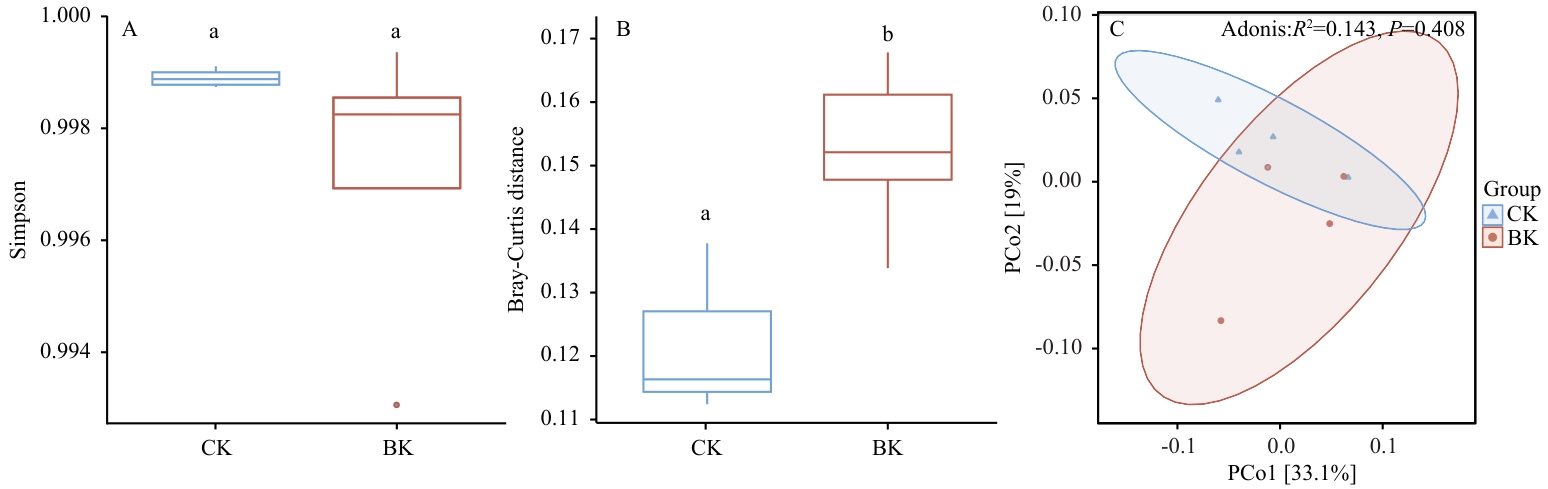
图4 对照组与大球盖菇基质处理组群落功能多样性A:对照组与大球盖菇基质处理组群落功能Simpson 多样性指数;B:对照组与大球盖菇基质处理组群落功能布雷-柯蒂斯距离;C:对照组与大球盖菇基质处理组群落功能主坐标分析图(PCoA)
Fig. 4 Community functional diversity of control group and S. rugosoannulata substrate treatment groupA: Community functional Simpson diversity index of the control group and S. rugosoannulata substrate treatment group. B: Community functional Bray-Curtis distance of the control group and S. rugosoannulata substrate treatment group. C: Community functional principal coordinate analysis (PCoA) plot of the control group and S. rugosoannulata substrate treatment group
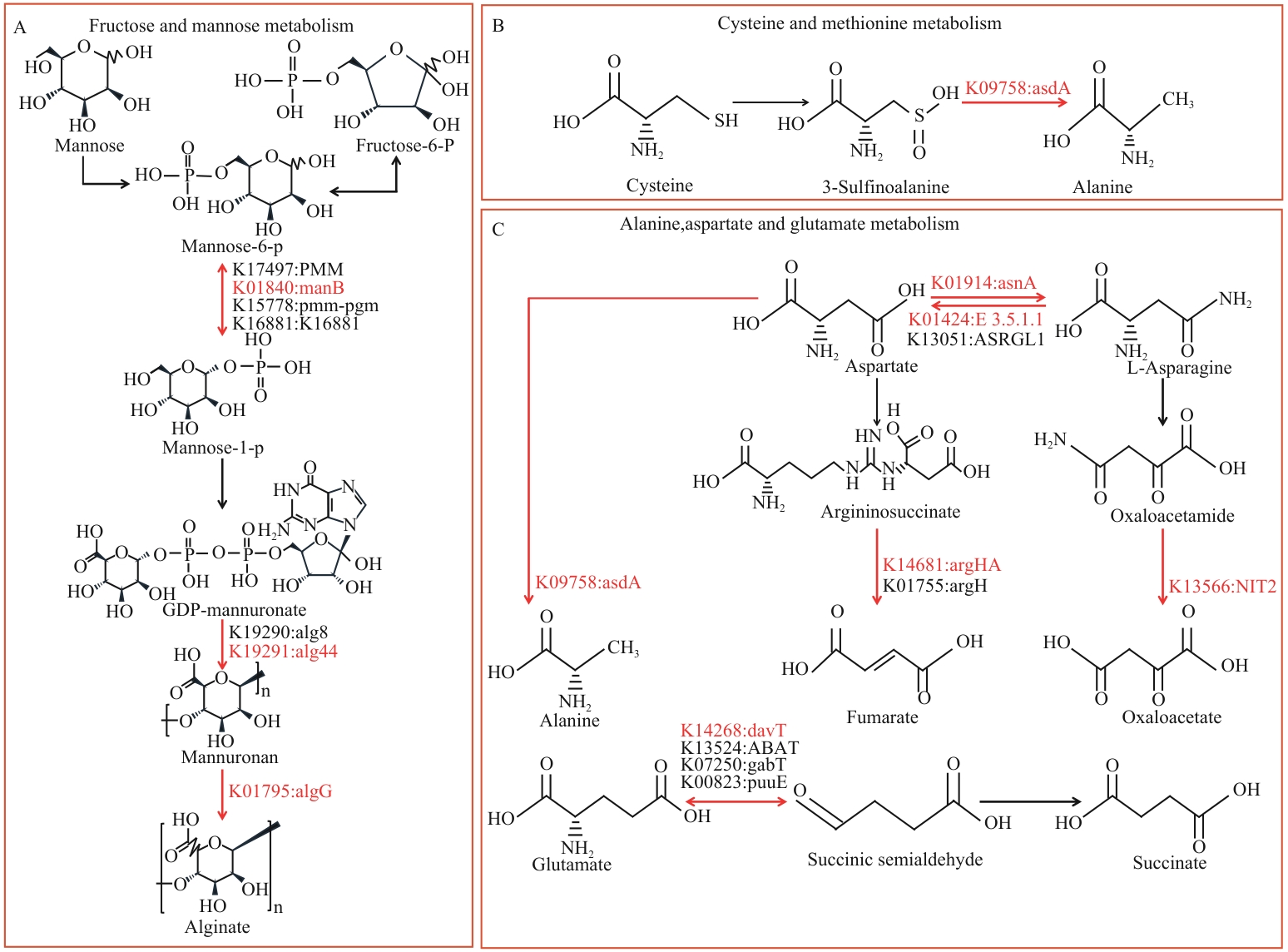
图5 果糖与甘露糖代谢通路(A)、半胱氨酸和甲硫氨酸代谢通路(B)以及丙氨酸、天冬氨酸和谷氨酸代谢通路(C)
Fig. 5 Fructose and mannose metabolism pathway (A), cysteine and methionine metabolism pathway (B), and alanine, aspartate and glutamate metabolism pathway (C)
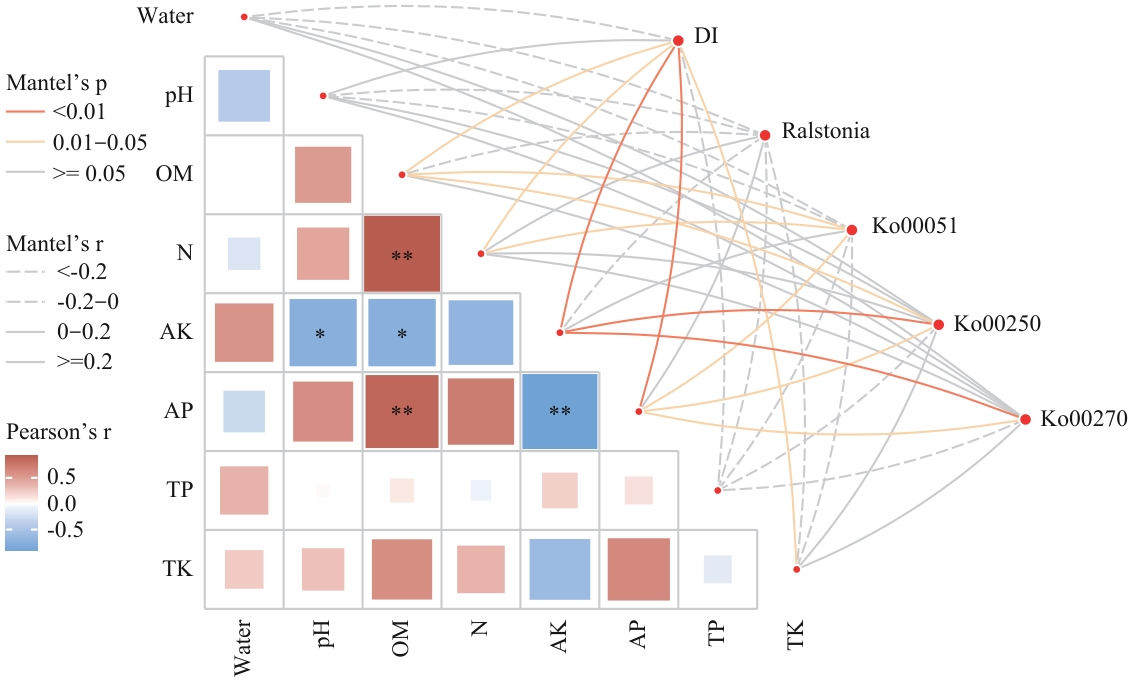
图6 Mantel检验与Pearson相关性热图DI:病情指数;Ralstonia:青枯雷尔氏菌;Ko00051:果糖与甘露糖代谢通路;Ko00250:半胱氨酸和甲硫氨酸代谢通路;Ko00270:丙氨酸、天冬氨酸和谷氨酸代谢通路
Fig. 6 Mantel test and Pearson correlation heatmapDI: Disease index; Ralstonia: Ralstonia solanacearum; Ko00051: fructose and mannose metabolism pathway; Ko00250: cysteine and methionine metabolism pathway; Ko00270: alanine, aspartate, and glutamate metabolism pathway
| [1] | Yang BY, Feng CC, Jiang H, et al. Effects of long-term continuous cropping on microbial community structure and function in tobacco rhizosphere soil [J]. Front Microbiol, 2025, 16: 1496385. |
| [2] | Chen YD, Yang L, Zhang LM, et al. Autotoxins in continuous tobacco cropping soils and their management [J]. Front Plant Sci, 2023, 14: 1106033. |
| [3] | Gu KY, Liu XL, Liu M, et al. Tobacco intercropping enhances soil fertility by improving synergic interactions between soil physicochemical and microbial properties [J]. Front Microbiol, 2025, 16: 1647493. |
| [4] | Xia H, Jiang CQ, Riaz M, et al. Impacts of continuous cropping on soil fertility, microbial communities, and crop growth under different tobacco varieties in a field study [J]. Environ Sci Eur, 2025, 37(1): 5. |
| [5] | Gong B, He Y, Luo ZB, et al. Response of rhizosphere soil physicochemical properties and microbial community structure to continuous cultivation of tobacco [J]. Ann Microbiol, 2024, 74(1): 4. |
| [6] | Li GT, Gong PF, Zhou J, et al. The succession of rhizosphere microbial community in the continuous cropping soil of tobacco [J]. Front Environ Sci, 2024, 11: 1251938. |
| [7] | Ding MJ, Dai HX, He Y, et al. Continuous cropping system altered soil microbial communities and nutrient cycles [J]. Front Microbiol, 2024, 15: 1374550. |
| [8] | Zhang ZH, Huang WX, Jia W, et al. Combined use of green fertilizer and soil conditioner improves soil physicochemical properties, microbial community structure and tobacco quality in continuous cropping [J]. Ann Microbiol, 2025, 75(1): 14. |
| [9] | Feng JN, Chen LL, Xia TY, et al. Microbial fertilizer regulates C: N: P stoichiometry and alleviates phosphorus limitation in flue-cured tobacco planting soil [J]. Sci Rep, 2023, 13: 10276. |
| [10] | Jiang GF, Wei Z, Xu J, et al. Bacterial wilt in China: history, current status, and future perspectives [J]. Front Plant Sci, 2017, 8: 1549. |
| [11] | Li YY, Feng J, Liu HL, et al. Genetic diversity and pathogenicity of Ralstonia solanacearum causing tobacco bacterial wilt in China [J]. Plant Dis, 2016, 100(7): 1288-1296. |
| [12] | Wang ZJ, Luo WB, Cheng SJ, et al. Ralstonia solanacearum-A soil borne hidden enemy of plants: Research development in management strategies, their action mechanism and challenges [J]. Front Plant Sci, 2023, 14: 1141902. |
| [13] | Wang ZB, Zhang YZ, Bo GD, et al. Ralstonia solanacearum infection disturbed the microbiome structure throughout the whole tobacco crop niche as well as the nitrogen metabolism in soil [J]. Front Bioeng Biotechnol, 2022, 10: 903555. |
| [14] | Yao T, Wang CY, Ren Q, et al. Bacterial wilt alters the microbial community characteristics of tobacco root and rhizosphere soil [J]. Rhizosphere, 2024, 32: 100995. |
| [15] | 苏文英, 纪伟, 刘晓梅, 等. 连云港地区秸秆-大球盖菇-大豆-豆丹种养循环模式 [J]. 农业与技术, 2022, 42(22): 53-56. |
| Su WY, Ji W, Liu XM, et al. Cyclic model of straw-Stropharia rugoso-soybean-Doudan in Lianyungang area [J]. Agric & Technol, 2022, 42(22): 53-56. | |
| [16] | 杨晓波, 郑光耀, 徐丽红, 等. 不同栽培料栽培大球盖菇比较试验 [J]. 食用菌, 2021, 43(2): 32-33, 9. |
| Yang XB, Zheng GY, Xu LH, et al. Comparative test on cultivating Stropharia rugosoannulata with different cultivation materials [J]. Edible Fungi, 2021, 43(2): 32-33, 9. | |
| [17] | He Y, Gong B, Liang TB, et al. Effects of reductive soil disinfestation on microbiological and physicochemical properties of continuous cropping soils in karst areas of Guizhou Province [J]. Ann Microbiol, 2025, 75(1): 6. |
| [18] | Shi HQ, Xu PW, Wu SX, et al. Analysis of rhizosphere bacterial communities of tobacco resistant and non-resistant to bacterial wilt in different regions [J]. Sci Rep, 2022, 12: 18309. |
| [19] | Wang XJ, Li ZH, Jin HG, et al. Effects of mushroom-tobacco rotation on microbial community structure in continuous cropping tobacco soil [J]. J Appl Microbiol, 2023: lxad088. |
| [20] | 国家质量监督检验检疫总局 中国国家标准化管理委员会. 烟草病虫害分级及调查方法: [S]. 北京: 中国标准出版社, 2009. |
| Standardization Administration of the People's Republic of China. Grade and investigation method of tobacco diseases and insect pests: [S]. Beijing: Standards Press of China, 2009. | |
| [21] | Chen SF, Zhou YQ, Chen YR, et al. Fastp: an ultra-fast all-in-one FASTQ preprocessor [J]. Bioinformatics, 2018, 34(17): i884-i890. |
| [22] | Li DH, Liu CM, Luo RB, et al. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph [J]. Bioinformatics, 2015, 31(10): 1674-1676. |
| [23] | Seemann T. Prokka: rapid prokaryotic genome annotation [J]. Bioinformatics, 2014, 30(14): 2068-2069. |
| [24] | Steinegger M, Söding J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets [J]. Nat Biotechnol, 2017, 35(11): 1026-1028. |
| [25] | Aroney STN, Newell RJP, Nissen JN, et al. CoverM: read alignment statistics for metagenomics [J]. Bioinformatics, 2025, 41(4): btaf147. |
| [26] | Cao YF, Shen ZZ, Zhang N, et al. Phosphorus availability influences disease-suppressive soil microbiome through plant-microbe interactions [J]. Microbiome, 2024, 12(1): 185. |
| [27] | Wu K, Yuan SF, Wang LL, et al. Effects of bio-organic fertilizer plus soil amendment on the control of tobacco bacterial wilt and composition of soil bacterial communities [J]. Biol Fertil Soils, 2014, 50(6): 961-971. |
| [28] | Wang S, Wang L, Li SC, et al. The win-win effects of an invasive plant biochar on a soil-crop system: controlling a bacterial soilborne disease and stabilizing the soil microbial community network [J]. Microorganisms, 2024, 12(3): 447. |
| [29] | Yang BY, Zhang CQ, Guan CW, et al. Analysis of the composition and function of rhizosphere microbial communities in plants with tobacco bacterial wilt disease and healthy plants [J]. Microbiol Spectr, 2024, 12(12): e0055924. |
| [30] | Suresh P, Shanmugaiah V, Rajagopal R, et al. Pseudomonas fluorescens VSMKU3054 mediated induced systemic resistance in tomato against Ralstonia solanacearum [J]. Physiol Mol Plant Pathol, 2022, 119: 101836. |
| [31] | Jiao WL, Wen J, Li N, et al. The biocontrol potentials of rhizospheric bacterium Bacillus velezensis K0T24 against mulberry bacterial wilt disease [J]. Arch Microbiol, 2024, 206(5): 213. |
| [32] | Wang NQ, Ping L, Mei XL, et al. Succinic acid reduces tomato bacterial wilt disease by recruiting Sphingomonas sp. [J]. Environ Microbiome, 2025, 20(1): 85. |
| [33] | Posas MB, Toyota K, Islam TM. Inhibition of bacterial wilt of tomato caused by Ralstonia solanacearum by sugars and amino acids [J]. Microb Environ, 2007, 22(3): 290-296. |
| [34] | Chen Z, Lin EQ, Lin X, et al. Metabolomic profiling of tomato root exudates induced by Ralstonia solanacearum strains of different pathogenicity: screening for metabolites conferring bacterial wilt resistance [J]. J Microbiol Biotechnol, 2025, 35: e2501033. |
| [35] | Wen T, Xie PH, Liu HW, et al. Tapping the rhizosphere metabolites for the prebiotic control of soil-borne bacterial wilt disease [J]. Nat Commun, 2023, 14: 4497. |
| [36] | Asimakis E, Shehata AA, Eisenreich W, et al. Algae and their metabolites as potential bio-pesticides [J]. Microorganisms, 2022, 10(2): 307. |
| [37] | Yang L, Wei ZL, Valls M, et al. Metabolic profiling of resistant and susceptible tobaccos response incited by Ralstonia pseudosolanacearum causing bacterial wilt [J]. Front Plant Sci, 2022, 12: 780429. |
| [38] | Murti RH, Afifah EN, Nuringtyas TR. Metabolomic response of tomatoes (Solanum lycopersicum L.) against bacterial wilt (Ralstonia solanacearum) using (1)H-NMR spectroscopy [J]. Plants, 2021, 10(6): 1143. |
| [39] | Yang XH, Huang J, Yang HX, et al. Transcriptome and metabolome profiling in different stages of infestation of Eucalyptus urophylla clones by Ralstonia solanacearum [J]. Mol Genet Genom, 2022, 297(4): 1081-1100. |
| [40] | Niu LL, Hu W, Wang FZ, et al. Integrated transcriptomic and metabolomic analyses elucidate the mechanism by which grafting impacts potassium utilization efficiency in tobacco [J]. BMC Plant Biol, 2025, 25(1): 94. |
| [1] | 彭善麟, 廖卓诚, 王涛, 刘志宇, 刘海忆, 王婷婷, 杨琴, 王哲, 邰欢欢. 玉米茎腐病不同抗性品种根际微生物群落多样性及功能差异[J]. 生物技术通报, 2026, 42(5): 1-13. |
| [2] | 张津浩, 邓辉, 张清壮, 陶禹, 周池, 李鑫. 贝莱斯芽胞杆菌XY40-1对百合球茎生长、品质及镉含量的调控作用[J]. 生物技术通报, 2025, 41(7): 281-291. |
| [3] | 李晨莹, 孔大帅, 李若楠, 张玉波, 阎萍, 李奎, 孔思远. 前沿组学技术创新助力畜禽生物育种[J]. 生物技术通报, 2025, 41(6): 71-86. |
| [4] | 陈才锭, 宋云洁, 田梦青. 四大主粮作物不同生育期根际微生物菌群变化研究进展[J]. 生物技术通报, 2025, 41(6): 49-60. |
| [5] | 叶柳健, 蒙健宗, 覃福方, 何双, 朱绮霞, 王小虎, 韦圣博, 周礼芹. 古茶树林菌株D2的鉴定、酶学特性及基因组学分析[J]. 生物技术通报, 2025, 41(5): 267-279. |
| [6] | 唐游, 赵俊伟, 孙兰茜, 李翔. 多组学技术在植物代谢通路解析中的联合应用[J]. 生物技术通报, 2025, 41(4): 76-87. |
| [7] | 李旭娟, 李纯佳, 刘洪博, 徐超华, 林秀琴, 陆鑫, 刘新龙. 甘蔗腋芽形成发育过程的转录组分析[J]. 生物技术通报, 2025, 41(3): 202-218. |
| [8] | 司旭鹏, 崔秀文, 欧阳荔枝, 房丹丹, 裴龙英, 方贵平, 濮希蕾, 马艺沔, 张争. 新疆阿魏不同部位转录组特征分析及倍半萜合成相关基因挖掘[J]. 生物技术通报, 2025, 41(12): 280-293. |
| [9] | 冯小虎, 张文梅, 徐婧, 熊书斌, 王利兵, 宋文静. 一组靶向烟草青枯病的噬菌体鸡尾酒筛选与防控效果[J]. 生物技术通报, 2025, 41(11): 311-318. |
| [10] | 慕雪男, 吴桐, 郑子薇, 张越, 王志刚, 徐伟慧. 一株番茄青枯病生防细菌的筛选、鉴定及其生防潜力分析[J]. 生物技术通报, 2025, 41(1): 276-286. |
| [11] | 於莉军, 王桥美, 彭文书, 严亮, 杨瑞娟. 景迈山古茶园与现代有机茶园根际土壤微生物群落研究[J]. 生物技术通报, 2024, 40(5): 237-247. |
| [12] | 王颢杰, 常栋, 李俊营, 孟颢光, 蒋士君, 周硕野, 崔江宽. 不同生境下烤烟三段式育苗微生物群落变化及抗逆酶活分析[J]. 生物技术通报, 2024, 40(4): 242-254. |
| [13] | 刘佳宁, 李梦, 杨新森, 吴伟, 裴新梧, 袁潜华. 不同水分管理栽培方式对山栏稻根际土壤细菌群落的影响[J]. 生物技术通报, 2024, 40(3): 242-250. |
| [14] | 高云云, 杨海飞, 吕虎杰, 刘永鑫. 微生物组分析方法与功能挖掘[J]. 生物技术通报, 2024, 40(10): 98-107. |
| [15] | 王雨晴, 马子奇, 侯嘉欣, 宗钰琪, 郝晗睿, 刘国元, 魏辉, 连博琳, 陈艳红, 张健. 盐胁迫下植物根系分泌物的成分分析与生态功能研究进展[J]. 生物技术通报, 2024, 40(1): 12-23. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||