








生物技术通报 ›› 2026, Vol. 42 ›› Issue (3): 283-293.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0704
罗威1,2( ), 宫奥1, 仲阳3, 胡迪1, 周洪源1, 张泓欣1, 艾菊1, 罗有卫4, 高冬丽1(
), 宫奥1, 仲阳3, 胡迪1, 周洪源1, 张泓欣1, 艾菊1, 罗有卫4, 高冬丽1( )
)
收稿日期:2025-07-01
出版日期:2026-03-26
发布日期:2026-04-23
通讯作者:
高冬丽,女,博士,研究员,研究方向 :果实发育调控;E-mail: gdongli@126.com作者简介:罗威,男,博士研究生,研究方向 :基因功能解析;E-mail: 1136689701@qq.com基金资助:
LUO Wei1,2( ), GONG Ao1, ZHONG Yang3, HU Di1, ZHOU Hong-yuan1, ZHANG Hong-xin1, AI Ju1, LUO You-wei4, GAO Dong-li1(
), GONG Ao1, ZHONG Yang3, HU Di1, ZHOU Hong-yuan1, ZHANG Hong-xin1, AI Ju1, LUO You-wei4, GAO Dong-li1( )
)
Received:2025-07-01
Published:2026-03-26
Online:2026-04-23
摘要:
目的 SEPALLATA2(SEP2)编码MADS家族转录因子,利用Cssep2突变体揭示其在黄瓜(Cucumis sativus L.)果实和疣状结构发育中的作用,并探究其调控机制,为黄瓜重要性状的深度解析提供理论支持。 方法 通过实时荧光定量分析CsSEP2的表达模式;亚细胞定位分析CsSEP2在细胞内的具体位置;通过稳定遗传转化创制Cssep2敲除突变体,从细胞学和转录组层面探究其表型出现的分子机制;利用酵母双杂交和萤火虫荧光素酶互补实验分析CsSEP2和其他蛋白之间的相互作用。 结果 CsSEP2的mRNA特异性积累于花和果实组织,其编码蛋白定位于细胞核。利用CRISPR/Cas9技术构建功能缺失突变体。突变体果实因细胞分裂与伸长进程受阻而显著缩短,且呈现开裂表型;果实疣状结构发育异常,表现为果刺密度增加、果刺基部缩小及果瘤萎缩。外果皮与胎座的转录组测序分析显示,CsSEP2功能缺失显著改变基因表达谱,尤其是与果实大小和疣状结构发育相关基因表达发生显著变化。CsSEP2与多个已知的调控疣状结构发育的蛋白存在互作关系。 结论 CsSEP2突变对果实和疣状结构发育产生了多效性影响。CsSEP2通过影响一系列基因的表达和蛋白间的互作来调控果实长度和疣状结构发育。
罗威, 宫奥, 仲阳, 胡迪, 周洪源, 张泓欣, 艾菊, 罗有卫, 高冬丽. 黄瓜SEPALLATA2基因敲除对果实及疣状结构发育的多效性影响[J]. 生物技术通报, 2026, 42(3): 283-293.
LUO Wei, GONG Ao, ZHONG Yang, HU Di, ZHOU Hong-yuan, ZHANG Hong-xin, AI Ju, LUO You-wei, GAO Dong-li. Pleiotropic Effects of SEPALLATA2 Knock-out on Fruit and Wart Development in Cucumber[J]. Biotechnology Bulletin, 2026, 42(3): 283-293.
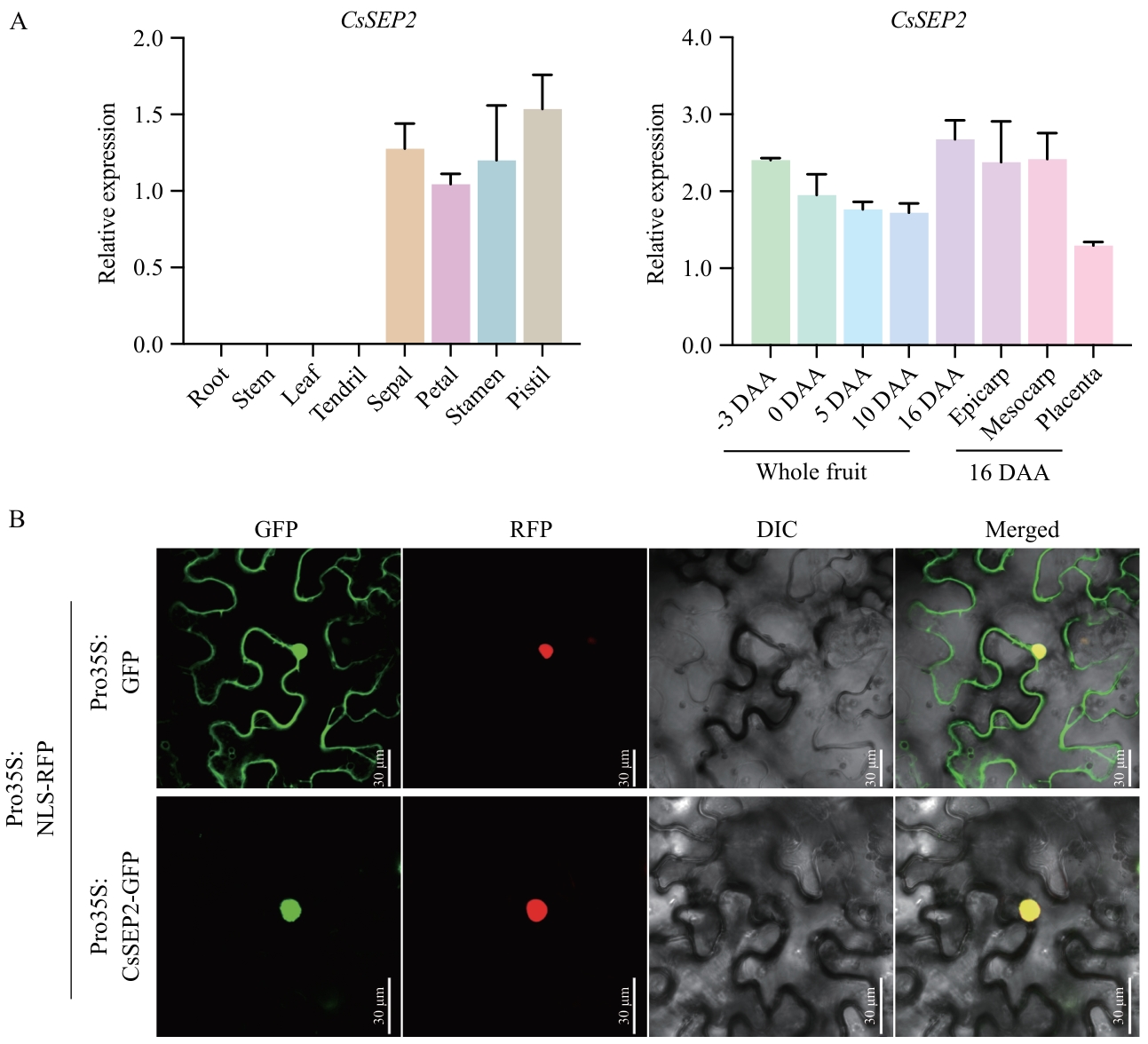
图1 CsSEP2基因表达分析(A)与亚细胞定位(B)图A中,检测CsSEP2在-3、0、5、10、16 DAA的果实,以及16 DAA果实中部外果皮、中果皮和胎座组织中的基因表达。数据以“均值±标准差”表示,3次生物学重复。DAA表示花后天数,下同
Fig. 1 Expression analyses (A) and subcellular localization (B) of gene CsSEP2In Fig. A, the expressions of CsSEP2 in thefruits of -3, 0, 5, 10, and 16 DAA and epicarp, mesocarp and placenta from the central parts of fruits at 16 DAA were examined. Data are means ± SD, n = 3. DAA refers to days after anthesis, the same below
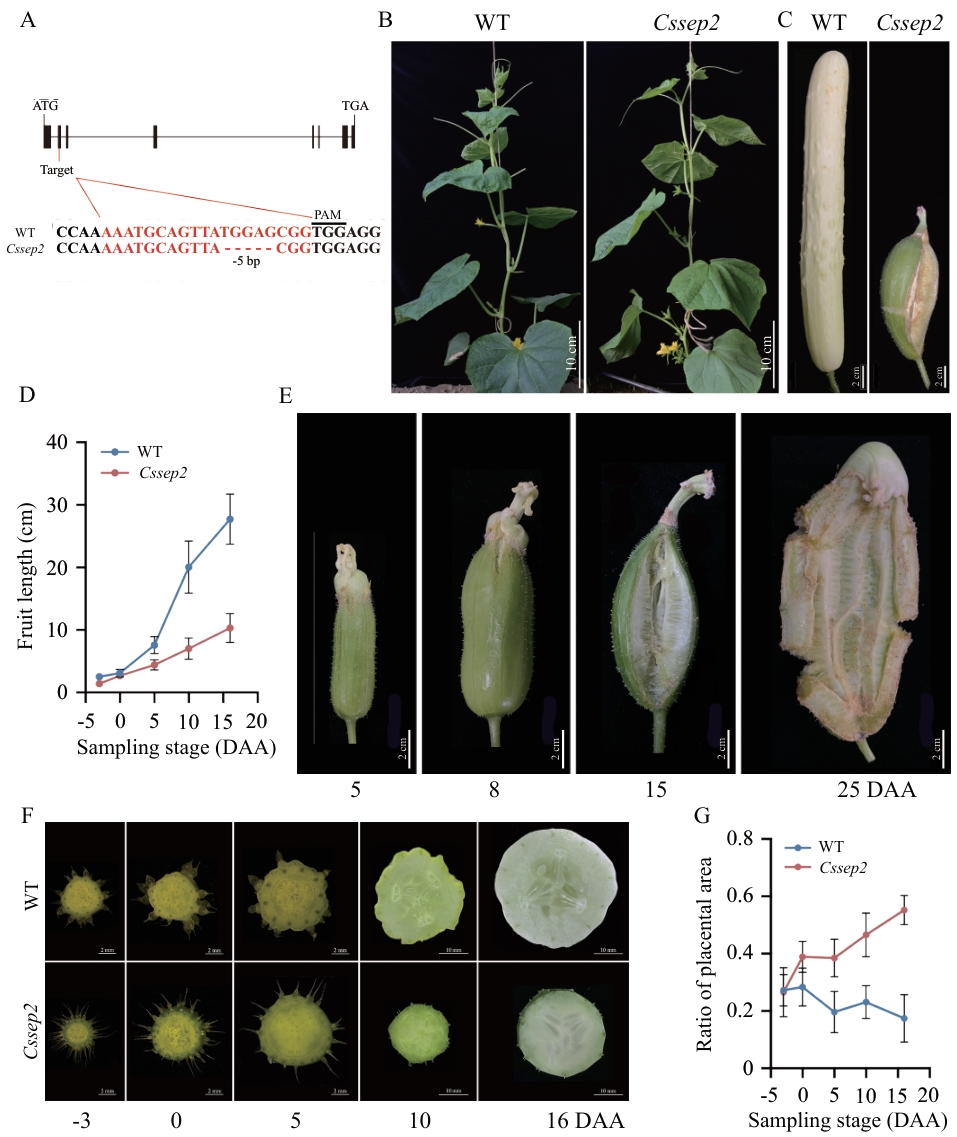
图2 CsSEP2基因敲除导致果实发育异常A:CsSEP2基因结构及Cssep2突变体的突变位点示意图;外显子和内含子分别以黑色方框和直线表示,红色核苷酸序列为基因编辑靶点,下划线标注的3个碱基为前间隔序列邻近基序(PAM);B、C:WT和Cssep2突变体植株(B)及果实(C)的代表性表型;D:WT和Cssep2突变体果实长度测定,5次生物学重复;E:Cssep2突变体果实开裂的表型;F:WT和Cssep2突变体果实胎座的对比;G:WT和Cssep2突变体果实胎座面积与横截面积的比值分析,样本量n ≥ 5,数据以“均值 ± 标准差”表示
Fig. 2 Knock-out of gene CsSEP2 resulted in abnormal fruit developmentA: Structure of the CsSEP2 gene and the mutated loci in the Cssep2 mutants. Exons and introns are denoted by black boxes and lines, respectively. The red nucleotide sequences are the editing target site of the gene, and the three underlined bases indicate the protospacer adjacent motif (PAM). B, C: Representative plants (B) and fruits (C) of WT and Cssep2 mutants. D: Measurement of the fruit lengths of WT and Cssep2 mutants. Data are means ± SD, n = 5. E: Macroscopic observation of fruit dehiscence of the Cssep2 mutants. F: Comparative observation of the fruit placentas of WT and Cssep2 mutants. G: The ratio of placenta areas to fruit cross areas of WT and Cssep2 mutants. Values are means ± SD, n ≥ 5
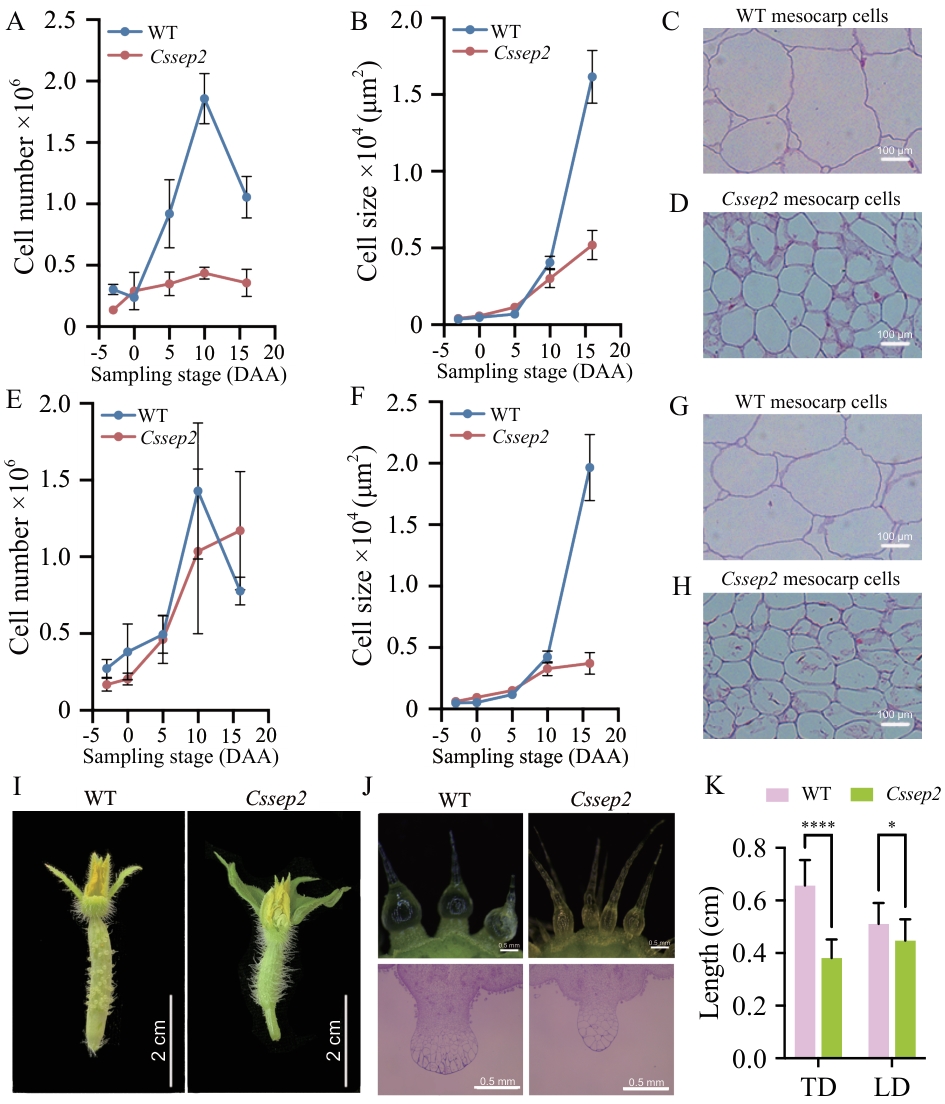
图3 Cssep2突变体果实及疣状结构的细胞学分析A、B:中果皮纵向细胞数目与细胞大小测定,样本量n = 540;C、D:16 DAA中果皮纵向切片观察;E、F:中果皮横向细胞数目与细胞大小测定,样本量n = 540;G、H:16 DAA中果皮横向切片观察:I:-3 DAA WT和Cssep2突变体果刺表型;J:-3 DAA WT和Cssep2突变体疣状结构表型;K:-3 DAA WT和Cssep2突变体果刺基部尺寸测量,TD:横向直径,LD:纵向直径;样本量n = 18;* P < 0.05;**** P < 0.000 1。数据以“均值 ± 标准差”表示。下同
Fig. 3 Cellular dissection of the fruit and wart phenotype of the Cssep2 mutantsA, B: Cell number and size of fruit mesocarp along the longitudinal axis. Values are means ± SD, n = 540. C, D: Longitudinal sections of the fruit mesocarp at 16 DAA. E, F: Cell number and size of fruit mesocarp along the transversal axis. Values are means ± SD, n = 540. G, H: Transversal sections of the fruit mesocarp at 16 DAA. I: Fruit spine of WT and Cssep2 mutants at -3 DAA. J: Macroscopic observation of the tubercule at -3 DAA. K: Measurement of the spine base of WT and Cssep2 mutants at -3 DAA. TD: Traverse diameter, LD: longitudinal diameter; * P < 0.05; **** P < 0.000 1. Values are means ± SD, n = 18. The same below
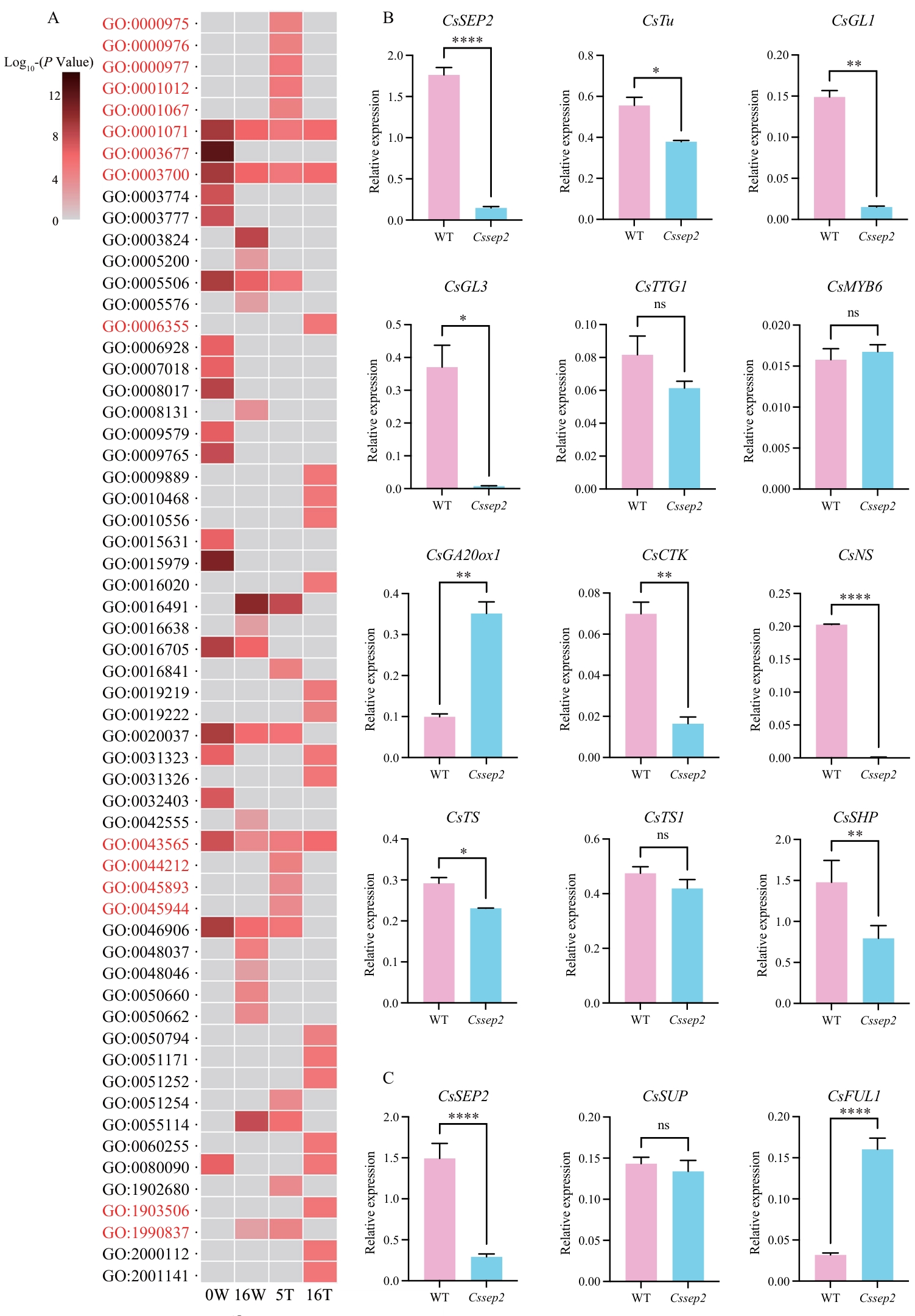
图4 差异表达基因的GO富集分析与RT-qPCR验证A:5T(5 DAA胎座)、16T(16 DAA胎座)、0W(0 DAA外果皮)和16W(16 DAA外果皮)样本中差异表达基因(DEGs)的GO富集分析结果;与DNA结合和转录调控相关的GO条目以红色高亮标注;B:WT和Cssep2突变体0 DAA外果皮组织中关键基因的表达分析;C:WT和Cssep2突变体5 DAA胎座组织中关键基因的表达分析;**** P < 0.000 1;** P < 0.01;* P < 0.05;ns:无显著差异
Fig. 4 GO enrichment and RT-qPCR analysis of differentially expressed genesA: GO enrichment analysis of DEGs at 5T (placenta at 5 DAA), 16T (placenta at 16 DAA), 0W (epicarp at 0 DAA) and 16W (epicarp at 16 DAA). GO terms related with DNA binding and regulation of transcription are highlighted with red. B: Expression analysis of genes in the epicarp tissues of WT and Cssep2 mutants. C: Expression analysis of genes in the placenta tissues of WT and Cssep2 mutants; **** P < 0.000 1; ** P < 0.01; * P < 0.05; ns: not significant difference
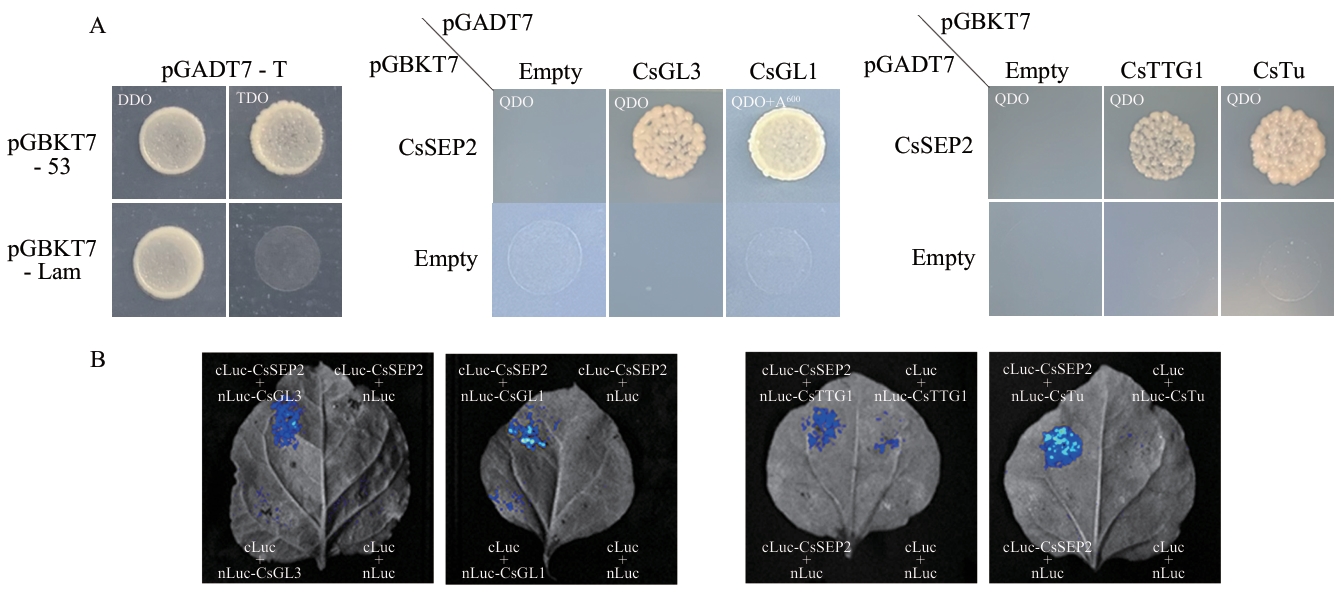
图5 与CsSEP2互作的蛋白鉴定A:酵母双杂交实验结果;QDO:四缺培养基(SD/-Leu-Trp-His-Ade),A600:终浓度为600 ng/mL的Aureobasidin A抗性筛选;B:萤火虫荧光素酶互补成像结果
Fig. 5 Identification of proteins interacting with CsSEP2A: Yeast two-hybrid assay results. QDO: Quadruple dropout, SD/-Leu-Trp-His-Ade. A600: Addition of Aureobasidin A with a final concentration of 600 ng/mL. B: Firefly luciferase complementation imaging analysis
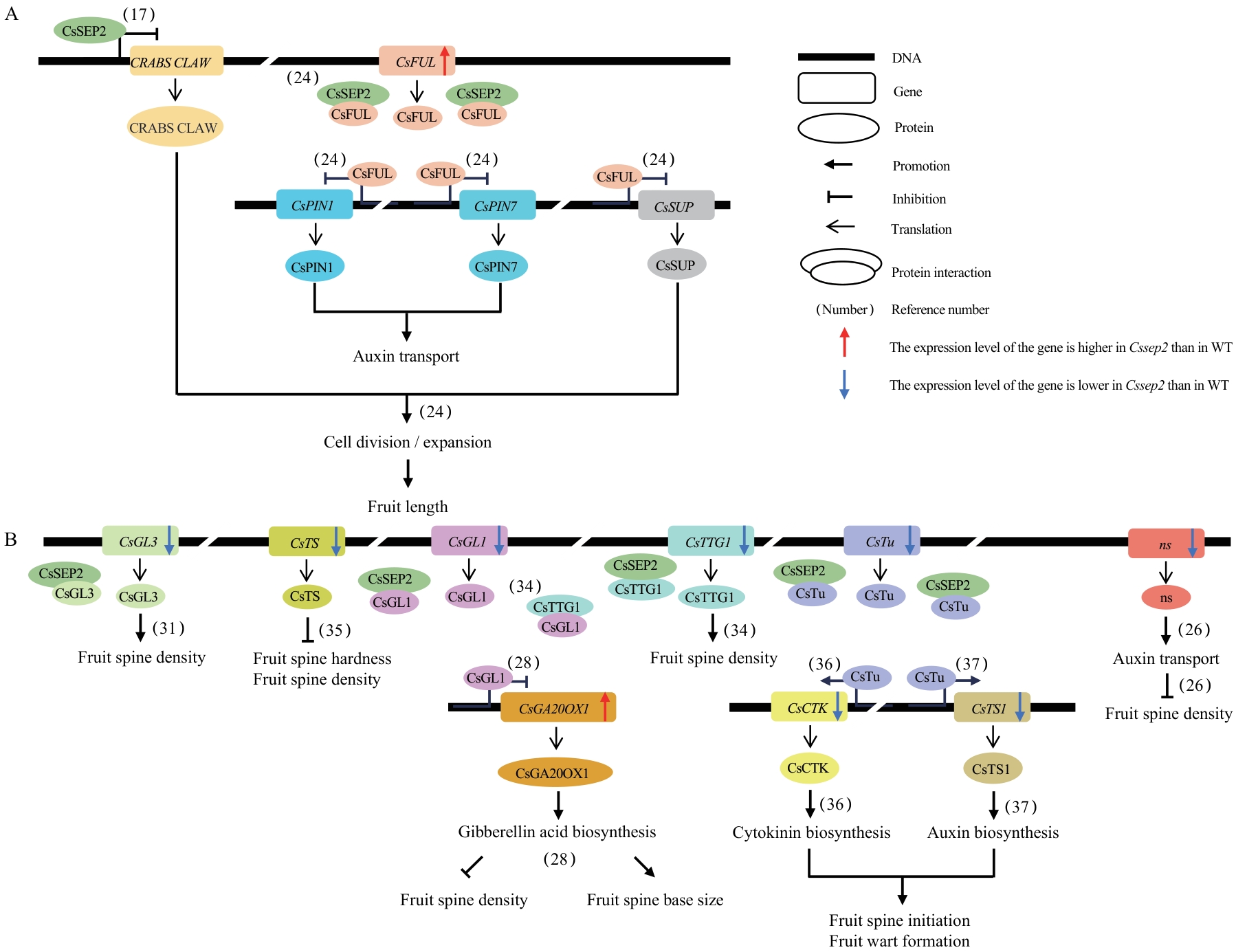
图6 CsSEP2调控果实相关性状的分子网络A、B:CsSEP2参与调控果实发育(A)及果疣性状(B)的分子网络示意图。图例位于右上角。CsSEP2通过抑制CRABS CLAW的表达负调控黄瓜果实长度[17];CsFUL通过抑制CsPIN1、CsPIN7和CsSUP的表达调控果实长度,且可与CsSEP2形成蛋白复合体发挥功能[21]。值得注意的是,Cssep2突变体中CsFUL和CsSUP的表达水平均发生显著变化。与WT相比,Cssep2突变体中CsGL3[22]、CsTS[23]、CsGL1[24]、CsTTG1[25]、CsTu[26-27]和ns[28]的表达水平下调,这些基因已被证实参与果刺与疣状结构的形成和发育调控。其中,ns基因编码生长素转运蛋白,负调控果刺密度[28];CsGL1不仅抑制CsGA20OX1的表达,影响赤霉素合成并改变果刺密度和果刺基部大小[24],还能与CsTTG1互作形成复合体调控果刺密度[25];CsTu通过促进CsCTK[26]和CsTS1[27]的表达,调节细胞分裂素和生长素合成,进而促进果刺起始和疣状结构形成。在Cssep2突变体中,CsGA20OX1、CsCTK和CsTS1的表达水平也发生不同程度的改变。此外,本研究证实CsSEP2可与CsGL3、CsGL1、CsTTG1和CsTu发生蛋白互作
Fig. 6 Molecular network of fruit-related traits regulated by CsSEP2A, B: Schematic diagram of the CsSEP2-invovled molecular network of fruit development (A) and warty traits (B). The captions of the figure are in the upper right corner. CsSEP2 inhibits the lengths of cucumber fruits by suppressing the expression of CRABS CLAW[17]. CsFUL regulates fruit length by inhibiting the expressions of CsPIN1, CsPIN7 and CsSUP, and can also form protein complex with CsSEP2 to exert its functions[21]. Notably, the expressions of both CsFUL and CsSUP changed significantly in Cssep2. Compared to WT, the expressions of CsGL3[22], CsTS[23], CsGL1[24], CsTTG1[25], CsTu[26-27], and ns[28] were lower in Cssep2 mutants. These genes have been confirmed to be involved in the regulation of the formation and development of fruit spines and warts. Among them, ns encodes an auxin transporter and inhibits the density of fruit spines[28]. CsGL1 can not only inhibit the expression of CsGA20OX1, affecting the synthesis of GA and altering the density of fruit spines and the size of the fruit spine base[24], but also interacts with CsTTG1 to form a complex to regulate the density of fruit spines[25]. CsTu promotes the expressions of CsCTK[26] and CsTS1[27], altering the synthesis of cytokinin and auxin and facilitating the initiation of fruit spines and the formation of fruit warts. In Cssep2 mutants, the expressions of CsGA20OX1, CsCTK, and CsTS1 also changed to varying degrees. In addition, this study further revealed that CsSEP2 interacts with CsGL3, CsGL1, CsTTG,1 and CsTu
| [1] | Bowman JL, Moyroud E. Reflections on the ABC model of flower development [J]. Plant Cell, 2024, 36(5): 1334-1357. |
| [2] | Lin ZY, Zhu GF, Lu CQ, et al. Functional conservation and divergence of SEPALLATA-like genes in floral development in Cymbidium sinense [J]. Front Plant Sci, 2023, 14: 1209834. |
| [3] | Ditta G, Pinyopich A, Robles P, et al. The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity [J]. Curr Biol, 2004, 14(21): 1935-1940. |
| [4] | Jack T. Relearning our ABCs: new twists on an old model [J]. Trends Plant Sci, 2001, 6(7): 310-316. |
| [5] | Tan XM, Li YR, Song MR, et al. The molecular mechanism of interaction between SEPALLATA3 and APETALA1 in Arabidopsis thaliana [J]. Plant Direct, 2025, 9(4): e70052. |
| [6] | Hugouvieux V, Blanc-Mathieu R, Janeau A, et al. SEPALLATA-driven MADS transcription factor tetramerization is required for inner whorl floral organ development [J]. Plant Cell, 2024, 36(9): 3435-3450. |
| [7] | Gan ZC, Wu XX, Biahomba SAM, et al. Genome-wide identification, evolution, and expression characterization of the pepper (Capsicum spp.) MADS-box gene family [J]. Genes, 2022, 13(11): 2047. |
| [8] | Cui ZB, Wang XW, Dai YD, et al. Transcription factor OsNF-YC1 regulates grain size by coordinating the transcriptional activation of OsMADS1 in Oryza sativa L [J]. Plant J, 2024, 119(3): 1465-1480. |
| [9] | Ampomah-Dwamena C, Morris BA, Sutherland P, et al. Down-regulation of TM29, a tomato SEPALLATA homolog, causes parthenocarpic fruit development and floral reversion [J]. Plant Physiol, 2002, 130(2): 605-617. |
| [10] | Vallarino JG, Merchante C, Sánchez-Sevilla JF, et al. Characterizing the involvement of FaMADS9 in the regulation of strawberry fruit receptacle development [J]. Plant Biotechnol J, 2020, 18(4): 929-943. |
| [11] | Ireland HS, Yao JL, Tomes S, et al. Apple SEPALLATA1/2-like genes control fruit flesh development and ripening [J]. Plant J, 2013, 73(6): 1044-1056. |
| [12] | Upadhyay RK, Tucker ML, Mattoo AK. Ethylene and RIPENING INHIBITOR modulate expression of SlHSP17.7A, B class I small heat shock protein genes during tomato fruit ripening [J]. Front Plant Sci, 2020, 11: 975. |
| [13] | Ito Y, Kitagawa M, Ihashi N, et al. DNA-binding specificity, transcriptional activation potential, and the rin mutation effect for the tomato fruit-ripening regulator RIN [J]. Plant J, 2008, 55(2): 212-223. |
| [14] | Zhou Y, Hu LF, Song JB, et al. Isolation and characterization of a MADS-box gene in cucumber (Cucumis sativus L.) that affects flowering time and leaf morphology in transgenic Arabidopsis [J]. Biotechnol Biotechnol Equip, 2019, 33(1): 54-63. |
| [15] | Cheng YW, He P, Jiang LW, et al. Identification and characterization of a SEPALLATA-like MADS-box gene from cucumber (Cucumis sativus L.) [J]. Not Bot Horti Agrobo, 2019, 47(4): 1168-1177. |
| [16] | Wang X, Gao DL, Sun JJ, et al. An exon skipping in a SEPALLATA-Like gene is associated with perturbed floral and fruits development in cucumber [J]. J Integr Plant Biol, 2016, 58(9): 766-771. |
| [17] | Song WY, Xie Y, Liu B, et al. Single nucleotide polymorphisms in SEPALLATA 2 underlie fruit length variation in cucurbits [J]. Plant Cell, 2024, 36(10): 4607-4621. |
| [18] | Xin TX, Tian HJ, Ma YL, et al. Targeted creating new mutants with compact plant architecture using CRISPR/Cas9 genome editing by an optimized genetic transformation procedure in cucurbit plants [J]. Hortic Res, 2022, 9: uhab086. |
| [19] | Kanehisa M, Furumichi M, Sato Y, et al. KEGG: biological systems database as a model of the real world [J]. Nucleic Acids Res, 2025, 53(D1): D672-D677. |
| [20] | Wang LP, Yu G, Macho AP, et al. Split-luciferase complementation imaging assay to study protein-protein interactions in Nicotiana benthamiana [J]. Bio Protoc, 2021, 11(23): e4237. |
| [21] | Zhao JY, Jiang L, Che G, et al. A functional allele of CsFUL1 regulates fruit length through repressing CsSUP and inhibiting auxin transport in cucumber [J]. Plant Cell, 2019, 31(6): 1289-1307. |
| [22] | Pan YP, Bo KL, Cheng ZH, et al. The loss-of-function GLABROUS 3 mutation in cucumber is due to LTR-retrotransposon insertion in a class IV HD-ZIP transcription factor gene CsGL3 that is epistatic over CsGL1 [J]. BMC Plant Biol, 2015, 15: 302. |
| [23] | Guo CL, Yang XQ, Wang YL, et al. Identification and mapping of ts (tender spines), a gene involved in soft spine development in Cucumis sativus [J]. Theor Appl Genet, 2018, 131(1): 1-12. |
| [24] | Qiang Li CC. The identification of Cucumis sativus Glabrous 1 (CsGL1) required for the formation of trichomes uncovers a novel function for the homeodomain-leucine zipper I gene [J]. J Exp Bot, 2015, 66(9): 2515-2526. |
| [25] | Chunhua Chen SY. The WD-repeat protein CsTTG1 regulates fruit wart formation through interaction with the homeodomain-leucine zipper I protein mict [J]. Plant Physiol, 2016, 171(2): 1156-1168. |
| [26] | Yang XQ, Zhang WW, He HL, et al. Tuberculate fruit gene Tu encodes a C2H2 zinc finger protein that is required for the warty fruit phenotype in cucumber (Cucumis sativus L.) [J]. Plant J, 2014, 78(6): 1034-1046. |
| [27] | Yang S, Wen CL, Liu B, et al. A CsTu-TS1 regulatory module promotes fruit tubercule formation in cucumber [J]. Plant Biotechnol J, 2019, 17(1): 289-301. |
| [28] | Xie Q, Liu PN, Shi LX, et al. Combined fine mapping, genetic diversity, and transcriptome profiling reveals that the auxin transporter gene ns plays an important role in cucumber fruit spine development [J]. Theor Appl Genet, 2018, 131(6): 1239-1252. |
| [29] | Zhang ZH, Zou WH, Lin PX, et al. Evolution and function of MADS-box transcription factors in plants [J]. Int J Mol Sci, 2024, 25(24): 13278. |
| [30] | Lai XL, Stigliani A, Lucas J, et al. Genome-wide binding of SEPALLATA3 and AGAMOUS complexes determined by sequential DNA-affinity purification sequencing [J]. Nucleic Acids Res, 2020, 48(17): 9637-9648. |
| [31] | Lai XL, Vega-Léon R, Hugouvieux V, et al. The intervening domain is required for DNA-binding and functional identity of plant MADS transcription factors [J]. Nat Commun, 2021, 12(1): 4760. |
| [32] | Yang S, Cai YL, Liu XW, et al. A CsMYB6-CsTRY module regulates fruit trichome initiation in cucumber [J]. J Exp Bot, 2018, 69(8): 1887-1902. |
| [33] | Chen CH, Liu ML, Jiang L, et al. Transcriptome profiling reveals roles of meristem regulators and polarity genes during fruit trichome development in cucumber (Cucumis sativus L.) [J]. J Exp Bot, 2014, 65(17): 4943-4958. |
| [34] | Zhao JL, Pan JS, Guan Y, et al. Micro-trichome as a class I homeodomain-leucine zipper gene regulates multicellular trichome development in Cucumis sativus [J]. J Integr Plant Biol, 2015, 57(11): 925-935. |
| [35] | Cui JY, Miao H, Ding LH, et al. A new glabrous gene (csgl3) identified in trichome development in cucumber (Cucumis sativus L.) [J]. PLoS One, 2016, 11(2): e0148422. |
| [36] | Wang YL, Nie JT, Chen HM, et al. Identification and mapping of Tril, a homeodomain-leucine zipper gene involved in multicellular trichome initiation in Cucumis sativus [J]. Theor Appl Genet, 2016, 129(2): 305-316. |
| [37] | Zhang HY, Wang LN, Zheng SS, et al. A fragment substitution in the promoter of CsHDZIV11/CsGL3 is responsible for fruit spine density in cucumber (Cucumis sativus L.) [J]. Theor Appl Genet, 2016, 129(7): 1289-1301. |
| [38] | Ferrándiz C, Fourquin C. Role of the FUL-SHP network in the evolution of fruit morphology and function [J]. J Exp Bot, 2014, 65(16): 4505-4513. |
| [39] | Ferrándiz C, Liljegren SJ, Yanofsky MF. Negative regulation of the SHATTERPROOF genes by FRUITFULL during Arabidopsis fruit development [J]. Science, 2000, 289(5478): 436-438. |
| [40] | Cheng ZH, Zhuo SB, Liu XF, et al. The MADS-box gene CsSHP participates in fruit maturation and floral organ development in cucumber [J]. Front Plant Sci, 2019, 10: 1781. |
| [41] | Wang LN, Cao CX, Zheng SS, et al. Transcriptomic analysis of short-fruit 1 (sf1) reveals new insights into the variation of fruit-related traits in Cucumis sativus [J]. Sci Rep, 2017, 7(1): 2950. |
| [42] | Wang J, Tian PP, Sun JJ, et al. CsMYC2 is involved in the regulation of phenylpropanoid biosynthesis induced by trypsin in cucumber (Cucumis sativus) during storage [J]. Plant Physiol Biochem, 2023, 196: 65-74. |
| [1] | 尹跃, 秦小雅, 米佳, 安巍, 何军, 张锋锋. 枸杞FBN基因家族鉴定及与类胡萝卜素代谢的相关性分析[J]. 生物技术通报, 2026, 42(3): 338-348. |
| [2] | 李天源, 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏. 欧李SPL基因家族的鉴定及在果实发育过程中的表达分析[J]. 生物技术通报, 2026, 42(3): 362-373. |
| [3] | 刘娜, 曾宝珍, 贾兆星, 祝英方. 表观遗传调控番茄果实发育及成熟的研究进展[J]. 生物技术通报, 2026, 42(3): 37-47. |
| [4] | 胡秋玲, 陈灵, 黄嘉怡, 赵梓乔, 潘璐怡, 刘慧丽, 刘太波. 多胺调控果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 203-212. |
| [5] | 马世杰, 李铮, 李蔚, 郭仰东, 张娜. 光信号调控园艺作物果实发育的研究进展[J]. 生物技术通报, 2026, 42(3): 5-18. |
| [6] | 张高翔, 吴玉碧, 郭亚静, 纪薇, 杨忠义. 葡萄WD40基因家族鉴定及表达量分析[J]. 生物技术通报, 2026, 42(3): 324-337. |
| [7] | 李成泉, 史庆华, 杨晓玉. 园艺作物果实发育的miRNA调控网络:从分子机制到种质创新[J]. 生物技术通报, 2026, 42(3): 19-36. |
| [8] | 龙林茜, 曾银萍, 王茜, 邓玉萍, 葛敏茜, 陈彦灼, 李鑫娟, 杨军, 邹建. 向日葵GH3基因家族鉴定及其在花发育中的功能分析[J]. 生物技术通报, 2026, 42(1): 125-138. |
| [9] | 刘佳丽, 宋经荣, 赵文宇, 张馨元, 赵子洋, 曹一博, 张凌云. 蓝莓R2R3-MYB基因鉴定及类黄酮调控基因表达分析[J]. 生物技术通报, 2025, 41(9): 124-138. |
| [10] | 豆飞飞, 任毓昭, 王石磊, 刘春颖, 王晓东, 王昭懿, 刘彩霞, 刘凤楼, 王掌军, 李清峰. 宁春4号小麦EMS突变体库的构建及表型变异分析[J]. 生物技术通报, 2025, 41(8): 92-101. |
| [11] | 王斌, 李健荣, 占朝霞, 袁晓. CsGR-RBP3克隆及其在采后黄瓜耐冷性中的功能[J]. 生物技术通报, 2025, 41(6): 155-166. |
| [12] | 罗嗣芳, 张祖铭, 谢丽芳, 郭紫晶, 陈兆星, 杨月华, 严翔, 张洪铭. 山金柑GATA基因家族全基因组鉴定及在果实发育中的表达分析[J]. 生物技术通报, 2025, 41(5): 218-230. |
| [13] | 杜量衡, 唐黄磊, 张治国. 控制水稻光响应基因ELM1的图位克隆[J]. 生物技术通报, 2025, 41(5): 82-89. |
| [14] | 杨宗辉, 李利斌, 孟昭娟, 高天, 祝利霞, 杜海梅, 董伟伟, 曹齐卫. 比较转录组学揭示乙烯信号与表观遗传协同调控黄瓜性别决定[J]. 生物技术通报, 2025, 41(12): 139-155. |
| [15] | 庞梦真, 徐汉琴, 刘海燕, 宋娟, 王佳涵, 孙丽娜, 姬佩梅, 尹泽芝, 胡又川, 赵晓萌, 梁闪闪, 张泗举, 栾维江. 水稻黄化早抽穗突变体 hz1 的基因鉴定及功能分析[J]. 生物技术通报, 2024, 40(7): 125-136. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||