








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (2): 197-206.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0890
Previous Articles Next Articles
WANG Shang-feng1( ), CHENG Bin2,3, WANG Ruo-ruo3,4, DING Yan-qing2,3, XU Jian-xia2,3, CAO Ning2,3, GAO Xu2,3, LI Wen-zhen2,3, ZHANG Li-yi2,3(
), CHENG Bin2,3, WANG Ruo-ruo3,4, DING Yan-qing2,3, XU Jian-xia2,3, CAO Ning2,3, GAO Xu2,3, LI Wen-zhen2,3, ZHANG Li-yi2,3( )
)
Received:2025-08-18
Online:2026-02-26
Published:2026-03-17
Contact:
ZHANG Li-yi
E-mail:619684541@qq.com;lyzhang1997@hotmail.com
WANG Shang-feng, CHENG Bin, WANG Ruo-ruo, DING Yan-qing, XU Jian-xia, CAO Ning, GAO Xu, LI Wen-zhen, ZHANG Li-yi. Gene Mapping of Sorghum Flowering Time and Prediction of Candidate Genes Based on BSA-seq[J]. Biotechnology Bulletin, 2026, 42(2): 197-206.
| 引物名称 Primer name | orward primer (5′‒3′) 正向引物 F | everse primer (5′‒3′) 反向引物 R |
|---|---|---|
| V1f1 | CAACTTCAGTGCAACGGTCG | AGCAAGGATTGTGTGGTGCT |
| V423 | TCAGTGCAACGGTCGACAAA | TGAATGGTGAACAGCCGACC |
| Hd8800 | CACATCAGTGTGTGCCGTTG | TTAGGGGCGTTCTTCTGCTC |
| V5f1 | CATCCACGGGGTTGAAGTCT | AGCGGTCCATTGCTCAAAGT |
Table 1 Primer information of the 4 InDel markers
| 引物名称 Primer name | orward primer (5′‒3′) 正向引物 F | everse primer (5′‒3′) 反向引物 R |
|---|---|---|
| V1f1 | CAACTTCAGTGCAACGGTCG | AGCAAGGATTGTGTGGTGCT |
| V423 | TCAGTGCAACGGTCGACAAA | TGAATGGTGAACAGCCGACC |
| Hd8800 | CACATCAGTGTGTGCCGTTG | TTAGGGGCGTTCTTCTGCTC |
| V5f1 | CATCCACGGGGTTGAAGTCT | AGCGGTCCATTGCTCAAAGT |
| 样本Sample | 平均测序深度Mean depth (X) | 短片段总数Number of total Reads | 总碱基读数Clean bases (bp) | GC (%) | Q30 (%) | 比对Mapped reads (%) |
|---|---|---|---|---|---|---|
| 红缨子 | 67.04 | 369 858 258 | 55 478 738 700 | 44.63 | 91.96 | 99.41 |
| SAP001 | 70.27 | 355 144 466 | 53 271 669 900 | 43.54 | 96.28 | 98.88 |
| 晚熟池 | 77.31 | 387 856 226 | 58 178 433 900 | 43.67 | 97.29 | 99.34 |
| 早熟池 | 86.91 | 443 492 672 | 66 523 900 800 | 43.53 | 97.36 | 98.95 |
Table 2 Sequencing and mapping results of different samples
| 样本Sample | 平均测序深度Mean depth (X) | 短片段总数Number of total Reads | 总碱基读数Clean bases (bp) | GC (%) | Q30 (%) | 比对Mapped reads (%) |
|---|---|---|---|---|---|---|
| 红缨子 | 67.04 | 369 858 258 | 55 478 738 700 | 44.63 | 91.96 | 99.41 |
| SAP001 | 70.27 | 355 144 466 | 53 271 669 900 | 43.54 | 96.28 | 98.88 |
| 晚熟池 | 77.31 | 387 856 226 | 58 178 433 900 | 43.67 | 97.29 | 99.34 |
| 早熟池 | 86.91 | 443 492 672 | 66 523 900 800 | 43.53 | 97.36 | 98.95 |
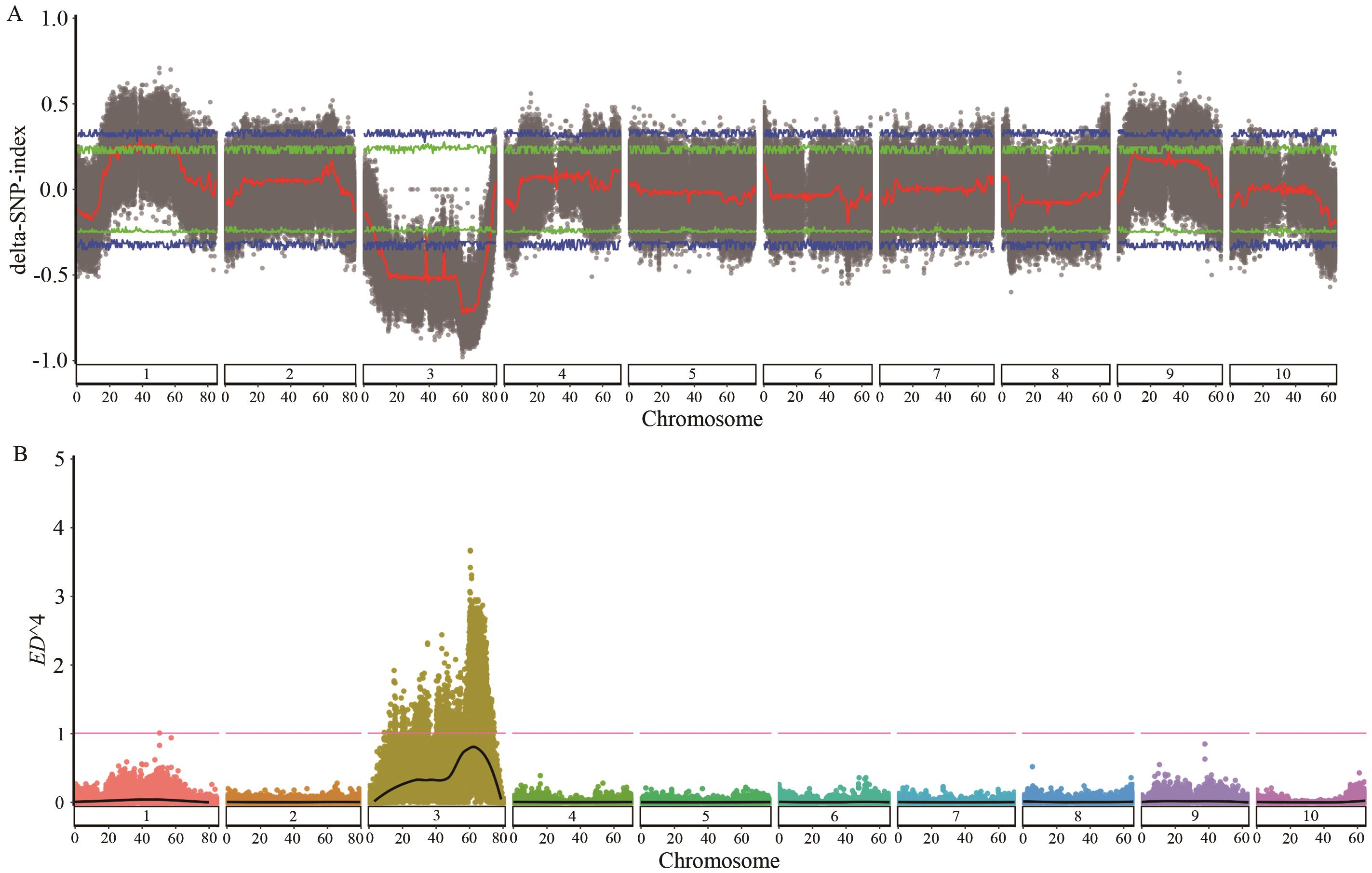
Fig. 1 Distribution of SNP-index and ED4 association values on chromosome 3A: Manhattan plot generated based on SNP-index algorithm. The blue and green lines indicate the 99% and 95% confidence intervals, respectively. The black dots denote the ΔSNP-index for each SNP, and the red line is the fitted curve of the mean values of ΔSNP-index derived from the sliding window analysis. B: Manhattan plot generated based on ED algorithm. The red line indicates the 99% threshold; and the black line is the fitted curve of the ED4 values
基因 Gene | 变异位置 Variation position (bp) | 参考基因组 Reference | 等位变异Alternative | CDS突变类型Type of CDS mutation | 启动子突变Promoter mutation |
|---|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 930 660 | A | - | - | 碱基缺失 |
| SbiHYZ.03G216700 | 59 972 784 | C | A | 非同义突变 | |
| SbiHYZ.03G216800 | 59 974 642 | C | A | - | 碱基突变 |
| SbiHYZ.03G216900 | 59 980 499 | A | - | 缺失突变/移码 | |
| 59 980 657 | G | A | 非同义突变 | ||
| 59 980 792 | G | T | 非同义突变 | ||
| 59 980 816 | G | C | 非同义突变 | ||
| SbiHYZ.03G217000 | 60 060 314 | A | - | - | 碱基缺失 |
| SbiHYZ.03G217200 | 60 093 586 | A | C | 非同义突变 | |
| SbiHYZ.03G217700 | 60 140 432 | G | A | 非同义突变 | |
| SbiHYZ.03G218800 | 60 365 936 | C | T | - | 碱基突变 |
| SbiHYZ.03G218900 | 60 375 442 | C | T | 非同义突变 |
Table 3 Sequence variation analysis of nine candidate genes
基因 Gene | 变异位置 Variation position (bp) | 参考基因组 Reference | 等位变异Alternative | CDS突变类型Type of CDS mutation | 启动子突变Promoter mutation |
|---|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 930 660 | A | - | - | 碱基缺失 |
| SbiHYZ.03G216700 | 59 972 784 | C | A | 非同义突变 | |
| SbiHYZ.03G216800 | 59 974 642 | C | A | - | 碱基突变 |
| SbiHYZ.03G216900 | 59 980 499 | A | - | 缺失突变/移码 | |
| 59 980 657 | G | A | 非同义突变 | ||
| 59 980 792 | G | T | 非同义突变 | ||
| 59 980 816 | G | C | 非同义突变 | ||
| SbiHYZ.03G217000 | 60 060 314 | A | - | - | 碱基缺失 |
| SbiHYZ.03G217200 | 60 093 586 | A | C | 非同义突变 | |
| SbiHYZ.03G217700 | 60 140 432 | G | A | 非同义突变 | |
| SbiHYZ.03G218800 | 60 365 936 | C | T | - | 碱基突变 |
| SbiHYZ.03G218900 | 60 375 442 | C | T | 非同义突变 |
基因 Gene | 物理位置 Physical position (bp) | 同源基因 Paralogous gene | 功能注释 Functional annotation | 相似率 Similarity (%) |
|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 931 181‒59 935 674 | LOC_Os01g41250 | F-box/FBD结构域 F-box/FBD domain | 66 |
| SbiHYZ.03G216700 | 59 971 690‒59 974 228 | LOC_Os01g41260 | F-box/FBD结构域 F-box/FBD domain | 63 |
| SbiHYZ.03G216800 | 59 974 820‒59 977 137 | Zm00001d011276 | F-box/FBD结构域 F-box/FBD domain | 80 |
| SbiHYZ.03G216900 | 59 978 600‒59 981 036 | LOC_Os01g41310 | F-box/FBD结构域 F-box/FBD domain | 51 |
| SbiHYZ.03G217000 | 60 061 333‒60 063 755 | Zm00001d044385 | F-box/FBD结构域 F-box/FBD domain | 83 |
| SbiHYZ.03G217200 | 60 090 312‒60 093 766 | Zm00001d044386 | F-box/FBD结构域 F-box/FBD domain | 75 |
| SbiHYZ.03G217700 | 60 139 306‒60 141 819 | LOC_Os01g41530 | F-box/FBD结构域 F-box/FBD domain | 50 |
| SbiHYZ.03G218800 | 60 366 296‒60 367 982 | ZM00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡchlorophyll a/b-binding protein | 97 |
| SbiHYZ.03G218900 | 60 374 509‒60 375 642 | Zm00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡ chlorophyll a/b-binding protein | 98 |
Table 4 Information of nine candidate genes
基因 Gene | 物理位置 Physical position (bp) | 同源基因 Paralogous gene | 功能注释 Functional annotation | 相似率 Similarity (%) |
|---|---|---|---|---|
| SbiHYZ.03G216400 | 59 931 181‒59 935 674 | LOC_Os01g41250 | F-box/FBD结构域 F-box/FBD domain | 66 |
| SbiHYZ.03G216700 | 59 971 690‒59 974 228 | LOC_Os01g41260 | F-box/FBD结构域 F-box/FBD domain | 63 |
| SbiHYZ.03G216800 | 59 974 820‒59 977 137 | Zm00001d011276 | F-box/FBD结构域 F-box/FBD domain | 80 |
| SbiHYZ.03G216900 | 59 978 600‒59 981 036 | LOC_Os01g41310 | F-box/FBD结构域 F-box/FBD domain | 51 |
| SbiHYZ.03G217000 | 60 061 333‒60 063 755 | Zm00001d044385 | F-box/FBD结构域 F-box/FBD domain | 83 |
| SbiHYZ.03G217200 | 60 090 312‒60 093 766 | Zm00001d044386 | F-box/FBD结构域 F-box/FBD domain | 75 |
| SbiHYZ.03G217700 | 60 139 306‒60 141 819 | LOC_Os01g41530 | F-box/FBD结构域 F-box/FBD domain | 50 |
| SbiHYZ.03G218800 | 60 366 296‒60 367 982 | ZM00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡchlorophyll a/b-binding protein | 97 |
| SbiHYZ.03G218900 | 60 374 509‒60 375 642 | Zm00001d011285 | 光捕获复合物Ⅱ叶绿素a/b结合蛋白 Light-harvesting complexⅡ chlorophyll a/b-binding protein | 98 |
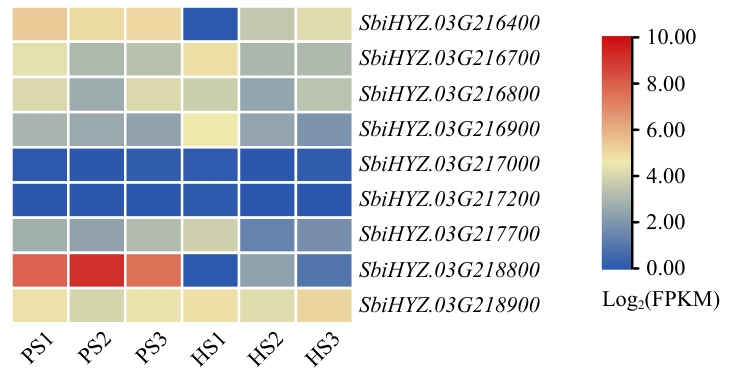
Fig. 3 Expressions of candidate genes in the parent in different tissues at different timesHeatmap of candidate gene expression based on FPKM values from RNA-seq analysis after log2 transformation.S1: Booting stage. S2: 5 cm heading. S3: First flowering day. P: SAP001. H: Hongyingzi
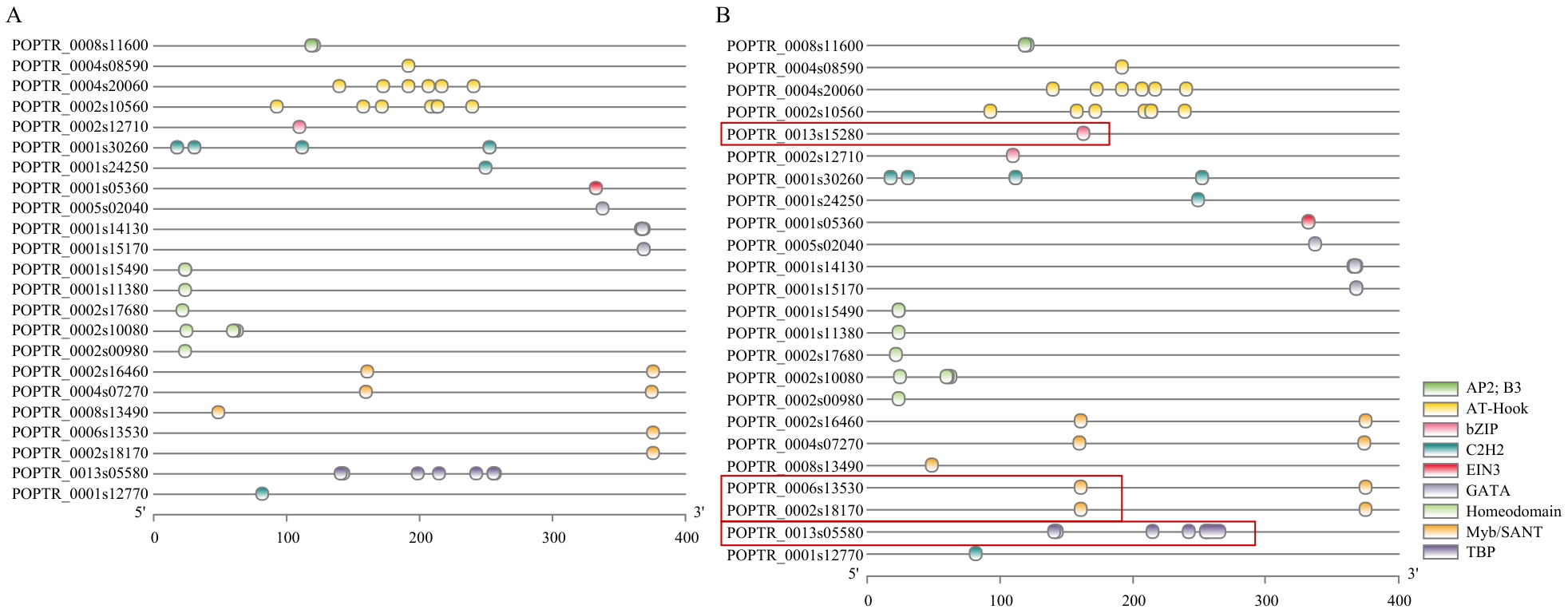
Fig. 4 Comparison of cis-elements in the promoter regions of key candidate genes in parentA: Visualized cis-acting elements in the promoter region of Hongyingzi. B: Visualized cis-acting elements in the promoter region of SAP001, with red boxes highlighting element divergence
| [1] | 王艳秋, 张飞, 朱凯, 等. 抗旱型高粱开花前期和后期应对水分亏缺的生理调节研究 [J]. 山西农业大学学报: 自然科学版, 2020, 40(3): 37-44. |
| Wang YQ, Zhang F, Zhu K, et al. Physiological response to water-deficience of drought-tolerant sorghum variety prior to and post of flowering [J]. J Shanxi Agric Univ Nat Sci Ed, 2020, 40(3): 37-44. | |
| [2] | 高士杰, 陈冰嬬, 李继洪. 对粒用高粱育种的思考 [J]. 现代农业科技, 2013(10): 31-32, 37. |
| Gao SJ, Chen BR, Li JH. Thoughts on breeding of grain sorghum [J]. Mod Agric Sci Technol, 2013(10): 31-32, 37. | |
| [3] | 陈冰嬬, 李继洪, 王阳, 等. 高粱(Sorghum bicolor (L.) Moench)种质资源研究进展 [J]. 西北农林科技大学学报: 自然科学版, 2013, 41(1): 67-72, 77. |
| Chen BR, Li JH, Wang Y, et al. Advances in germplasm resources of sorghum (Sorghum bicolor (L.) Moench) [J]. J Northwest A&F Univ Nat Sci Ed, 2013, 41(1): 67-72, 77. | |
| [4] | 丁延庆, 徐建霞, 汪灿, 等. 基于Super-GBS技术的高粱籽粒酿造相关性状QTL定位 [J]. 核农学报, 2023, 37(2): 241-250. |
| Ding YQ, Xu JX, Wang C, et al. QTL mapping of grain traits related to brewing in sorghum based on super-GBS technology [J]. J Nucl Agric Sci, 2023, 37(2): 241-250. | |
| [5] | 阳志锐, 冉红艳. 贵州省酱香型白酒全产业链发展现状及能级提升路径研究 [J]. 中国酿造, 2025, 44(3): 285-291. |
| Yang ZR, Ran HY. Development status and capacity enhancement path of the whole industry chain of sauce-flavor Baijiu in Guizhou province [J]. China Brew, 2025, 44(3): 285-291. | |
| [6] | Mace ES, Hunt CH, Jordan DR. Supermodels: sorghum and maize provide mutual insight into the genetics of flowering time [J]. Theor Appl Genet, 2013, 126(5): 1377-1395. |
| [7] | Upadhyaya HD, Reddy KN, Vetriventhan M, et al. Latitudinal adaptation of flowering response to photoperiod and temperature in the world collection of sorghum landraces [J]. Crop Sci, 2018, 58(3): 1265-1276. |
| [8] | Rooney WL, Aydin S. Genetic control of a photoperiod-sensitive response in Sorghum bicolor (L.) moench [J]. Crop Sci, 1999, 39(2): 397-400. |
| [9] | Smith CW, Frederiksen RA. Sorghum: Origin, History, Technology, and Production[M]. Wiley, 2000: 240-242. |
| [10] | Murphy RL, Klein RR, Morishige DT, et al. Coincident light and clock regulation of pseudoresponse regulator protein 37 (PRR37) controls photoperiodic flowering in sorghum [J]. Proc Natl Acad Sci USA, 2011, 108(39): 16469-16474. |
| [11] | Casto AL, Mattison AJ, Olson SN, et al. Maturity2, a novel regulator of flowering time in Sorghum bicolor, increases expression of SbPRR37 and SbCO in long days delaying flowering [J]. PLoS One, 2019, 14(4): e0212154. |
| [12] | Childs KL, Miller FR, Cordonnier-Pratt MM, et al. The sorghum photoperiod sensitivity gene, Ma3, encodes a phytochrome B [J]. Plant Physiol, 1997, 113(2): 611-619. |
| [13] | Liu HH, Liu HQ, Zhou LN, et al. Parallel domestication of the heading date 1 gene in cereals [J]. Mol Biol Evol, 2015, 32(10): 2726-2737. |
| [14] | Murphy RL, Morishige DT, Brady JA, et al. Ghd7 (Ma6) represses sorghum flowering in long days: Ghd7 alleles enhance biomass accumulation and grain production [J]. Plant Genome, 2014, 7(2): plantgenome2013.11.0040. |
| [15] | Yang SS, Murphy RL, Morishige DT, et al. Sorghum phytochrome B inhibits flowering in long days by activating expression of SbPRR37 and SbGHD7, repressors of SbEHD1, SbCN8 and SbCN12 [J]. PLoS One, 2014, 9(8): e105352. |
| [16] | Song YH, Ito S, Imaizumi T. Similarities in the circadian clock and photoperiodism in plants [J]. Curr Opin Plant Biol, 2010, 13(5): 594-603. |
| [17] | Yang MK, Lin WJ, Xu YR, et al. Flowering-time regulation by the circadian clock: From Arabidopsis to crops [J]. Crop J, 2024, 12(1): 17-27. |
| [18] | Huang XZ, Yang YF, Xu C. Biomolecular condensation programs floral transition to orchestrate flowering time and inflorescence architecture [J]. New Phytol, 2025, 245(1): 88-94. |
| [19] | Kong WQ, Kim C, Zhang D, et al. Genotyping by sequencing of 393 Sorghum bicolor BTx623×IS3620C recombinant inbred lines improves sensitivity and resolution of QTL detection [J]. G3, 2018, 8(8): 2563-2572. |
| [20] | Ding YQ, Wang YL, Xu JX, et al. A telomere-to-telomere genome assembly of Hongyingzi, a sorghum cultivar used for Chinese Baijiu production [J]. Crop J, 2024, 12(2): 635-640. |
| [21] | Bangbol Sangma H. Genetic characterization of flowering time in sorghum [D]. Queensland: The University of Queensland, 2013. |
| [22] | Parh DK. DNA-based markers for ergot resistance in sorghum [D]. Queensland: The University of Queensland, 2005. |
| [23] | Srinivas G, Satish K, Madhusudhana R, et al. Identification of quantitative trait loci for agronomically important traits and their association with genic-microsatellite markers in sorghum [J]. Theor Appl Genet, 2009, 118(8): 1439-1454. |
| [24] | Wang XM, Mace E, Hunt C, et al. Two distinct classes of QTL determine rust resistance in sorghum [J]. BMC Plant Biol, 2014, 14: 366. |
| [25] | Mantilla Perez MB, Zhao J, Yin YH, et al. Association mapping of brassinosteroid candidate genes and plant architecture in a diverse panel of Sorghum bicolor [J]. Theor Appl Genet, 2014, 127(12): 2645-2662. |
| [26] | Guindo D, Teme N, Vaksmann M, et al. Quantitative trait loci for sorghum grain morphology and quality traits: Toward breeding for a traditional food preparation of West-Africa [J]. J Cereal Sci, 2019, 85: 256-272. |
| [27] | Reddy RN, Madhusudhana R, Mohan SM, et al. Mapping QTL for grain yield and other agronomic traits in post-rainy sorghum [Sorghum bicolor (L.) Moench [J]. Theor Appl Genet, 2013, 126(8): 1921-1939. |
| [28] | Upadhyaya HD, Wang YH, Sharma R, et al. SNP markers linked to leaf rust and grain mold resistance in sorghum [J]. Mol Breed, 2013, 32(2): 451-462. |
| [29] | Feltus FA, Hart GE, Schertz KF, et al. Alignment of genetic maps and QTLs between inter- and intra-specific sorghum populations [J]. Theor Appl Genet, 2006, 112(7): 1295-1305. |
| [30] | Bouchet S, Olatoye MO, Marla SR, et al. Increased power to dissect adaptive traits in global sorghum diversity using a nested association mapping population [J]. Genetics, 2017, 206(2): 573-585. |
| [31] | Shiringani AL, Frisch M, Friedt W. Genetic mapping of QTLs for sugar-related traits in a RIL population of Sorghum bicolor L. Moench [J]. Theor Appl Genet, 2010, 121(2): 323-336. |
| [32] | Olatoye MO, Marla SR, Hu ZB, et al. Dissecting adaptive traits with nested association mapping: genetic architecture of inflorescence morphology in sorghum [J]. G3, 2020, 10(5): 1785-1796. |
| [33] | Boycheva I, Vassileva V, Revalska M, et al. Cyclin-like F-box protein plays a role in growth and development of the three model species Medicago truncatula, Lotus japonicus, and Arabidopsis thaliana [J]. Res Rep Biol, 2015, 6: 117-130. |
| [34] | Lechner E, Achard P, Vansiri A, et al. F-box proteins everywhere [J]. Curr Opin Plant Biol, 2006, 9(6): 631-638. |
| [35] | Kuroda H, Takahashi N, Shimada H, et al. Classification and expression analysis of Arabidopsis F-box-containing protein genes [J]. Plant Cell Physiol, 2002, 43(10): 1073-1085. |
| [36] | Cheng CH, Wang ZJ, Ren ZY, et al. SCFAtPP2-B11 modulates ABA signaling by facilitating SnRK2.3 degradation in Arabidopsis thaliana [J]. PLoS Genet, 2017, 13(8): e1006947. |
| [37] | 曾冰洁, 阎晋东, 杨飘, 等. 拟南芥ZTL/FKF1/LKP2蛋白家族功能研究进展 [J]. 生物信息学, 2019, 17(3): 145-150. |
| Zeng BJ, Yan JD, Yang P, et al. Progress of function studies of ZTL/FKF1/LKP2 proteins family in Arabidopsis [J]. Chin J Bioinform, 2019, 17(3): 145-150. | |
| [38] | Zoltowski BD, Imaizumi T. Structure and function of the ZTL/FKF1/LKP2 group proteins in Arabidopsis [J]. Enzymes, 2014, 35: 213-239. |
| [39] | Lee HG, Kim J, Park KH, et al. High-temperature-induced FKF1 accumulation promotes flowering through the dispersion of GI and degradation of SVP [J]. Nat Plants, 2025, 11: 1282-1297. |
| [40] | Hicks KA, Millar AJ, Carré IA, et al. Conditional circadian dysfunction of the Arabidopsis early-flowering 3 mutant [J]. Science, 1996, 274(5288): 790-792. |
| [41] | Makino S, Kiba T, Imamura A, et al. Genes encoding pseudo-response regulators: insight into his-to-asp phosphorelay and circadian rhythm in Arabidopsis thaliana [J]. Plant Cell Physiol, 2000, 41(6): 791-803. |
| [42] | Sreekantan L, Mathiason K, Grimplet J, et al. Differential floral development and gene expression in grapevines during long and short photoperiods suggests a role for floral genes in dormancy transitioning [J]. Plant Mol Biol, 2010, 73(1/2): 191-205. |
| [43] | Alabadí D, Oyama T, Yanovsky MJ, et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock [J]. Science, 2001, 293(5531): 880-883. |
| [44] | Quail PH. Phytochrome photosensory signalling networks [J]. Nat Rev Mol Cell Biol, 2002, 3(2): 85-93. |
| [45] | 彭凌涛. 控制拟南芥和水稻开花时间光周期途径的分子机制 [J]. 植物生理学通讯, 2006, 42(6): 1021-1031. |
| Peng LT. Molecular mechanism of flowering time controlling photoperiod pathway in Arabidopsis and rice [J]. Plant Physiol Commun, 2006, 42(6): 1021-1031. | |
| [46] | 贺军虎, 李唯正, 陈华蕊, 等. ‘金煌’杧果MiCAB2的克隆及表达与花期调控关系分析 [J]. 园艺学报, 2017, 44(7): 1275-1286. |
| He JH, Li WZ, Chen HR, et al. Cloning of the light harvesting chlorophyll a/b binding protein gene (MiCAB2) from ‘Jinhuang’ mango, and correlation analysis between its expression and flowering regulation [J]. Acta Hortic Sin, 2017, 44(7): 1275-1286. | |
| [47] | Zhao YG, Kong H, Guo YL, et al. Light-harvesting chlorophyll a/b-binding protein-coding genes in Jatropha and the comparison with Castor, cassava and Arabidopsis [J]. PeerJ, 2020, 8: e8465. |
| [48] | 阳江华, 张希财, 邹智. 橡胶树捕光叶绿素a/b结合蛋白基因CAB2的克隆与分析 [J]. 西南林业大学学报: 自然科学, 2019, 39(1): 88-94. |
| Yang JH, Zhang XC, Zou Z. Molecular cloning and analysis of HbCAB2, a chlorophyll a/b-binding protein-encoding gene from Hevea brasiliensis [J]. J Southwest For Univ Nat Sci, 2019, 39(1): 88-94. | |
| [49] | Zou Z, Li MY, Jia RZ, et al. Genes encoding light-harvesting chlorophyll a/b-binding proteins in papaya (Carica papaya L.) and insight into lineage-specific evolution in Brassicaceae [J]. Gene, 2020, 748: 144685. |
| [50] | 孔德元. 光周期调控甘菊成花过程中ClRVE基因家族功能分析 [D]. 北京: 北京林业大学, 2022. |
| Kong DY. Functional analysis of ClRVE gene family during photoperiod regulation of chamomile flower formation [D]. Beijing: Beijing Forestry University, 2022. | |
| [51] | Mishal R, Luna-Arias JP. Role of the TATA-box binding protein (TBP) and associated family members in transcription regulation [J]. Gene, 2022, 833: 146581. |
| [1] | ZHANG Chao-chao, HAN Kai-yuan, WANG Tong, CHEN Zhong. Cloning and Functional Analysis of PtoYABBY2 and PtoYABBY12 in Populus tomentosa [J]. Biotechnology Bulletin, 2025, 41(9): 256-264. |
| [2] | HU Lu, WANG Kai, XU Jing-yi, YE Li-hui, WANG Yong-fei, WANG Li-hua, LI Jie-qin. Research Progress in Genetic Transformation Technologies of Maize and Sorghum [J]. Biotechnology Bulletin, 2025, 41(9): 32-43. |
| [3] | WU Xia-ming, ZHOU Chen-ping, YANG Min, XU Ze, KUANG Rui-bin, LIU Chuan-he, HE Han, WEI Yue-rong. Gene Mapping of Plant Height in Papaya Based on BSA-seq [J]. Biotechnology Bulletin, 2025, 41(9): 54-61. |
| [4] | LIU Ze-zhou, DUAN Nai-bin, YUE Li-xin, WANG Qing-hua, YAO Xing-hao, GAO Li-min, KONG Su-ping. Analysis of Wax Components and Screening of Wax-deficient Gene Ggl-1 in Garlic (Allium sativum L.) [J]. Biotechnology Bulletin, 2025, 41(9): 219-231. |
| [5] | YAN Meng-yang, LIANG Xiao-yang, DAI Jun-ang, ZHANG Yan, GUAN Tuan, ZHANG Hui, LIU Liang-bo, SUN Zhi-hua. Screening of Amoxicillin-degrading Bacteria and Study on Its Degradation Mechanisms [J]. Biotechnology Bulletin, 2025, 41(9): 314-325. |
| [6] | LI Ya-qiong, GESANG La-mao, CHEN Qi-di, YANG Yu-huan, HE Hua-zhuan, ZHAO Yao-fei. Heterologous Overexpression of Sorghum SbSnRK2.1 Enhances the Resistance to Salt Stress in Arabidopsis [J]. Biotechnology Bulletin, 2025, 41(8): 115-123. |
| [7] | WANG Yue-chen, HAN Xin-qi, WEI Wen-min, CUI Zhao-lan, LUO Yang-mei, CHEN Peng-ru, WANG Hai-gang, LIU long-long, ZHANG Li, WANG Lun. Biological Basis Study for Grain Shattering in Proso Millet and Identification of Genes Regulating Grain Shattering [J]. Biotechnology Bulletin, 2025, 41(7): 164-171. |
| [8] | ZHAO Qiang, CHEN Si-yu, PENG Fang-li, WANG Can, GAO Jie, ZHOU Ling-bo, ZHANG Guo-bing, JIANG Yu-wen, SHAO Ming-bo. Effects of Intercropping and Nitrogen Application on the Diversity and Functions of Soil Bacteria around Sorghum Rhizosphere [J]. Biotechnology Bulletin, 2025, 41(6): 307-316. |
| [9] | LIU Zhuo-jun, CHAI Wen-ting, REN Yi-le, WANG Xin-yu, ZHU Li-xun, ZHAO Shan-shan, YANG Bo-hui, FAN Jia-li, LI Xin-feng, ZHAO Wei-jun, LYU Jin-hui, ZHANG Chun-lai. Analysis on Expression and DNA Variation of TGA Genes in Sorghum (Sorghum bicolor) in Response to Sporisorium reilianum Infection [J]. Biotechnology Bulletin, 2025, 41(5): 90-103. |
| [10] | PENG Shao-zhi, WANG Deng-ke, ZHANG Xiang, DAI Xiong-ze, XU Hao, ZOU Xue-xiao. Cloning, Expression Characteristics and Functional Verification of the Pepper CaFD1 Gene [J]. Biotechnology Bulletin, 2025, 41(5): 153-164. |
| [11] | LI Xin-peng, ZHANG Wu-han, ZHANG Li, SHU Fu, HE Qiang, GUO Yang, DENG Hua-feng, WANG Yue, SUN Ping-yong. Creation of Rice Mutant by Gamma-ray and Its Molecular Identification [J]. Biotechnology Bulletin, 2025, 41(3): 35-43. |
| [12] | DU Pin-ting, WU Guo-jiang, WANG Zhen-guo, LI Yan, ZHOU Wei, ZHOU Ya-xing. Identification and Expression Analysis of CPP Gene Family in Sorghum [J]. Biotechnology Bulletin, 2025, 41(1): 132-142. |
| [13] | GAO Meng-meng, ZHAO Tian-yu, JIAO Xin-yue, LIN Chun-jing, GUAN Zhe-yun, DING Xiao-yang, SUN Yan-yan, ZHANG Chun-bao. Comparative Transcriptome Analysis of Cytoplasmic Male Sterile Line and Its Restorer Line in Soybean [J]. Biotechnology Bulletin, 2024, 40(7): 137-149. |
| [14] | BAI Zhi-yuan, XU Fei, YANG Wu, WANG Ming-gui, YANG Yu-hua, ZHANG Hai-ping, ZHANG Rui-jun. Transcriptome Analysis of Fertility Transformation in Weakly Restoring Hybrid F1 of Soybean Cytoplasmic Male Sterility [J]. Biotechnology Bulletin, 2024, 40(6): 134-142. |
| [15] | GAO Yu-kun, ZHANG Jian-dong, YANG Pu-yuan, CHEN Dong-ming, WANG Zhi-bo, TIAN Yi-jin, Zakey Eldinn. E. A. Khlid, CUI Jiang-hui, CHANG Jin-hua. Responses of Sorghum Rhizosphere Soil Bacterial Communities to Salt Stress [J]. Biotechnology Bulletin, 2024, 40(4): 203-216. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||