








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 263-274.doi: 10.13560/j.cnki.biotech.bull.1985.2026-0160
Previous Articles Next Articles
LI Ying-hui1( ), WANG Yang-bo-han2, ZHOU Hao-bo3, LU Xin-ru3, ZHANG Ke-xin3, YU Yang4, LI Chuan-you4, SUN Chuan-long2(
), WANG Yang-bo-han2, ZHOU Hao-bo3, LU Xin-ru3, ZHANG Ke-xin3, YU Yang4, LI Chuan-you4, SUN Chuan-long2( )
)
Received:2026-02-01
Online:2026-03-26
Published:2026-04-23
Contact:
SUN Chuan-long
E-mail:liyinghui@hainanu.edu.cn;clsun@sdau.edu.cn
LI Ying-hui, WANG Yang-bo-han, ZHOU Hao-bo, LU Xin-ru, ZHANG Ke-xin, YU Yang, LI Chuan-you, SUN Chuan-long. Identification of VPE Gene Family and Their Functional Analysis under Abiotic Stress in Tomato[J]. Biotechnology Bulletin, 2026, 42(3): 263-274.
基因名称 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids | 分子量 Molecular weight (kD) | 疏水性 GRAVY | 等电点 pI |
|---|---|---|---|---|---|
| SlVPE1 | Solyc08g065790.2.1 | 460 | 51.77 | -0.49 | 6.92 |
| SlVPE2 | Solyc08g065780.1.1 | 451 | 50.09 | -0.12 | 5.76 |
| SlVPE3 | Solyc08g065610.2.1 | 470 | 52.11 | -0.36 | 7.27 |
| SlVPE4 | Solyc08g079160.2.1 | 469 | 52.00 | -0.33 | 7.59 |
| SlVPE5 | Solyc12g095910.1.1 | 463 | 51.04 | -0.20 | 5.05 |
| SlVPE6 | Solyc08g065530.1.1 | 483 | 53.88 | -0.31 | 5.83 |
| SlVPE7 | Solyc08g065570.1.1 | 428 | 47.54 | -0.28 | 7.49 |
| SlVPE8 | Solyc08g065590.1.1 | 446 | 49.35 | -0.30 | 7.26 |
| SlVPE9 | Solyc08g065690.1.1 | 450 | 49.50 | -0.20 | 7.27 |
| SlVPE10 | Solyc08g065710.1.1 | 451 | 50.68 | -0.44 | 7.78 |
| SlVPE11 | Solyc08g065720.1.1 | 482 | 53.22 | -0.34 | 5.67 |
| SlVPE12 | Solyc08g065740.1.1 | 497 | 55.21 | -0.37 | 5.49 |
| SlVPE13 | Solyc08g065750.1.1 | 480 | 52.97 | -0.27 | 6.10 |
Table 1 Basic information about SlVPEs
基因名称 Gene name | 基因ID Gene ID | 氨基酸数量 Number of amino acids | 分子量 Molecular weight (kD) | 疏水性 GRAVY | 等电点 pI |
|---|---|---|---|---|---|
| SlVPE1 | Solyc08g065790.2.1 | 460 | 51.77 | -0.49 | 6.92 |
| SlVPE2 | Solyc08g065780.1.1 | 451 | 50.09 | -0.12 | 5.76 |
| SlVPE3 | Solyc08g065610.2.1 | 470 | 52.11 | -0.36 | 7.27 |
| SlVPE4 | Solyc08g079160.2.1 | 469 | 52.00 | -0.33 | 7.59 |
| SlVPE5 | Solyc12g095910.1.1 | 463 | 51.04 | -0.20 | 5.05 |
| SlVPE6 | Solyc08g065530.1.1 | 483 | 53.88 | -0.31 | 5.83 |
| SlVPE7 | Solyc08g065570.1.1 | 428 | 47.54 | -0.28 | 7.49 |
| SlVPE8 | Solyc08g065590.1.1 | 446 | 49.35 | -0.30 | 7.26 |
| SlVPE9 | Solyc08g065690.1.1 | 450 | 49.50 | -0.20 | 7.27 |
| SlVPE10 | Solyc08g065710.1.1 | 451 | 50.68 | -0.44 | 7.78 |
| SlVPE11 | Solyc08g065720.1.1 | 482 | 53.22 | -0.34 | 5.67 |
| SlVPE12 | Solyc08g065740.1.1 | 497 | 55.21 | -0.37 | 5.49 |
| SlVPE13 | Solyc08g065750.1.1 | 480 | 52.97 | -0.27 | 6.10 |
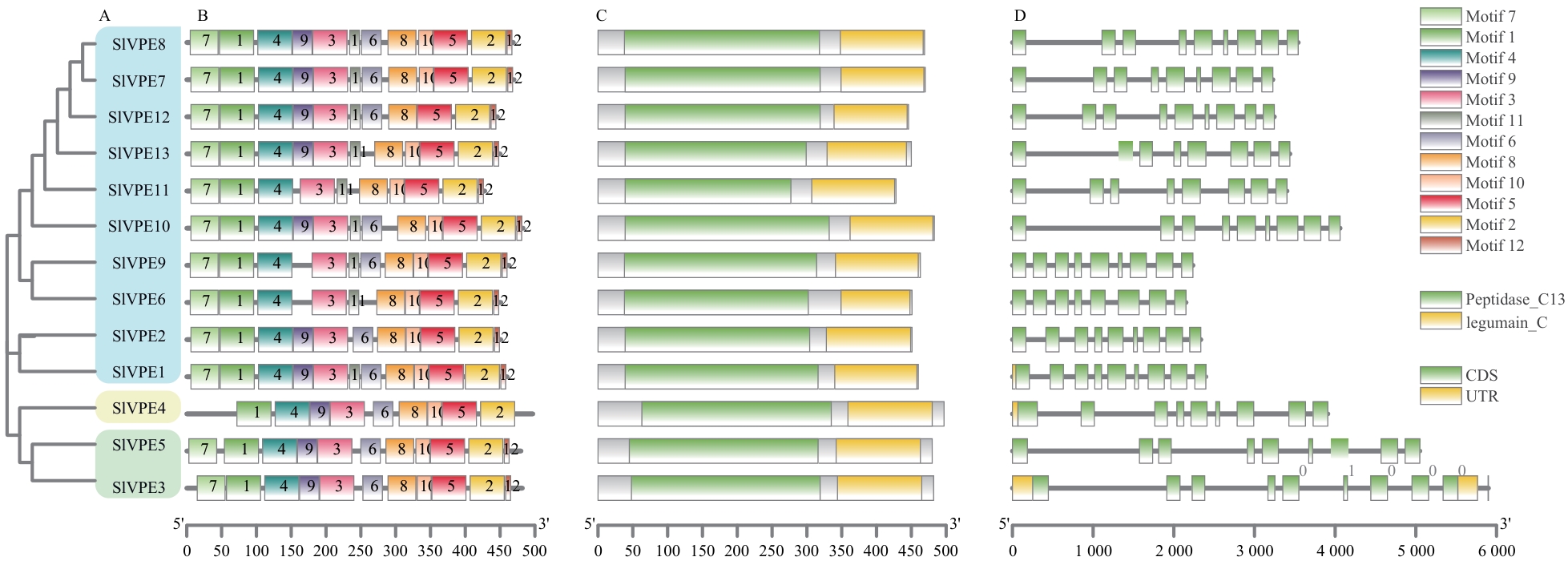
Fig. 2 Motifs, conserved domains and gene structure analysis of SlVPE proteinA: Phylogenetic tree of SlVPE proteins. B: Analysis of conserved motifs in SlVPE proteins. C: Analysis of conserved domains of SlVPE proteins, with green and yellow boxes representing Peptidase_C13 and legumain_C domains, respectively. D: Analysis of SlVPE gene structures. CDS, UTRs, and introns are indicated by green boxes, yellow boxes, and gray lines, respectively
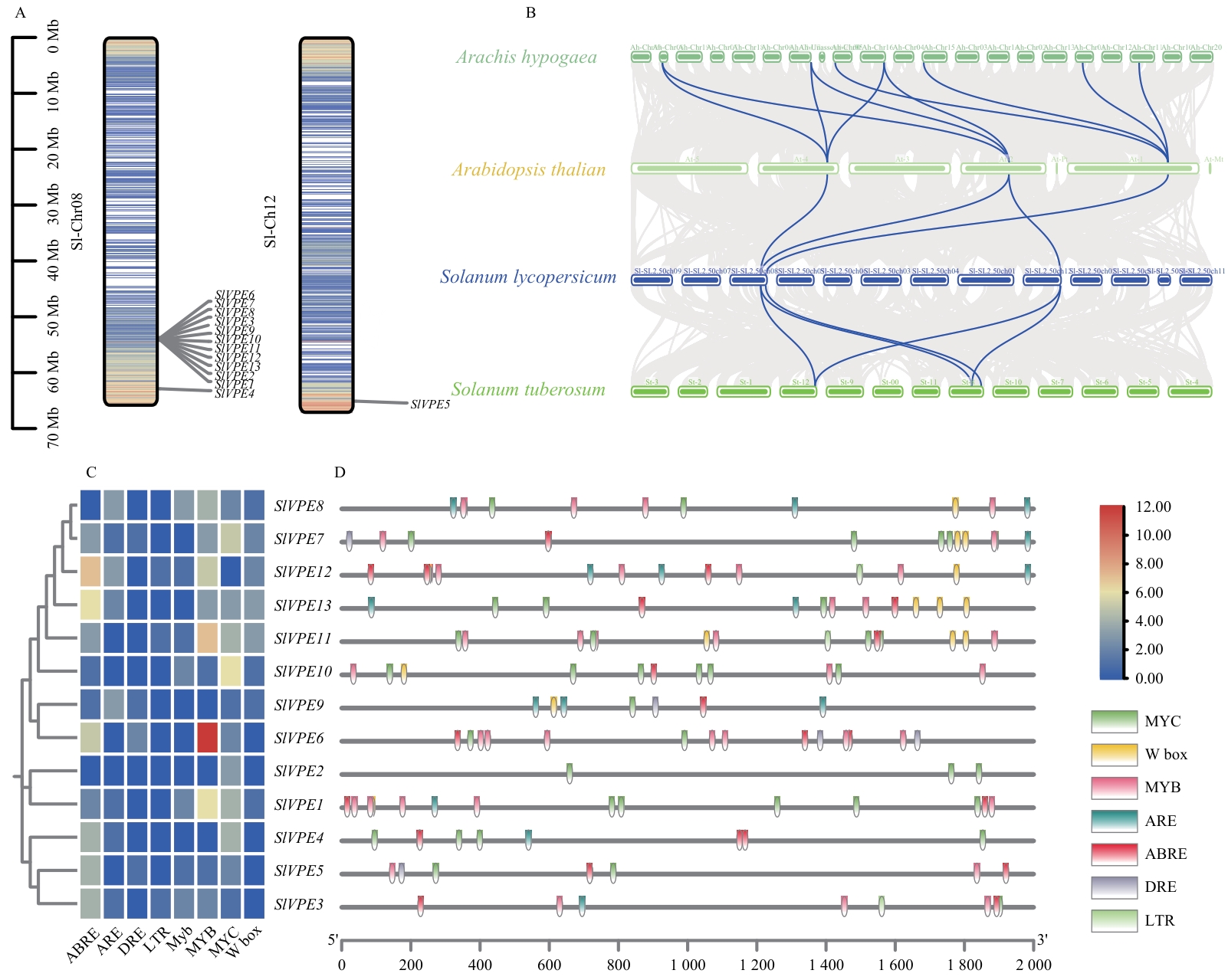
Fig. 3 Analysis of chromosomal localization, interspecies collinearity, and cis-acting elements of SlVPE genesA: Chromosomal locations of the SlVPE genes. B: Collinear relationships of SlVPE genes among tomato, Arabidopsis, potato, and peanut genomes. Gray and blue lines indicate the collinearity of all genes and VPE genes, respectively. C: Heatmap of cis-acting elements in SlVPEs promoters. D: Visualization of the types, positions, and numbers of cis-acting elements in SlVPEs promoters
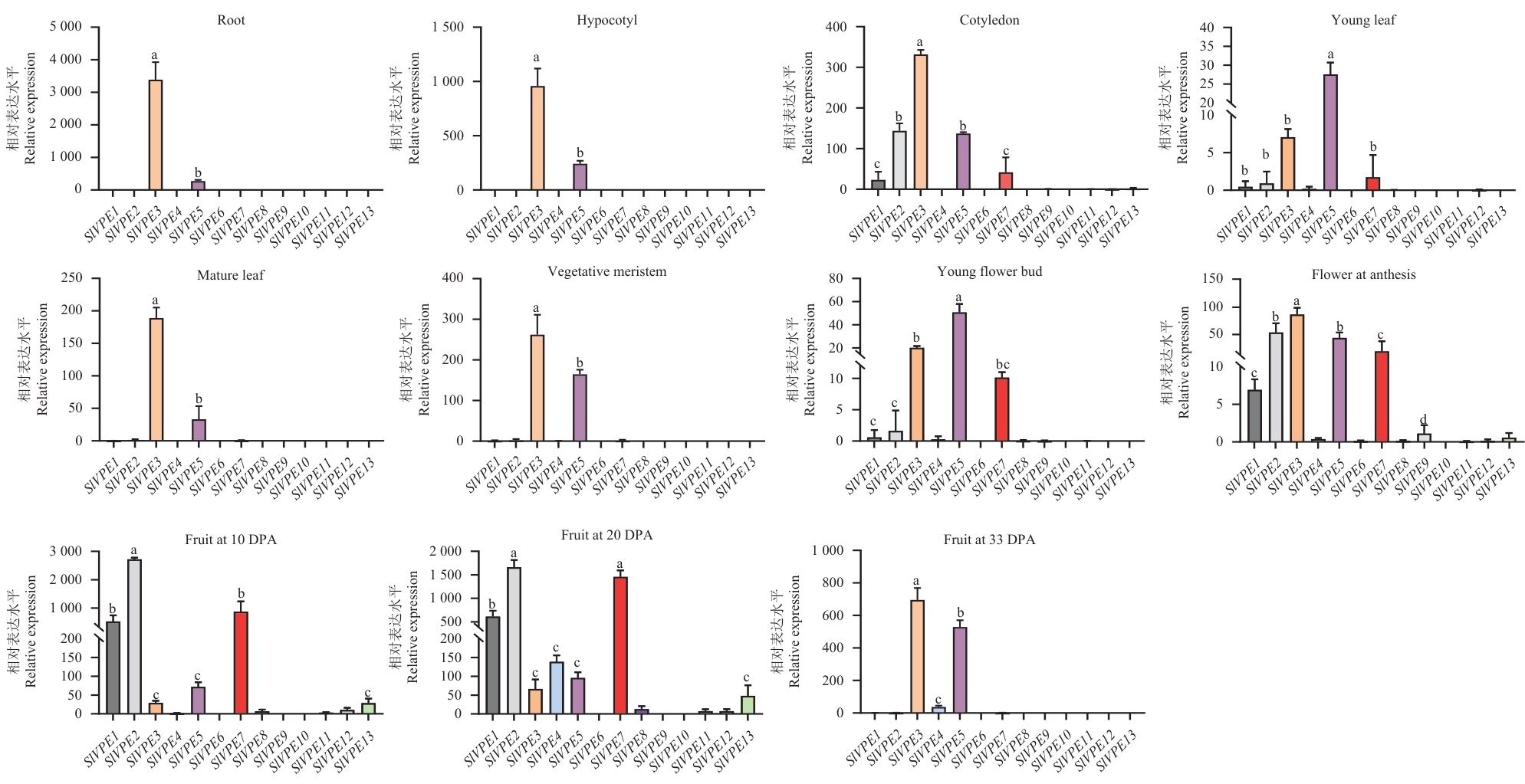
Fig. 4 Expression profile analysis of SlVPE gene in different tissues of tomatoData are presented as mean ± SE from four biological replicates (n=4). Different lowercase letters indicate significant differences (P<0.05)
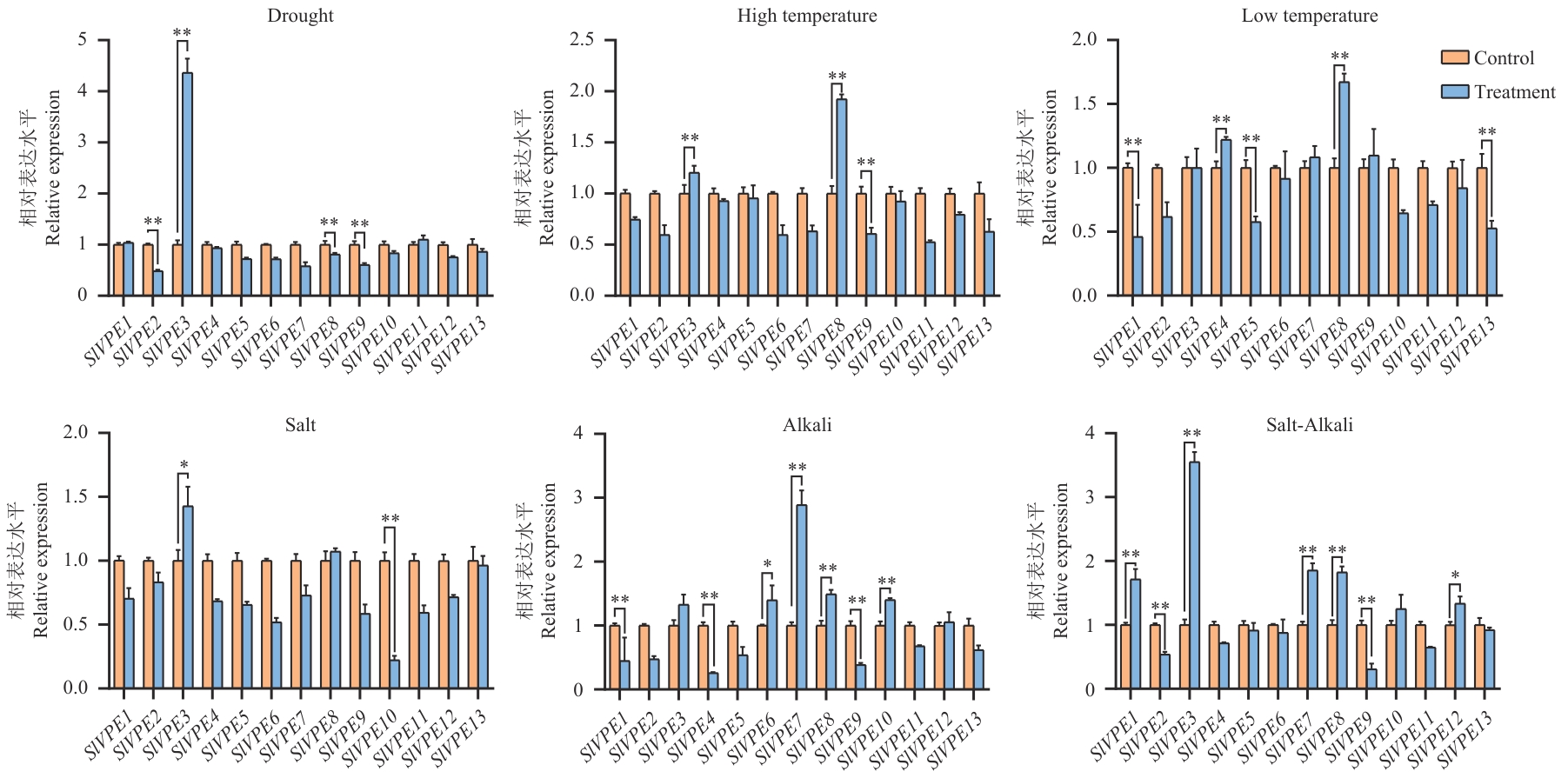
Fig. 6 Expression analysis of SlVPE genes under various abiotic stressesDrought stress: 20% PEG6000; salt stress: 150 mmol/L NaCl; alkali stress: 100 mmol/L NaHCO₃; salt-alkali combined stress: 60 mmol/L NaCl+60 mmol/L NaHCO3; low-temperature stress: 12 ℃; high-temperature stress: 38 ℃. Samples were collected at 7 d after stress treatment. Data are presented as mean±SE of three biological replicates (n=3), each with three technical replicates. *P<0.05, **P<0.01
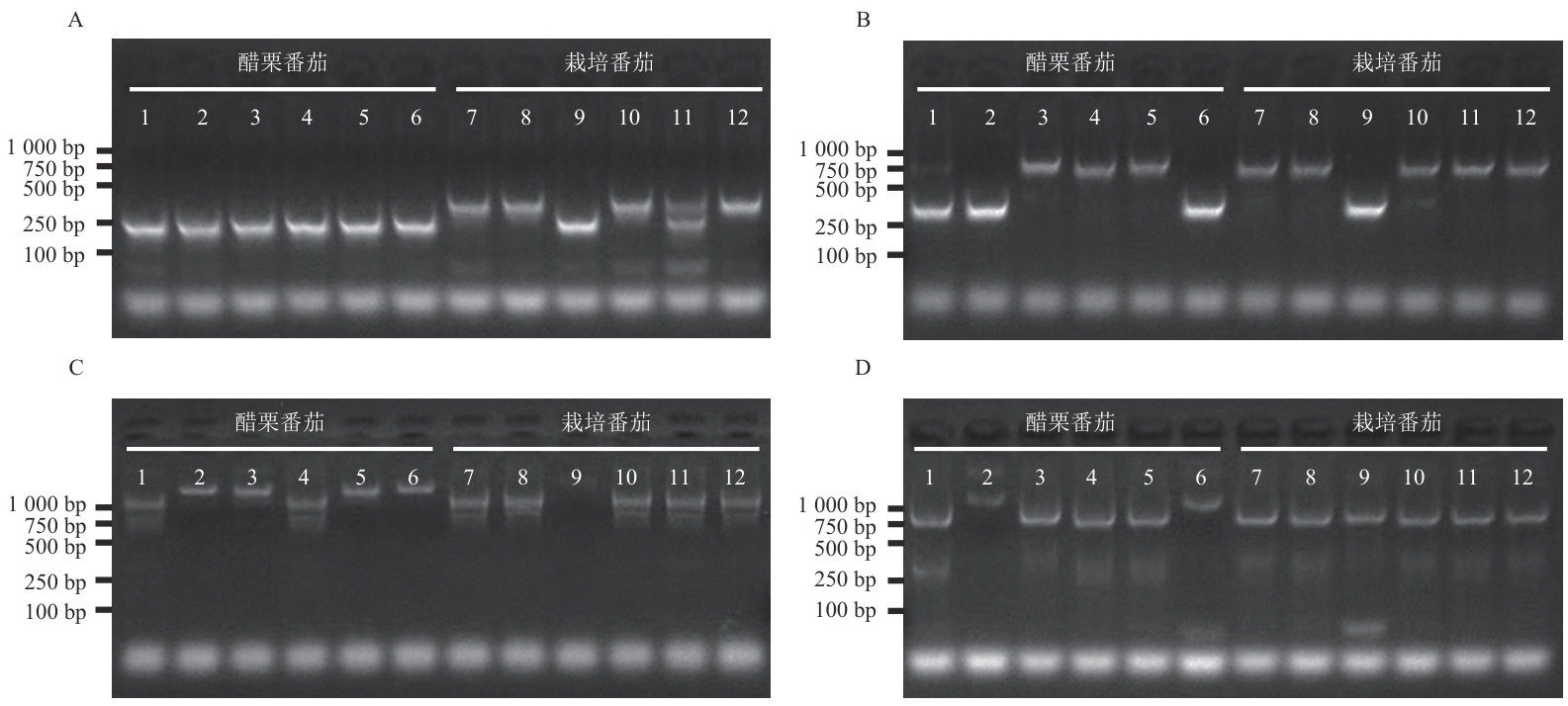
Fig. 7 Polymorphism detection of SlVPE gene molecular markersAgarose gel electrophoresis was used to examine polymorphisms between Solanum pimpinellifolium and cultivated tomato in the promoter region of SlVPE6 (A), the downstream region of SlVPE11 (B), the promoter region of SlVPE13 (C), and the promoter region of SlVPE10 (D). Lane 1-6: S. pimpinellifolium accessions, LA0533, LA1375, LA1420, LA2873, LA4763, and LA1589; lane 7-12: cultivated tomato materials, AC, M82, TS-418, Hawaii7996, MoneyMaker, and Heinz1706
| [1] | Hara-Nishimura I, Hatsugai N. The role of vacuole in plant cell death [J]. Cell Death Differ, 2011, 18(8): 1298-1304. |
| [2] | Teper-Bamnolker P, Danieli R, Peled-Zehavi H, et al. Vacuolar processing enzyme translocates to the vacuole through the autophagy pathway to induce programmed cell death [J]. Autophagy, 2021, 17(10): 3109-3123. |
| [3] | Sarwar R, Yu J, Zekraoui M, et al. Unraveling the molecular functions of multifaced plant-vacuolar processing enzymes [J]. Physiol Plant, 2024, 176: e14131. |
| [4] | Gao XX, Tang YL, Shi QY, et al. Vacuolar processing enzyme positively modulates plant resistance and cell death in response to Phytophthora parasitica infection [J]. J Integr Agric, 2023, 22(5): 1424-1433. |
| [5] | Duan XB, Zhang K, Yu Y, et al. Genome-wide analysis of soybean vacuolar processing enzyme gene family reveals their roles in plant development and response to stress conditions [J]. J Plant Growth Regul, 2024, 43(8): 2817-2829. |
| [6] | Shimada T, Yamada K, Kataoka M, et al. Vacuolar processing enzymes are essential for proper processing of seed storage proteins in Arabidopsis thaliana [J]. J Biol Chem, 2003, 278(34): 32292-32299. |
| [7] | Tang CN, Wan Abdullah WMAN, Wee CY, et al. Promoter cis-element analyses reveal the function of αVPE in drought stress response of Arabidopsis [J]. Biology, 2023, 12(3): 430. |
| [8] | Nakaune S, Yamada K, Kondo M, et al. A vacuolar processing enzyme, δVPE, is involved in seed coat formation at the early stage of seed development [J]. Plant Cell, 2005, 17(3): 876-887. |
| [9] | Kuroyanagi M, Yamada K, Hatsugai N, et al. Vacuolar processing enzyme is essential for mycotoxin-induced cell death in Arabidopsis thaliana [J]. J Biol Chem, 2005, 280(38): 32914-32920. |
| [10] | Lu WY, Deng MJ, Guo F, et al. Suppression of OsVPE3 enhances salt tolerance by attenuating vacuole rupture during programmed cell death and affects stomata development in rice [J]. Rice, 2016, 9: 65. |
| [11] | Deng HB, Cao S, Zhang GL, et al. OsVPE2, a member of vacuolar processing enzyme family, decreases chilling tolerance of rice [J]. Rice, 2024, 17: 5. |
| [12] | Zhang JH, Yue Y, Hu MJ, et al. Dynamic transcriptome landscape of maize pericarp development [J]. Plant J, 2024, 117(5): 1574-1591. |
| [13] | Wang TH, Guan MH, Zheng YH, et al. Genome-wide molecular characterization and expression profiling of the cysteine protease gene family in maize [J]. BMC Genomics, 2025, 26(1): 789. |
| [14] | Quadros IPS, Madeira NN, Loriato VAP, et al. Cadmium-mediated toxicity in plant cells is associated with the DCD/NRP-mediated cell death response [J]. Plant Cell Environ, 2022, 45(2): 556-571. |
| [15] | Radchuk V, Tran V, Radchuk R, et al. Vacuolar processing enzyme 4 contributes to maternal control of grain size in barley by executing programmed cell death in the pericarp [J]. New Phytol, 2018, 218(3): 1127-1142. |
| [16] | Wang BK, Li N, Huang SY, et al. Enhanced soluble sugar content in tomato fruit using CRISPR/Cas9-mediated SlINVINH1 and SlVPE5 gene editing [J]. PeerJ, 2021, 9: e12478. |
| [17] | Wang WH, Cai JH, Wang PW, et al. Post-transcriptional regulation of fruit ripening and disease resistance in tomato by the vacuolar protease SlVPE3 [J]. Genome Biol, 2017, 18: 47. |
| [18] | Zhu LF, Wang XP, Tian J, et al. Genome-wide analysis of VPE family in four Gossypium species and transcriptional expression of VPEs in the upland cotton seedlings under abiotic stresses [J]. Funct Integr Genomics, 2022, 22(2): 179-192. |
| [19] | Song JF, Yang F, Xun M, et al. Genome-wide identification and characterization of vacuolar processing enzyme gene family and diverse expression under stress in apple (Malus × domestic) [J]. Front Plant Sci, 2020, 11: 626. |
| [20] | Bombarely A, Menda N, Tecle IY, et al. The Sol Genomics Network (solgenomics.net): growing tomatoes using Perl [J]. Nucleic Acids Res, 2011, 39(): D1149-D1155. |
| [21] | Consortium The Tomato Genome. The tomato genome sequence provides insights into fleshy fruit evolution [J]. Nature, 2012, 485(7400): 635-641. |
| [22] | Finn RD, Bateman A, Clements J, et al. Pfam: the protein families database [J]. Nucleic Acids Res, 2014, 42(D1): D222-D230. |
| [23] | Berardini TZ, Reiser L, Li DH, et al. The Arabidopsis information resource: Making and mining the “gold standard” annotated reference plant genome [J]. Genesis, 2015, 53(8): 474-485. |
| [24] | Bertioli DJ, Cannon SB, Froenicke L, et al. The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut [J]. Nat Genet, 2016, 48(4): 438-446. |
| [25] | Kumar S, Stecher G, Li M, et al. MEGA X: molecular evolutionary genetics analysis across computing platforms [J]. Mol Biol Evol, 2018, 35(6): 1547-1549. |
| [26] | Chen CJ, Chen H, Zhang Y, et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data [J]. Mol Plant, 2020, 13(8): 1194-1202. |
| [27] | Bailey TL, Boden M, Buske FA, et al. MEME Suite: tools for motif discovery and searching [J]. Nucleic Acids Res, 2009, 37(): W202-W208. |
| [28] | Lescot M, Déhais P, Thijs G, et al. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences [J]. Nucleic Acids Res, 2002, 30(1): 325-327. |
| [29] | Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method [J]. Methods, 2001, 25(4): 402-408. |
| [30] | Wleklik K, Borek S. Vacuolar processing enzymes in plant programmed cell death and autophagy [J]. Int J Mol Sci, 2023, 24(2): 1198. |
| [31] | Pat Heslop-Harrison JS, Schwarzacher T, Liu Q. Polyploidy: its consequences and enabling role in plant diversification and evolution [J]. Ann Bot, 2023, 131(1): 1-10. |
| [32] | Fang YP, Jiang JM, Hou XL, et al. Plant protein-coding gene families: Their origin and evolution [J]. Front Plant Sci, 2022, 13: 995746. |
| [33] | Zheng P, Zheng CY, Otegui MS, et al. Endomembrane mediated-trafficking of seed storage proteins: from Arabidopsis to cereal crops [J]. J Exp Bot, 2022, 73(5): 1312-1326. |
| [34] | Baral R, Vainer A, Melzer S, et al. ‘Bud to fruit’—hormonal interactions governing early fruit development [J]. J Exp Bot, 2025, 76(22): 6657-6673. |
| [35] | Wang RF, de Maagd RA. Transcriptional control of tomato fruit development and ripening [J]. J Exp Bot, 2025, 76(21): 6311-6326. |
| [36] | Wei SB, Li X, Lu ZF, et al. A transcriptional regulator that boosts grain yields and shortens the growth duration of rice [J]. Science, 2022, 377(6604): eabi8455. |
| [37] | Tan WR, Chen JH, Yue XL, et al. The heat response regulators HSFA1s promote Arabidopsis thermomorphogenesis via stabilizing PIF4 during the day [J]. Sci Adv, 2023, 9(44): eadh1738. |
| [38] | Xu L, Yang LJ, Li AP, et al. An AP2/ERF transcription factor confers chilling tolerance in rice [J]. Sci Adv, 2024, 10(35): eado4788. |
| [39] | Zhu JK. Abiotic stress signaling and responses in plants [J]. Cell, 2016, 167(2): 313-324. |
| [1] | YIN Ya-long, ZHANG Ming-yang, WANG Jie-min, MIAO Xue-xue, CHEN Jin, WANG Wei-ping. Advances in Coordinated Tolerance Mechanisms to Abiotic Stresses in Rice [J]. Biotechnology Bulletin, 2026, 42(4): 26-37. |
| [2] | CHEN Deng-ke, LAN Gang, XIA Zhi, HOU Bao-guo, YANG Liu-liu, CAO Cai-rong, LI Peng-bo, WU Cui-cui. Identification of ZF-HD Gene Family in Arachis hypogaea and Analysis in Response to Abiotic Stress [J]. Biotechnology Bulletin, 2026, 42(4): 114-128. |
| [3] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [4] | DU Dan, GUO Xiang, HU Xin, PAN Yu. Advances in the Regulatory Mechanisms of Plastid Development on Fruit Ripening and Quality [J]. Biotechnology Bulletin, 2026, 42(3): 48-59. |
| [5] | JIANG Zhe-hui, WANG Xiao-long, WANG Shou-chuang, ZHOU Ke. Advances in the Elucidation of Metabolic Pathways and Molecular Breeding for Tomato Flavor [J]. Biotechnology Bulletin, 2026, 42(3): 60-78. |
| [6] | WANG Xiao-yi, LI Jin-yan, XING Xing, ZHU Hong-liang. Screening and Functional Analysis of Ethylene-responsive Genes Regulating Tomato Fruit Ripening and Respiration [J]. Biotechnology Bulletin, 2026, 42(3): 275-282. |
| [7] | LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development [J]. Biotechnology Bulletin, 2026, 42(3): 362-373. |
| [8] | LIU Miao, LIN Tao, JIA Le-song, HU Feng, LI Tao, LI Zhi-wan, LIU Mei-fang, ZHENG Fang-yan, CUI Long. From Wild to Cultivated: Evolution and Regulatory Mechanisms of Tomato Fruit Color [J]. Biotechnology Bulletin, 2026, 42(3): 187-202. |
| [9] | LI Ya-ni, HAN Hong-yu, GENG Meng-shuang, MI Ruo-lan, Wang Wei-qi, Yu Wen-jing, MENG Xian-wen, LI Chuan-you. Mechanistic Study on ChiC-mediated Regulation Mechanism of Tomato Resistance to Botrytis cinerea [J]. Biotechnology Bulletin, 2026, 42(3): 255-262. |
| [10] | CHENG Yun-xia, ZHANG Jun-hong, YE Jie. Advances in the Genetic Regulation of Soluble Solid Accumulation in Tomato Fruits [J]. Biotechnology Bulletin, 2026, 42(3): 145-155. |
| [11] | YAN Chen-lin, LI Fan, YAN Chun-ting, CHENG Jiao-wen, HU Kai-lin, YE Zhi-biao, SONG Jian-wen. Advances in Genes Related to Tomato Fruit Morphogenesis [J]. Biotechnology Bulletin, 2026, 42(3): 172-186. |
| [12] | LONG Lin-xi, ZENG Yin-ping, WANG Qian, DENG Yu-ping, GE Min-qian, CHEN Yan-zhuo, LI Xin-juan, YANG Jun, ZOU Jian. Identification of Sunflower GH3 Gene Family and Analysis of Their Function in Flower Development [J]. Biotechnology Bulletin, 2026, 42(1): 125-138. |
| [13] | FEI Si-tian, HOU Ying-xiang, LI Lan, ZHANG Chao. Biological Functions and Regulatory Network of SLR1, a Negative Regulator of Gibberellin Signaling in Rice [J]. Biotechnology Bulletin, 2026, 42(1): 13-30. |
| [14] | LI Zheng, QIU Wei-yue, SUN Rui-xue, ZHAO Xiao. Overexpression of the xylR Gene Driven by Different Promoters Enhances Glucose-xylose Co-utilization Capability in Escherichia coli [J]. Biotechnology Bulletin, 2026, 42(1): 329-337. |
| [15] | ZHANG Chi-hao, LIU Jin-nan, CHAO Yue-hui. Cloning and Functional Analysis of a bZIP Transcription Factor MtbZIP29 from Medicago truncatula [J]. Biotechnology Bulletin, 2026, 42(1): 241-250. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||