








Biotechnology Bulletin ›› 2021, Vol. 37 ›› Issue (6): 154-162.doi: 10.13560/j.cnki.biotech.bull.1985.2020-1165
Previous Articles Next Articles
ZHANG Ting-huan( ), ZHANG Li-juan, CHEN Si-qing, GUO Zong-yi(
), ZHANG Li-juan, CHEN Si-qing, GUO Zong-yi( )
)
Received:2020-09-15
Online:2021-06-26
Published:2021-07-08
Contact:
GUO Zong-yi
E-mail:ztinghuan@163.com;guozongyi@sina.com
ZHANG Ting-huan, ZHANG Li-juan, CHEN Si-qing, GUO Zong-yi. Effects of the Polymorphism of the Seed Sequence in Porcine miR-378 on Its Function and Carcass Traits[J]. Biotechnology Bulletin, 2021, 37(6): 154-162.
| 引物名称 Name | 引物序列 Sequence | 长度Length/bp |
|---|---|---|
| ap2-F | GGCGTGACTTCCACAAGAGTTTA | 23 |
| ap2-R | GCCTCTTCCTTTGGCTCATG | 20 |
| Ppar-γ-F | GTGAAGGATGCAAGGGTT | 18 |
| Ppar-γ-R | CCTGATGGCATTGTGAGA | 18 |
| C/EBP-α-R | TGGACAAGAACAGCAACGAG | 20 |
| C/EBP-α-F | TCACTGGTCAACTCCAGCAC | 20 |
| PGC-1α-F | AGCGCCGTGTGATTTACGTT | 20 |
| PGC-1α-R | CCGCAGATTTACGGTGCATT | 20 |
| PGC-1β-F | CAGACGTGAGAGCAGAGGGC | 20 |
| PGC-1β-R | CGAATGTATACCACACGGCCT | 21 |
| Resistin-F | CAACTCCCTGTTTCCAAATGC | 21 |
| Resistin-R | GTCCAGCAATTTAAGCCAATGTT | 23 |
| HSL-F | AACCAACCCTAGGCCAACTG | 20 |
| HSL-R | GCTGTGTGCACCAAACTACG | 20 |
| ATGL-F | AGGCCAATGTCTGCAGCACAT | 21 |
| ATGL-R | CAAGTTGTCTGAAATGCCGCC | 21 |
| CGI-58-F | CCCTCAGGTTGGACAAAATGA | 21 |
| CGI-58-R | AGGAAAACCCCATGGCTCTAC | 21 |
| GAPDH-F | ACCACAGTCCATGCCATCAC | 20 |
| GAPDH-R | TCCACCACCCTGTTGCTGTA | 20 |
| U6-F | CGCTTCGGCAGCACATATA | 19 |
| U6-R | TTCACGAATTTGCGTGTCAT | 20 |
| miR-378W | ACTGGACTTGGAGTCAGAAGGC | 22 |
| miR-378M | ACTGGGCTTGGAGTCAGAAGGC | 22 |
Table 1 RT-PCR primers
| 引物名称 Name | 引物序列 Sequence | 长度Length/bp |
|---|---|---|
| ap2-F | GGCGTGACTTCCACAAGAGTTTA | 23 |
| ap2-R | GCCTCTTCCTTTGGCTCATG | 20 |
| Ppar-γ-F | GTGAAGGATGCAAGGGTT | 18 |
| Ppar-γ-R | CCTGATGGCATTGTGAGA | 18 |
| C/EBP-α-R | TGGACAAGAACAGCAACGAG | 20 |
| C/EBP-α-F | TCACTGGTCAACTCCAGCAC | 20 |
| PGC-1α-F | AGCGCCGTGTGATTTACGTT | 20 |
| PGC-1α-R | CCGCAGATTTACGGTGCATT | 20 |
| PGC-1β-F | CAGACGTGAGAGCAGAGGGC | 20 |
| PGC-1β-R | CGAATGTATACCACACGGCCT | 21 |
| Resistin-F | CAACTCCCTGTTTCCAAATGC | 21 |
| Resistin-R | GTCCAGCAATTTAAGCCAATGTT | 23 |
| HSL-F | AACCAACCCTAGGCCAACTG | 20 |
| HSL-R | GCTGTGTGCACCAAACTACG | 20 |
| ATGL-F | AGGCCAATGTCTGCAGCACAT | 21 |
| ATGL-R | CAAGTTGTCTGAAATGCCGCC | 21 |
| CGI-58-F | CCCTCAGGTTGGACAAAATGA | 21 |
| CGI-58-R | AGGAAAACCCCATGGCTCTAC | 21 |
| GAPDH-F | ACCACAGTCCATGCCATCAC | 20 |
| GAPDH-R | TCCACCACCCTGTTGCTGTA | 20 |
| U6-F | CGCTTCGGCAGCACATATA | 19 |
| U6-R | TTCACGAATTTGCGTGTCAT | 20 |
| miR-378W | ACTGGACTTGGAGTCAGAAGGC | 22 |
| miR-378M | ACTGGGCTTGGAGTCAGAAGGC | 22 |
| 引物名称 Name | 引物序列 Sequence | 长度 Length/bp |
|---|---|---|
| miR-378-F | TAACCCCTAGGTGGGTCTGAG | 21 |
| miR-378-R | TCAGATGAGCAGGACAGTTCAG | 22 |
Table 2 Amplification primer of miR-378
| 引物名称 Name | 引物序列 Sequence | 长度 Length/bp |
|---|---|---|
| miR-378-F | TAACCCCTAGGTGGGTCTGAG | 21 |
| miR-378-R | TCAGATGAGCAGGACAGTTCAG | 22 |
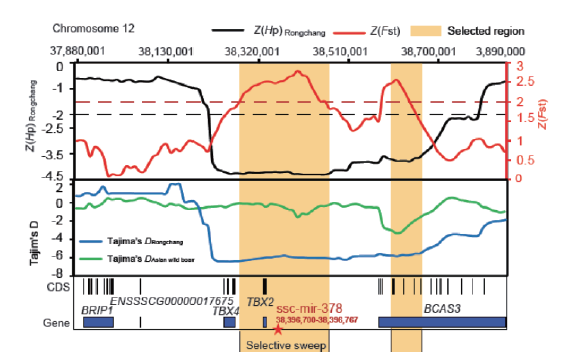
Fig.1 Population genetic selection analysis of miR-378 locus in Rongchang pig and Asian wild boar The areas marked in yellow indicate the selected regions and the red five-pointed star indicates miR-378 locus
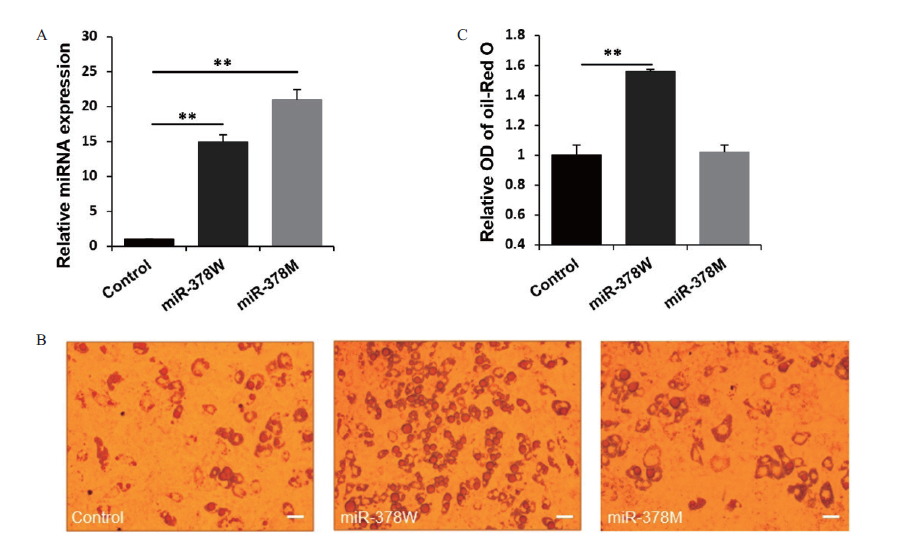
Fig.4 Effects of miR-378W and miR-378 M on the lipid production of adipocytes A:Transfection efficiency of miRNA mimics. B:Oil red O staining adipocytes,scale bar:50 μm. C:Quantitative analysis of the lipid content. **refers to significant difference(P < 0.01),the same below
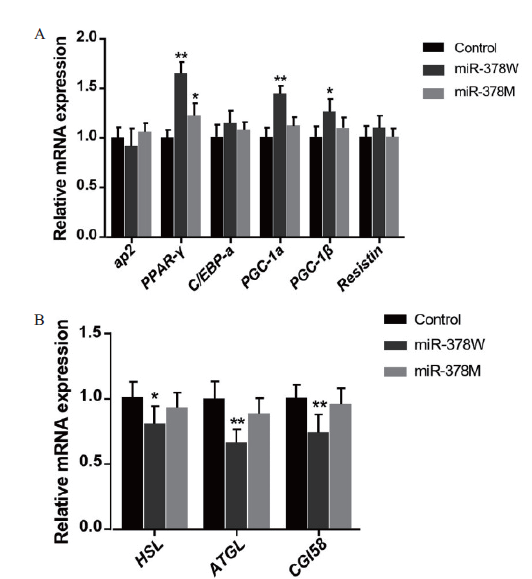
Fig.5 Effects of miR-378W and miR-378 M on the expres-sion of lipid related genes A:The expression of genes related to adipogenesis. B:The expression of genes related to lipolysis. * refers to significant difference(P < 0.05),the same below
| GG(24) | AG(51) | AA(32) | 加性效应Additive effect | 显性效应Dominant effect | |
|---|---|---|---|---|---|
| 最后肋骨处背膘厚 Backfat thickness at the last rib/cm | 2.77a±0.20 | 2.66b±0.13 | 2.63b±0.16 | 0.07 | -0.04 |
| 眼肌面积 Eye-muscle area/cm2 | 22.16b±1.08 | 23.25ab±0.63 | 23.80a±0.62 | 1.22 | 0.445 |
| 胴体瘦肉率 Lean percentage/% | 48.12±1.06 | 48.28±0.64 | 48.32±0.55 | 0.32 | 0.07 |
Table 3 Number of individuals for three genotypes of miR-378 seed sequence in the population and their association with slaughter traits
| GG(24) | AG(51) | AA(32) | 加性效应Additive effect | 显性效应Dominant effect | |
|---|---|---|---|---|---|
| 最后肋骨处背膘厚 Backfat thickness at the last rib/cm | 2.77a±0.20 | 2.66b±0.13 | 2.63b±0.16 | 0.07 | -0.04 |
| 眼肌面积 Eye-muscle area/cm2 | 22.16b±1.08 | 23.25ab±0.63 | 23.80a±0.62 | 1.22 | 0.445 |
| 胴体瘦肉率 Lean percentage/% | 48.12±1.06 | 48.28±0.64 | 48.32±0.55 | 0.32 | 0.07 |
| [1] | Yapijakis C. Regulatory role of microRNAs in brain development and function[J]. Adv Exp Med Biol, 2020, 1195:237-247. |
| [2] | Ameres SL, Zamore PD. Diversifying microRNA sequence and function[J]. Nat Rev Mol Cell Biol, 2013, 14(8):475-488. |
| [3] |
Michlewski G, Caceres JF. Post-transcriptional control of miRNA biogenesis[J]. RNA, 2019, 25(1):1-16.
doi: 10.1261/rna.068692.118 pmid: 30333195 |
| [4] | Hausser J, Zavolan M. Identification and consequences of miRNA-target interactions--beyond repression of gene expression[J]. Nat Rev Genet, 2014, 15(9):599-612. |
| [5] |
Dorn GW 2nd, Matkovich SJ, Eschenbacher WH, et al. A human 3’ miR-499 mutation alters cardiac mRNA targeting and function[J]. Circ Res, 2012, 110(7):958-967.
doi: 10.1161/CIRCRESAHA.111.260752 URL |
| [6] |
Kotani A, Ha D, Schotte D, et al. A novel mutation in the miR-128b gene reduces miRNA processing and leads to glucocorticoid resistance of MLL-AF4 acute lymphocytic leukemia cells[J]. Cell Cycle, 2010, 9(6):1037-1042.
pmid: 20237425 |
| [7] |
Sun W, Lan J, Chen L, et al. A mutation in porcine pre-miR-15b alters the biogenesis of MiR-15b\16-1 cluster and strand selection of MiR-15b[J]. PLoS One, 2017, 12(5):e0178045.
doi: 10.1371/journal.pone.0178045 URL |
| [8] |
Mencia A, Modamio-Hoybjor S, Redshaw N, et al. Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss[J]. Nat Genet, 2009, 41(5):609-613.
doi: 10.1038/ng.355 URL |
| [9] |
Duan R, Pak C, Jin P. Single nucleotide polymorphism associated with mature miR-125a alters the processing of pri-miRNA[J]. Hum Mol Genet, 2007, 16(9):1124-1131.
doi: 10.1093/hmg/ddm062 URL |
| [10] |
Carrer M, Liu N, Grueter CE, et al. Control of mitochondrial metabolism and systemic energy homeostasis by microRNAs 378 and 378*[J]. Proc Natl Acad Sci USA, 2012, 109(38):15330-15335.
doi: 10.1073/pnas.1207605109 URL |
| [11] |
Gerin I, Bommer GT, Mccoin CS, et al. Roles for miRNA-378/378* in adipocyte gene expression and lipogenesis[J]. Am J Physiol Endocrinol Metab, 2010, 299(2):E198-206.
doi: 10.1152/ajpendo.00179.2010 URL |
| [12] |
Gagan J, Dey BK, Layer R, et al. MicroRNA-378 targets the myogenic repressor MyoR during myoblast differentiation[J]. J Biol Chem, 2011, 286(22):19431-19438.
doi: 10.1074/jbc.M111.219006 URL |
| [13] |
Tong H, Jiang R, Liu T, et al. bta-miR-378 promote the differentiation of bovine skeletal muscle-derived satellite cells[J]. Gene, 2018, 668:246-251.
doi: 10.1016/j.gene.2018.03.102 URL |
| [14] |
Hupkes M, Sotoca AM, Hendriks JM, et al. MicroRNA miR-378 promotes BMP2-induced osteogenic differentiation of mesenchymal progenitor cells[J]. BMC Mol Biol, 2014, 15:1.
doi: 10.1186/1471-2199-15-1 URL |
| [15] |
Chai J, Chen L, Luo Z, et al. Spontaneous single nucleotide polymorphism in porcine microRNA-378 seed region leads to functional alteration[J]. Bioscience, Biotechnology, and Biochemistry, 2018, 82(7):1081-1089.
doi: 10.1080/09168451.2018.1459175 URL |
| [16] |
Xu LL, Shi CM, Xu GF, et al. TNF-alpha, IL-6, and leptin increase the expression of miR-378, an adipogenesis-related microRNA in human adipocytes[J]. Cell Biochem Biophys, 2014, 70(2):771-776.
doi: 10.1007/s12013-014-9980-x URL |
| [17] |
Pan D, Mao C, Quattrochi B, et al. MicroRNA-378 controls classical brown fat expansion to counteract obesity[J]. Nat Commun, 2014, 5:4725.
doi: 10.1038/ncomms5725 URL |
| [18] |
Lu J, Webb R, Richardson JA, et al. MyoR:a muscle-restricted basic helix-loop-helix transcription factor that antagonizes the actions of MyoD[J]. Proc Natl Acad Sci USA, 1999, 96(2):552-557.
doi: 10.1073/pnas.96.2.552 URL |
| [19] |
Hou X, Tang Z, Liu H, et al. Discovery of MicroRNAs associated with myogenesis by deep sequencing of serial developmental skeletal muscles in pigs[J]. PLoS One, 2012, 7(12):e52123.
doi: 10.1371/journal.pone.0052123 URL |
| [20] |
Knezevic I, Patel A, Sundaresan NR, et al. A novel cardiomyocyte-enriched microRNA, miR-378, targets insulin-like growth factor 1 receptor:implications in postnatal cardiac remodeling and cell survival[J]. J Biol Chem, 2012, 287(16):12913-12926.
doi: 10.1074/jbc.M111.331751 pmid: 22367207 |
| [21] |
Chai J, Chen L, Luo Z, et al. Spontaneous single nucleotide polymorphism in porcine microRNA-378 seed region leads to functional alteration[J]. Biosci Biotechnol Biochem, 2018, 82(7):1081-1089.
doi: 10.1080/09168451.2018.1459175 URL |
| [22] | Wang K, Cao Y, Rong Y, et al. A novel SNP in EIF2AK4 gene is associated with thermal tolerance traits in Chinesecattle[J]. Animals(Basel), 2019, 9(6):375. |
| [23] |
Chang JS, Ghosh S, Newman S, et al. A map of the PGC-1alpha- and NT-PGC-1alpha-regulated transcriptional network in brown adipose tissue[J]. Sci Rep, 2018, 8(1):7876.
doi: 10.1038/s41598-018-26244-4 pmid: 29777200 |
| [24] |
Song W, Zhong C, Yuan Y, et al. Peroxisome proliferator-activated receptor-coactivator 1-beta(PGC-1beta)modulates the expression of genes involved in adipogenesis during preadipocyte differentiation in chicken[J]. Gene, 2020, 741:144516.
doi: 10.1016/j.gene.2020.144516 URL |
| [25] |
Xu L, Ma X, Verma NK, et al. Ablation of PPARgamma in subcutaneous fat exacerbates age-associated obesity and metabolic decline[J]. Aging Cell, 2018, 17(2):e12721.
doi: 10.1111/acel.2018.17.issue-2 URL |
| [26] | Schreiber R, Xie H, Schweiger M. Of mice and men:The physiological role of adipose triglyceride lipase(ATGL)[J]. Biochim Biophys Acta Mol Cell Biol Lipids, 2019, 1864(6):880-899. |
| [27] |
Demine S, Tejerina S, Bihin B, et al. Mild mitochondrial uncoupling induces HSL/ATGL-independent lipolysis relying on a form of autophagy in 3T3-L1 adipocytes[J]. J Cell Physiol, 2018, 233(2):1247-1265.
doi: 10.1002/jcp.25994 URL |
| [28] |
Korbelius M, Vujic N, Sachdev V, et al. ATGL/CGI-58-dependent hydrolysis of a lipid storage pool in murine enterocytes[J]. Cell Rep, 2019, 28(7):1923-1934.e4.
doi: S2211-1247(19)30927-1 pmid: 31412256 |
| [29] |
Choy YH, Park BH, Choi TJ, et al. Estimation of relative economic weights of hanwoo carcass traits based on carcass market price[J]. Asian-Australas J Anim Sci, 2012, 25(12):1667-1673.
doi: 10.5713/ajas.2012.12397 URL |
| [30] |
Wei X, Shimizu T, Lai ZC. Mob as tumor suppressor is activated by Hippo kinase for growth inhibition in Drosophila[J]. EMBO J, 2007, 26(7):1772-1781.
doi: 10.1038/sj.emboj.7601630 URL |
| [31] |
Wang Z, Ma B, Ji X, et al. MicroRNA-378-5p suppresses cell proliferation and induces apoptosis in colorectal cancer cells by targeting BRAF[J]. Cancer Cell Int, 2015, 15:40.
doi: 10.1186/s12935-015-0192-2 URL |
| [32] |
Zeng M, Zhu L, Li L, et al. miR-378 suppresses the proliferation, migration and invasion of colon cancer cells by inhibiting SDAD1[J]. Cell Mol Biol Lett, 2017, 22:12.
doi: 10.1186/s11658-017-0041-5 URL |
| [33] |
Peng N, Miao Z, Wang L, et al. MiR-378 promotes the cell proliferation of osteosarcoma through down-regulating the expression of Kruppel-like factor 9[J]. Biochem Cell Biol, 2018, 96(5):515-521.
doi: 10.1139/bcb-2017-0186 URL |
| [34] |
Kuang X, Wei C, Zhang T, et al. miR-378 inhibits cell growth and enhances apoptosis in human myelodysplastic syndromes[J]. Int J Oncol, 2016, 49(5):1921-1930.
doi: 10.3892/ijo.2016.3689 URL |
| [35] |
Kulyte A, Lorente-Cebrian S, Gao H, et al. MicroRNA profiling links miR-378 to enhanced adipocyte lipolysis in human cancer cachexia[J]. Am J Physiol Endocrinol Metab, 2014, 306(3):E267-274.
doi: 10.1152/ajpendo.00249.2013 URL |
| [36] |
Li Y, Jiang J, Liu W, et al. microRNA-378 promotes autophagy and inhibits apoptosis in skeletal muscle[J]. Proc Natl Acad Sci USA, 2018, 115(46):E10849-E10858.
doi: 10.1073/pnas.1803377115 URL |
| [1] | YU Shi-zhou, CAO Ling-gai, WANG Shi-ze, LIU Yong, BIAN Wen-jie, REN Xue-liang. Development Core SNP Markers for Tobacco Germplasm Genotyping [J]. Biotechnology Bulletin, 2023, 39(3): 89-100. |
| [2] | ZHANG Ting-huan, GUO Zong-yi, CHAI Jie, PAN Hong-mei, ZHANG Liang, CHEN Lei, LONG Xi. Effects of Sequence Variation on the Biogenesis and Target Relationship of miR-378 [J]. Biotechnology Bulletin, 2022, 38(1): 205-214. |
| [3] | ZHANG Ting-huan, LONG Xi, GUO Zong-yi, CHAI Jie. miR-378 Promoting Lipogenesis and Identification of Target Genes [J]. Biotechnology Bulletin, 2021, 37(2): 80-87. |
| [4] | GUO Li-li, LI Yu-ying, GUO Da-long, HOU Xiao-gai. Research Progress on High-density Genetic Linkage Map Construction of Important Ornamental Plants:a Review [J]. Biotechnology Bulletin, 2021, 37(1): 246-254. |
| [5] | CHEN Yi-dan, ZHANG Yu, YANG Jie, ZHANG Qin, JIANG Li. Exploration of Key Functional Genes Affecting Milk Production Traits in Dairy Cattle Based on RNA-seq [J]. Biotechnology Bulletin, 2020, 36(9): 244-252. |
| [6] | HU Bin-yue, HU Yang, CHENG Wen-min, ZHAO Su-mei, ZHAO Hong-Ye, WEI Hong-Jiang. Lipid Droplet Formation in the Pre-adipocytes of Leptin-overexpressed Pig [J]. Biotechnology Bulletin, 2020, 36(8): 111-119. |
| [7] | ZHANG De-rong, MA Xiao-xia, LI Yu-fei, ZHAO Yong-qing, HUO Sheng-dong, MA Zhong-ren, BAI Jia-lin. Research Progress and Prospects of Adiponectin and Its Receptor in Mammal [J]. Biotechnology Bulletin, 2020, 36(6): 236-244. |
| [8] | YU Jun-jian, CHI Mei-li, JIA Yong-yi, LIU Shi-li, ZHU Jun-quan, GU Zhi-min. Tetra-primer Amplification Refractory Mutation System PCR and Its Application in Fauna and Flora Genetics and Breeding Research [J]. Biotechnology Bulletin, 2020, 36(5): 32-38. |
| [9] | LI Xiao-kai, FAN Yi-xing, QIAO Xian, ZHANG Lei, WANG Feng-hong, WANG Zhi-ying, WANG Rui-jun, ZHANG Yan-jun, LIU Zhi-hong, WANG Zhi-xin, HE Li-bing, LI Jin-quan, SU Rui, ZHANG Jia-xin. Research Progress of Goat Genome and Genetic Variation Map [J]. Biotechnology Bulletin, 2020, 36(4): 175-184. |
| [10] | LI Biao, ZHANG Rui-ying, WANG Xiao-qi, ZHANG Cun-fang, DUAN Zi-yuan. Microsatellite Polymorphism and Its Correlation Analysis with Body Size Traits of Tan Sheep [J]. Biotechnology Bulletin, 2019, 35(6): 131-137. |
| [11] | HUANG Long, WU Ben-li, HE Ji-xiang, CHEN Jing, SONG Guang-tong, WANG Xiang, ZHANG Ye, WU Song. SNP Identification of MyoD1 Gene and Its Correlation with Growth Traits in Pelodiscus sinensis [J]. Biotechnology Bulletin, 2019, 35(4): 76-81. |
| [12] | GUO Meng-meng, ZHOU Yan-qing, DUAN Hong-ying, YANG Ke, SHAO Lu-ying. Development of SNP Marker Based on the Rehmannia glutinosa Transcriptome Database and Construction of DNA Fingerprint in Rehmannia [J]. Biotechnology Bulletin, 2019, 35(11): 224-230. |
| [13] | WANG Yan-xin, LIAO Yuan-yuan, A Yimuguli, QI Ao-qiong, LI Hai-jian, XU Hong-wei, YANG Ju-tian, CAI Yong. Correlation Analysis Between LHX3 Gene Polymorphism and Growth Traits in Three Sheep Breeds [J]. Biotechnology Bulletin, 2019, 35(10): 162-168. |
| [14] | JI Hui, WANG Hui, CHAI Zhi-xin, WANG Ji-kun, LUO Xiao-lin, JI Qiu-mei, XIN Jin-wei, ZHONG Jin-cheng. Precursor Cloning and Tissue Expression Analysis of Yak miR-378 [J]. Biotechnology Bulletin, 2019, 35(1): 58-66. |
| [15] | LI Xiao-kai ,WANG Gui ,QIAO Xian ,FAN Yi-xing ,ZHANG Lei ,MA Yu-hao ,NIE Rui-xue ,WANG Rui-jun ,HE Li-bing ,SU Rui. Research Progress on Whole-genome Sequencing on Important Domesticated Animals [J]. Biotechnology Bulletin, 2018, 34(6): 11-21. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||