








生物技术通报 ›› 2024, Vol. 40 ›› Issue (3): 89-99.doi: 10.13560/j.cnki.biotech.bull.1985.2023-0684
收稿日期:2023-07-17
出版日期:2024-03-26
发布日期:2024-04-08
通讯作者:
马欣荣,博士,教授,研究方向:藻类分子生物学;E-mail: xinrong.ma@tust.edu.cn作者简介:沈天虹,硕士研究生,研究方向:藻类分子生物学;E-mail: shentianhong_2022@163.com
基金资助:
SHEN Tian-hong( ), QI Xiao-bo, ZHAO Rui-feng, MA Xin-rong(
), QI Xiao-bo, ZHAO Rui-feng, MA Xin-rong( )
)
Received:2023-07-17
Published:2024-03-26
Online:2024-04-08
摘要:
微藻是地球上光合微生物的原始种类之一,由于其生长周期较短、生长速率较快和生产高附加值产物的潜力而被广泛开发利用。然而,在微藻放大生产的过程中极易受到高盐等非生物胁迫的不利影响,极大地限制了微藻的生产力。因此,了解微藻盐胁迫响应的分子机制将有助于耐盐藻株的快速建立。本文总结了真核微藻和原核蓝藻响应盐胁迫的各种参与蛋白及其具体作用机制,包括转运蛋白维持离子稳态、积累渗透调节物质、抗氧化防御机制、信号蛋白和脂质积累等;同时综述了已被开发利用的天然耐盐藻包括杜氏盐藻(Dunaliella salina)、盐生隐杆藻(Aphanothece halophytica)、皮克绿球藻(Picochlorum sp.)和海洋绿藻(Chlamydomonas W80)等微藻及其耐盐基因的研究进展;最后讨论了典型盐响应基因在优良藻种选育中的价值与应用前景。
沈天虹, 齐孝博, 赵瑞丰, 马欣荣. 微藻盐胁迫响应分子机制研究进展[J]. 生物技术通报, 2024, 40(3): 89-99.
SHEN Tian-hong, QI Xiao-bo, ZHAO Rui-feng, MA Xin-rong. Research Progress in the Molecular Mechanisms of Microalgae Responding to Salt Stress[J]. Biotechnology Bulletin, 2024, 40(3): 89-99.
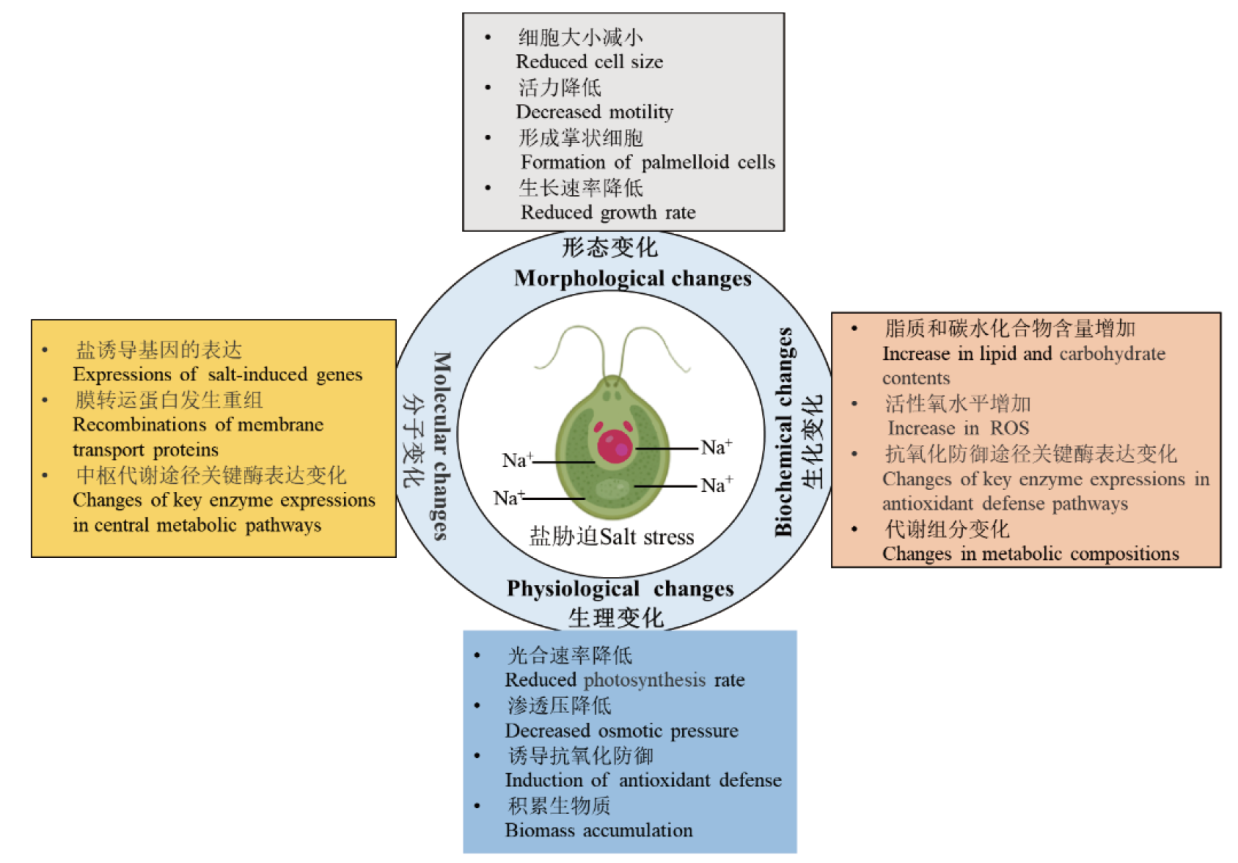
图2 莱茵衣藻(C. reinhardtii)在受到盐分胁迫时形态、生理、生化和分子水平上的变化
Fig. 2 Changes in morphology, physiology, biochemistry and molecular level of C. reinhardtii under salt stress
| [1] |
Siddiki SYA, Mofijur M, Kumar PS, et al. Microalgae biomass as a sustainable source for biofuel, biochemical and biobased value-added products: an integrated biorefinery concept[J]. Fuel, 2022, 307: 121782.
doi: 10.1016/j.fuel.2021.121782 URL |
| [2] |
Arora N, Jaiswal KK, Kumar V, et al. Small-scale phyco-mitigation of raw urban wastewater integrated with biodiesel production and its utilization for aquaculture[J]. Bioresour Technol, 2020, 297: 122489.
doi: 10.1016/j.biortech.2019.122489 URL |
| [3] |
Brar A, Kumar M, Soni T, et al. Insights into the genetic and metabolic engineering approaches to enhance the competence of microalgae as biofuel resource: a review[J]. Bioresour Technol, 2021, 339: 125597.
doi: 10.1016/j.biortech.2021.125597 URL |
| [4] |
Correa DF, Beyer HL, Possingham HP, et al. Microalgal biofuel production at national scales: reducing conflicts with agricultural lands and biodiversity within countries[J]. Energy, 2021, 215: 119033.
doi: 10.1016/j.energy.2020.119033 URL |
| [5] |
Ndimba BK, Ndimba RJ, Johnson TS, et al. Biofuels as a sustainable energy source: an update of the applications of proteomics in bioenergy crops and algae[J]. J Proteomics, 2013, 93: 234-244.
doi: 10.1016/j.jprot.2013.05.041 pmid: 23792822 |
| [6] |
Mallick N, Bagchi SK, Koley S, et al. Progress and challenges in microalgal biodiesel production[J]. Front Microbiol, 2016, 7: 1019.
doi: 10.3389/fmicb.2016.01019 pmid: 27446055 |
| [7] | 聂煜东, 耿媛媛, 张贤明, 等. 产油微藻胁迫培养策略研究综述[J]. 中国环境科学, 2021, 41(8): 3853-3866. |
| Nie YD, Geng YY, Zhang XM, et al. A review on stress cultivation strategies of oleginous microalgae[J]. China Environ Sci, 2021, 41(8): 3853-3866. | |
| [8] | 牛旭东, 李梅, 王宁, 等. 利用微藻处理废水研究进展[J]. 山东农业科学, 2022, 54(2): 146-152. |
| Niu XD, Li M, Wang N, et al. Research progress of treating wastewater with microalgae[J]. Shandong Agric Sci, 2022, 54(2): 146-152. | |
| [9] |
Wang Y, Guo WQ, Yen HW, et al. Cultivation of Chlorella vulgaris JSC-6 with swine wastewater for simultaneous nutrient/COD removal and carbohydrate production[J]. Bioresour Technol, 2015, 198: 619-625.
doi: 10.1016/j.biortech.2015.09.067 URL |
| [10] |
García D, Posadas E, Blanco S, et al. Evaluation of the dynamics of microalgae population structure and process performance during piggery wastewater treatment in algal-bacterial photobioreactors[J]. Bioresour Technol, 2018, 248(Pt B): 120-126.
doi: 10.1016/j.biortech.2017.06.079 URL |
| [11] |
Dahmani S, Zerrouki D, Ramanna L, et al. Cultivation of Chlorella pyrenoidosa in outdoor open raceway pond using domestic wastewater as medium in arid desert region[J]. Bioresour Technol, 2016, 219: 749-752.
doi: 10.1016/j.biortech.2016.08.019 URL |
| [12] |
Fal S, Aasfar A, Rabie R, et al. Salt induced oxidative stress alters physiological, biochemical and metabolomic responses of green microalga Chlamydomonas reinhardtii[J]. Heliyon, 2022, 8(1): e08811.
doi: 10.1016/j.heliyon.2022.e08811 URL |
| [13] |
Salama ES, Govindwar SP, Khandare RV, et al. Can omics approaches improve microalgal biofuels under abiotic stress?[J]. Trends Plant Sci, 2019, 24(7): 611-624.
doi: S1360-1385(19)30097-4 pmid: 31085124 |
| [14] |
Cui JY, Sun T, Chen L, et al. Engineering salt tolerance of photosynthetic cyanobacteria for seawater utilization[J]. Biotechnol Adv, 2020, 43: 107578.
doi: 10.1016/j.biotechadv.2020.107578 URL |
| [15] |
Zhang LY, Xing ZT, Chen LQ, et al. Comprehensive time-course transcriptome and co-expression network analyses identify salt stress responding mechanisms in Chlamydomonas reinhardtii strain GY-D55[J]. Front Plant Sci, 2022, 13: 828321.
doi: 10.3389/fpls.2022.828321 URL |
| [16] |
Ves-Urai P, Krobthong S, Thongsuk K, et al. Comparative secretome analysis between salinity-tolerant and control Chlamydomonas re-inhardtii strains[J]. Planta, 2021, 253(3): 68.
doi: 10.1007/s00425-021-03583-7 |
| [17] |
Hounslow E, Evans CA, Pandhal J, et al. Quantitative proteomic comparison of salt stress in Chlamydomonas reinhardtii and the snow alga Chlamydomonas nivalis reveals mechanisms for salt-triggered fatty acid accumulation via reallocation of carbon resources[J]. Biotechnol Biofuels, 2021, 14(1): 121.
doi: 10.1186/s13068-021-01970-6 pmid: 34022944 |
| [18] |
Raven JA, Girard-Bascou J. Algal model systems and the elucidation of photosynthetic metabolism[J]. J Phycol, 2001, 37(6): 943-950.
doi: 10.1046/j.1529-8817.2001.01079.x URL |
| [19] | 陈龙, 金阿南, 马香娟, 等. 微生物高盐渗透适应策略及其耐盐强化研究进展[J]. 微生物学报, 2022, 62(9): 3306-3317. |
| Chen L, Jin AN, Ma XJ, et al. Research progress on osmotic pressure adaptation strategy and salt tolerance enhancement of microorganisms under high salinity environment[J]. Acta Microbiol Sin, 2022, 62(9): 3306-3317. | |
| [20] |
Ullah MA, Abdullah-Zawawi MR, Zainal-Abidin RA, et al. A review of integrative omic approaches for understanding rice salt response mechanisms[J]. Plants, 2022, 11(11): 1430.
doi: 10.3390/plants11111430 URL |
| [21] |
Shelake RM, Kadam US, Kumar R, et al. Engineering drought and salinity tolerance traits in crops through CRISPR-mediated genome editing: targets, tools, challenges, and perspectives[J]. Plant Commun, 2022, 3(6): 100417.
doi: 10.1016/j.xplc.2022.100417 URL |
| [22] |
Chakraborty K, Bose J, Shabala L, et al. Difference in root K+ retention ability and reduced sensitivity of K+-permeable channels to reactive oxygen species confer differential salt tolerance in three Brassica species[J]. J Exp Bot, 2016, 67(15): 4611-4625.
doi: 10.1093/jxb/erw236 pmid: 27340231 |
| [23] |
Foflonker F, Price DC, Qiu H, et al. Genome of the halotolerant green alga Picochlorum sp. reveals strategies for thriving under fluctuating environmental conditions[J]. Environ Microbiol, 2015, 17(2): 412-426.
doi: 10.1111/1462-2920.12541 pmid: 24965277 |
| [24] |
Taylor AR, Brownlee C, Wheeler GL. Proton channels in algae: reasons to be excited[J]. Trends Plant Sci, 2012, 17(11): 675-684.
doi: 10.1016/j.tplants.2012.06.009 pmid: 22819465 |
| [25] |
Katz A, Pick U. Plasma membrane electron transport coupled to Na+ extrusion in the halotolerant alga Dunaliella[J]. Biochim Biophys Acta, 2001, 1504(2-3): 423-431.
pmid: 11245805 |
| [26] |
Shono M, Wada M, Hara Y, et al. Molecular cloning of Na(+)-ATPase cDNA from a marine alga, Heterosigma akashiwo[J]. Biochim Biophys Acta, 2001, 1511(1): 193-199.
pmid: 11248217 |
| [27] |
Kishimoto M, Shimajiri Y, Oshima A, et al. Functional expression of an animal type-Na+-ATPase gene from a marine red seaweed Porphyra yezoensis increases salinity tolerance in rice plants[J]. Plant Biotechnol, 2013, 30(4): 417-422.
doi: 10.5511/plantbiotechnology.13.0517a URL |
| [28] |
Pick U, Karni L, Avron M. Determination of ion content and ion fluxes in the halotolerant alga Dunaliella salina[J]. Plant Physiol, 1986, 81(1): 92-96.
doi: 10.1104/pp.81.1.92 pmid: 16664814 |
| [29] |
Shetty P, Gitau MM, Maróti G. Salinity stress responses and adaptation mechanisms in eukaryotic green microalgae[J]. Cells, 2019, 8(12): 1657.
doi: 10.3390/cells8121657 URL |
| [30] |
Brown AD, Simpson JR. Water relations of sugar-tolerant yeasts: the role of intracellular polyols[J]. J Gen Microbiol, 1972, 72(3): 589-591.
pmid: 4404634 |
| [31] |
Gunde-Cimerman N, Plemenitaš A, Oren A. Strategies of adaptation of microorganisms of the three domains of life to high salt concentrations[J]. FEMS Microbiol Rev, 2018, 42(3): 353-375.
doi: 10.1093/femsre/fuy009 pmid: 29529204 |
| [32] |
Lv HX, Kim M, Park S, et al. Comparative transcriptome analysis of short-term responses to salt and glycerol hyperosmotic stress in the green alga Dunaliella salina[J]. Algal Res, 2021, 53: 102147.
doi: 10.1016/j.algal.2020.102147 URL |
| [33] |
Ben-Amotz A, Avron M. Glycerol and β-carotene metabolism in the halotolerant alga Dunaliella: a model system for biosolar energy conversion[J]. Trends Biochem Sci, 1981, 6: 297-299.
doi: 10.1016/0968-0004(81)90106-7 URL |
| [34] |
Hellebust JA, Le Gresley SML. Growth characteristics of the marine rock pool flagellate Chlamydomonas pulsatilla Wollenweber(Chlorophyta)[J]. Phycologia, 1985, 24(2): 225-229.
doi: 10.2216/i0031-8884-24-2-225.1 URL |
| [35] |
Liu LJ, Huang L, Lin XY, et al. Hydrogen peroxide alleviates salinity-induced damage through enhancing proline accumulation in wheat seedlings[J]. Plant Cell Rep, 2020, 39(5): 567-575.
doi: 10.1007/s00299-020-02513-3 pmid: 32025801 |
| [36] |
Mohseni A, Fan LH, Roddick FA. Impact of microalgae species and solution salinity on algal treatment of wastewater reverse osmosis concentrate[J]. Chemosphere, 2021, 285: 131487.
doi: 10.1016/j.chemosphere.2021.131487 URL |
| [37] |
Ma XC, Wei HY, Zhang YD, et al. Glutathione peroxidase 5 deficiency induces lipid metabolism regulated by reactive oxygen species in Chlamydomonas reinhardtii[J]. Microb Pathog, 2020, 147: 104358.
doi: 10.1016/j.micpath.2020.104358 URL |
| [38] |
Moghimifam R, Niknam V, Ebrahimzadeh H, et al. The influence of different CO2 concentrations on the biochemical and molecular response of two isolates of Dunaliella sp.(ABRIINW-CH2 and ABRIINW-SH33)[J]. J Appl Phycol, 2020, 32(1): 175-187.
doi: 10.1007/s10811-019-01914-6 |
| [39] |
Pathak J, Singh PR, Häder DP, et al. UV-induced DNA damage and repair: a cyanobacterial perspective[J]. Plant Gene, 2019, 19: 100194.
doi: 10.1016/j.plgene.2019.100194 URL |
| [40] |
Mittler R. Oxidative stress, antioxidants and stress tolerance[J]. Trends Plant Sci, 2002, 7(9): 405-410.
doi: 10.1016/s1360-1385(02)02312-9 pmid: 12234732 |
| [41] |
Yeh HL, Lin TH, Chen CC, et al. Monodehydroascorbate reductase plays a role in the tolerance of Chlamydomonas reinhardtii to photooxidative stress[J]. Plant Cell Physiol, 2019, 60(10): 2167-2179.
doi: 10.1093/pcp/pcz110 URL |
| [42] |
Lin ST, Chiou CW, Chu YL, et al. Enhanced ascorbate regeneration via dehydroascorbate reductase confers tolerance to photo-oxidative stress in Chlamydomonas reinhardtii[J]. Plant Cell Physiol, 2016, 57(10): 2104-2121.
doi: 10.1093/pcp/pcw129 URL |
| [43] |
Choi BY, Kim H, Shim D, et al. The Chlamydomonas bZIP transcription factor BLZ8 confers oxidative stress tolerance by inducing the carbon-concentrating mechanism[J]. Plant Cell, 2022, 34(2): 910-926.
doi: 10.1093/plcell/koab293 URL |
| [44] |
Peng Z, Liu G, Huang KY. Cold adaptation mechanisms of a snow alga Chlamydomonas nivalis during temperature fluctuations[J]. Front Microbiol, 2021, 11: 611080.
doi: 10.3389/fmicb.2020.611080 URL |
| [45] |
Podgornaia AI, Laub MT. Determinants of specificity in two-component signal transduction[J]. Curr Opin Microbiol, 2013, 16(2): 156-162.
doi: 10.1016/j.mib.2013.01.004 pmid: 23352354 |
| [46] |
Lenzoni G, Liu JL, Knight MR. Predicting plant immunity gene expression by identifying the decoding mechanism of calcium signatures[J]. New Phytol, 2018, 217(4): 1598-1609.
doi: 10.1111/nph.14924 pmid: 29218709 |
| [47] |
Li YJ, Fei XW, Dai HF, et al. Genome-wide identification of calcium-dependent protein kinases in Chlamydomonas reinhardtii and functional analyses in nitrogen deficiency-induced oil accumulation[J]. Front Plant Sci, 2019, 10: 1147.
doi: 10.3389/fpls.2019.01147 URL |
| [48] |
Gomez-Osuna A, Calatrava V, Galvan A, et al. Identification of the MAPK cascade and its relationship with nitrogen metabolism in the green alga Chlamydomonas reinhardtii[J]. Int J Mol Sci, 2020, 21(10): 3417.
doi: 10.3390/ijms21103417 URL |
| [49] |
Tang ZY, Cao XY, Zhang YP, et al. Two splice variants of the DsMEK1 mitogen-activated protein kinase kinase(MAPKK)are involved in salt stress regulation in Dunaliella salina in different ways[J]. Biotechnol Biofuels, 2020, 13: 147.
doi: 10.1186/s13068-020-01786-w |
| [50] |
Waditee R, Hibino T, Nakamura T, et al. Overexpression of a Na+/H+ antiporter confers salt tolerance on a freshwater cyanobacterium, making it capable of growth in sea water[J]. Proc Natl Acad Sci USA, 2002, 99(6): 4109-4114.
doi: 10.1073/pnas.052576899 URL |
| [51] |
Soontharapirakkul K, Promden W, Yamada N, et al. Halotolerant cyanobacterium Aphanothece halophytica contains an Na+-dependent F1F0-ATP synthase with a potential role in salt-stress tolerance[J]. J Biol Chem, 2011, 286(12): 10169-10176.
doi: 10.1074/jbc.M110.208892 pmid: 21262962 |
| [52] |
Fukaya F, Promden W, Hibino T, et al. An mrp-like cluster in the halotolerant Cyanobacterium Aphanothece halophytica functions as a Na+/H+ antiporter[J]. Appl Environ Microbiol, 2009, 75(20): 6626-6629.
doi: 10.1128/AEM.01387-09 URL |
| [53] |
Checchetto V, Segalla A, Sato Y, et al. Involvement of potassium transport systems in the response of Synechocystis PCC 6803 cyanobacteria to external pH change, high-intensity light stress and heavy metal stress[J]. Plant Cell Physiol, 2016, 57(4): 862-877.
doi: 10.1093/pcp/pcw032 pmid: 26880819 |
| [54] |
Pade N, Michalik D, Ruth W, et al. Trimethylated homoserine functions as the major compatible solute in the globally significant oceanic cyanobacterium Trichodesmium[J]. Proc Natl Acad Sci USA, 2016, 113(46): 13191-13196.
doi: 10.1073/pnas.1611666113 URL |
| [55] |
Song K, Tan XM, Liang YJ, et al. The potential of Synechococcus elongatus UTEX 2973 for sugar feedstock production[J]. Appl Microbiol Biotechnol, 2016, 100(18): 7865-7875.
doi: 10.1007/s00253-016-7510-z pmid: 27079574 |
| [56] |
Kempf B, Bremer E. Uptake and synthesis of compatible solutes as microbial stress responses to high-osmolality environments[J]. Arch Microbiol, 1998, 170(5): 319-330.
pmid: 9818351 |
| [57] |
Chen THH, Murata N. Enhancement of tolerance of abiotic stress by metabolic engineering of betaines and other compatible solutes[J]. Curr Opin Plant Biol, 2002, 5(3): 250-257.
doi: 10.1016/s1369-5266(02)00255-8 pmid: 11960744 |
| [58] |
Rontein D, Basset G, Hanson AD. Metabolic engineering of osmoprotectant accumulation in plants[J]. Metab Eng, 2002, 4(1): 49-56.
pmid: 11800574 |
| [59] |
Waditee R, Tanaka Y, Aoki K, et al. Isolation and functional characterization of N-methyltransferases that catalyze betaine synthesis from glycine in a halotolerant photosynthetic organism Aphanothece halophytica[J]. J Biol Chem, 2003, 278(7): 4932-4942.
doi: 10.1074/jbc.M210970200 URL |
| [60] |
Waditee-Sirisattha R, Singh M, Kageyama H, et al. Anabaena sp. PCC7120 transformed with glycine methylation genes from Aphan-othece halophytica synthesized glycine betaine showing increased tolerance to salt[J]. Arch Microbiol, 2012, 194(11): 909-914.
doi: 10.1007/s00203-012-0824-z pmid: 22707090 |
| [61] |
Verma E, Singh S, Niveshika, et al. Salinity-induced oxidative stress-mediated change in fatty acids composition of cyanobacterium Synechococcus sp. PCC7942[J]. Int J Environ Sci Technol, 2019, 16(2): 875-886.
doi: 10.1007/s13762-018-1720-0 |
| [62] |
Huflejt ME, Tremolieres A, Pineau B, et al. Changes in membrane lipid composition during saline growth of the fresh water cyanobacterium Synechococcus 6311[J]. Plant Physiol, 1990, 94(4): 1512-1521.
doi: 10.1104/pp.94.4.1512 URL |
| [63] |
Ritter D, Yopp JH. Plasma membrane lipid composition of the halophilic cyanobacterium Aphanothece halophytica[J]. Arch Microbiol, 1993, 159(5): 435-439.
doi: 10.1007/BF00288590 URL |
| [64] |
Joset F, Jeanjean R, Hagemann M. Dynamics of the response of cyanobacteria to salt stress: Deciphering the molecular events[J]. Physiol Plant, 1996, 96(4): 738-744.
doi: 10.1111/ppl.1996.96.issue-4 URL |
| [65] |
Yamamori T, Kageyama H, Tanaka Y, et al. Requirement of alkanes for salt tolerance of Cyanobacteria: characterization of alkane synthesis genes from salt-sensitive Synechococcus elongatus PCC7942 and salt-tolerant Aphanothece halophytica[J]. Lett Appl Microbiol, 2018, 67(3): 299-305.
doi: 10.1111/lam.13038 pmid: 30039571 |
| [66] |
Agostoni M, Montgomery BL. Survival strategies in the aquatic and terrestrial world: the impact of second messengers on cyanobacterial processes[J]. Life, 2014, 4(4): 745-769.
doi: 10.3390/life4040745 URL |
| [67] |
Imashimizu M, Yoshimura H, Katoh H, et al. NaCl enhances cellular cAMP and upregulates genes related to heterocyst development in the cyanobacterium, Anabaena sp. strain PCC 7120[J]. FEMS Microbiol Lett, 2005, 252(1): 97-103.
pmid: 16182471 |
| [68] |
Cadoret JC, Rousseau B, Perewoska I, et al. Cyclic nucleotides, the photosynthetic apparatus and response to a UV-B stress in the Cyanobacterium synechocystis sp. PCC 6803[J]. J Biol Chem, 2005, 280(40): 33935-33944.
doi: 10.1074/jbc.M503153200 URL |
| [69] |
Agostoni M, Logan-Jackson AR, Heinz ER, et al. Homeostasis of second messenger cyclic-di-AMP is critical for cyanobacterial fitness and acclimation to abiotic stress[J]. Front Microbiol, 2018, 9: 1121.
doi: 10.3389/fmicb.2018.01121 pmid: 29896182 |
| [70] |
Angerer V, Schwenk P, Wallner T, et al. The protein Slr1143 is an active diguanylate cyclase in Synechocystis sp. PCC 6803 and interacts with the photoreceptor Cph2[J]. Microbiology, 2017, 163(6): 920-930.
doi: 10.1099/mic.0.000475 URL |
| [71] |
Xu CX, Sun T, Li SB, et al. Adaptive laboratory evolution of cadmium tolerance in Synechocystis sp. PCC 6803[J]. Biotechnol Biofuels, 2018, 11: 205.
doi: 10.1186/s13068-018-1205-x |
| [72] |
Perrineau MM, Zelzion E, Gross J, et al. Evolution of salt tolerance in a laboratory reared population of Chlamydomonas reinhard-tii[J]. Environ Microbiol, 2014, 16(6): 1755-1766.
doi: 10.1111/emi.2014.16.issue-6 URL |
| [73] |
Kato Y, Ho SH, Vavricka CJ, et al. Evolutionary engineering of salt-resistant Chlamydomonas sp. strains reveals salinity stress-activated starch-to-lipid biosynthesis switching[J]. Bioresour Technol, 2017, 245(Pt B): 1484-1490.
doi: 10.1016/j.biortech.2017.06.035 URL |
| [74] |
Chen H, Jiang JG. Osmotic responses of Dunaliella to the changes of salinity[J]. J Cell Physiol, 2009, 219(2): 251-258.
doi: 10.1002/jcp.21715 pmid: 19202552 |
| [75] |
Goyal A. Osmoregulation in Dunaliella, Part II: Photosynthesis and starch contribute carbon for glycerol synthesis during a salt stress in Dunaliella tertiolecta[J]. Plant Physiol Biochem, 2007, 45(9): 705-710.
doi: 10.1016/j.plaphy.2007.05.009 URL |
| [76] | Gong WF, Zhao LN, Hu B, et al. Identifying novel salt-tolerant genes from Dunaliella salina using a Haematococcus pluvialis expression system[J]. Plant Cell Tissue Organ Cult PCTOC, 2014, 117(1): 113-124. |
| [77] |
Miller DM, Jones JH, Yopp JH, et al. Ion metabolism in a halophilic blue-green alga, Aphanothece halophytica[J]. Arch Microbiol, 1976, 111(1/2): 145-149.
doi: 10.1007/BF00446561 URL |
| [78] |
Munns R, Tester M. Mechanisms of salinity tolerance[J]. Annu Rev Plant Biol, 2008, 59: 651-681.
doi: 10.1146/annurev.arplant.59.032607.092911 pmid: 18444910 |
| [79] |
Ono K, Hibino T, Kohinata T, et al. Overexpression of DnaK from a halotolerant Cyanobacterium Aphanothece halophytica enhances the high-temperatue tolerance of tobacco during germination and early growth[J]. Plant Sci, 2001, 160(3): 455-461.
pmid: 11166432 |
| [80] |
Scaife MA, Nguyen GTDT, Rico J, et al. Establishing Chla-mydomonas reinhardtii as an industrial biotechnology host[J]. Plant J, 2015, 82(3): 532-546.
doi: 10.1111/tpj.2015.82.issue-3 URL |
| [81] |
Ma XR, Kim EJ, Kook I, et al. Small interfering RNA-mediated translation repression alters ribosome sensitivity to inhibition by cycloheximide in Chlamydomonas reinhardtii[J]. Plant Cell, 2013, 25(3): 985-998.
doi: 10.1105/tpc.113.109256 URL |
| [82] |
Akella S, Ma XR, Bacova R, et al. Co-targeting strategy for precise, scarless gene editing with CRISPR/Cas9 and donor ssODNs in Chlamydomonas[J]. Plant Physiol, 2021, 187(4): 2637-2655.
doi: 10.1093/plphys/kiab418 URL |
| [1] | 李昊, 伍国强, 魏明, 韩悦欣. 甜菜BvBADH基因家族全基因组鉴定及其高盐胁迫下的表达分析[J]. 生物技术通报, 2024, 40(2): 233-244. |
| [2] | 徐扬, 张瑞英, 戴良香, 张冠初, 丁红, 张智猛. 盐胁迫下氮素对花生种子萌发和种子际细菌菌群结构的调控[J]. 生物技术通报, 2024, 40(2): 253-265. |
| [3] | 王雨晴, 马子奇, 侯嘉欣, 宗钰琪, 郝晗睿, 刘国元, 魏辉, 连博琳, 陈艳红, 张健. 盐胁迫下植物根系分泌物的成分分析与生态功能研究进展[J]. 生物技术通报, 2024, 40(1): 12-23. |
| [4] | 焦进兰, 王文文, 介欣芮, 王华忠, 岳洁瑜. 外源钙缓解小麦幼苗盐胁迫的作用机制[J]. 生物技术通报, 2024, 40(1): 207-221. |
| [5] | 王帅, 冯宇梅, 白苗, 杜维俊, 岳爱琴. 大豆GmHMGR基因响应外源激素及非生物胁迫功能研究[J]. 生物技术通报, 2023, 39(7): 131-142. |
| [6] | 魏茜雅, 秦中维, 梁腊梅, 林欣琪, 李映志. 褪黑素种子引发处理提高朝天椒耐盐性的作用机制[J]. 生物技术通报, 2023, 39(7): 160-172. |
| [7] | 王海龙, 李雨倩, 王勃, 邢国芳, 张杰伟. 谷子SiMAPK3基因的克隆和表达特性分析[J]. 生物技术通报, 2023, 39(3): 123-132. |
| [8] | 杜清洁, 周璐瑶, 杨思震, 张嘉欣, 陈春林, 李娟起, 李猛, 赵士文, 肖怀娟, 王吉庆. 过表达CaCP1提高转基因烟草对盐胁迫的敏感性[J]. 生物技术通报, 2023, 39(2): 172-182. |
| [9] | 叶红, 王玉昆. 植物PRR免疫受体功能研究进展[J]. 生物技术通报, 2023, 39(12): 1-15. |
| [10] | 汪明滔, 刘建伟, 赵春钊. 植物调控盐胁迫下细胞壁完整性的分子机制[J]. 生物技术通报, 2023, 39(11): 18-27. |
| [11] | 张玉娟, 黎冬华, 宫慧慧, 崔新晓, 高春华, 张秀荣, 游均, 赵军胜. 芝麻NAC转录因子基因SiNAC77的克隆及耐盐功能分析[J]. 生物技术通报, 2023, 39(11): 308-317. |
| [12] | 徐扬, 丁红, 张冠初, 郭庆, 张智猛, 戴良香. 盐胁迫下花生种子萌发期代谢组学分析[J]. 生物技术通报, 2023, 39(1): 199-213. |
| [13] | 陈光, 李佳, 杜瑞英, 王旭. 水稻盐敏感突变体ss2的鉴定与基因功能分析[J]. 生物技术通报, 2022, 38(9): 158-166. |
| [14] | 张斌, 杨昕霞. 水稻响应盐胁迫关键转录因子的鉴定[J]. 生物技术通报, 2022, 38(3): 9-15. |
| [15] | 张业猛, 朱丽丽, 陈志国. 藜麦NHX基因家族鉴定及盐胁迫下表达分析[J]. 生物技术通报, 2022, 38(12): 184-193. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||