








生物技术通报 ›› 2026, Vol. 42 ›› Issue (4): 92-100.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0903
许孟歌( ), 宋火焱, 罗佳, 苏亿, 周会汶, 王灿, 孔可可(
), 宋火焱, 罗佳, 苏亿, 周会汶, 王灿, 孔可可( )
)
收稿日期:2025-08-20
出版日期:2026-04-26
发布日期:2026-04-30
通讯作者:
孔可可,男,博士,讲师,研究方向 :大豆遗传育种及抗逆性分子机理;E-mail: kekekong2011011034@163.com作者简介:许孟歌,女,博士,讲师,研究方向 :大豆分子育种及优异性状遗传解析;E-mail: 943723370@qq.com
基金资助:
XU Meng-ge( ), SONG Huo-yan, LUO Jia, SU Yi, ZHOU Hui-wen, WANG Can, KONG Ke-ke(
), SONG Huo-yan, LUO Jia, SU Yi, ZHOU Hui-wen, WANG Can, KONG Ke-ke( )
)
Received:2025-08-20
Published:2026-04-26
Online:2026-04-30
摘要:
目的 针对前期挖掘到的百粒重候选基因GmRLK19,分析其表达模式并筛选互作蛋白,为进一步研究GmRLK19的生物学功能提供依据。 方法 利用生物信息学分析GmRLK19基因的结构特点;利用荧光定量PCR技术分析GmRLK19的组织表达模式;利用亚细胞定位确定其蛋白表达位置;利用酵母双杂交技术筛选GmRLK19的互作蛋白并预测其潜在功能。 结果 GmRLK19蛋白的二级结构主要为无规则卷曲和α-螺旋,可能存在的磷酸化位点共有145个。GmRLK19基因在被检测的组织中均有表达且在开花后30 d的籽粒中表达水平最高;亚细胞定位结果表明其蛋白主要定位在细胞膜。通过酵母双杂筛选互作蛋白,共获得187个阳性克隆,测序分析后得到148个候选互作蛋白;功能预测结果显示,互作蛋白主要在调控植物生长发育、信号转导、代谢过程等方面发挥作用。GO富集结果显示互作蛋白在光合作用和蛋白质翻译条目中显著富集。 结论 GmRLK19基因在多个组织中表达,其蛋白主要定位在细胞膜。筛选得到148个候选互作蛋白,推测GmRLK19通过植物激素信号转导、光合作用等途径影响大豆粒重。
许孟歌, 宋火焱, 罗佳, 苏亿, 周会汶, 王灿, 孔可可. 大豆GmRLK19基因表达分析及互作蛋白的筛选[J]. 生物技术通报, 2026, 42(4): 92-100.
XU Meng-ge, SONG Huo-yan, LUO Jia, SU Yi, ZHOU Hui-wen, WANG Can, KONG Ke-ke. Expression Analysis and Interaction Protein Screening of GmRLK19 in Soybean[J]. Biotechnology Bulletin, 2026, 42(4): 92-100.
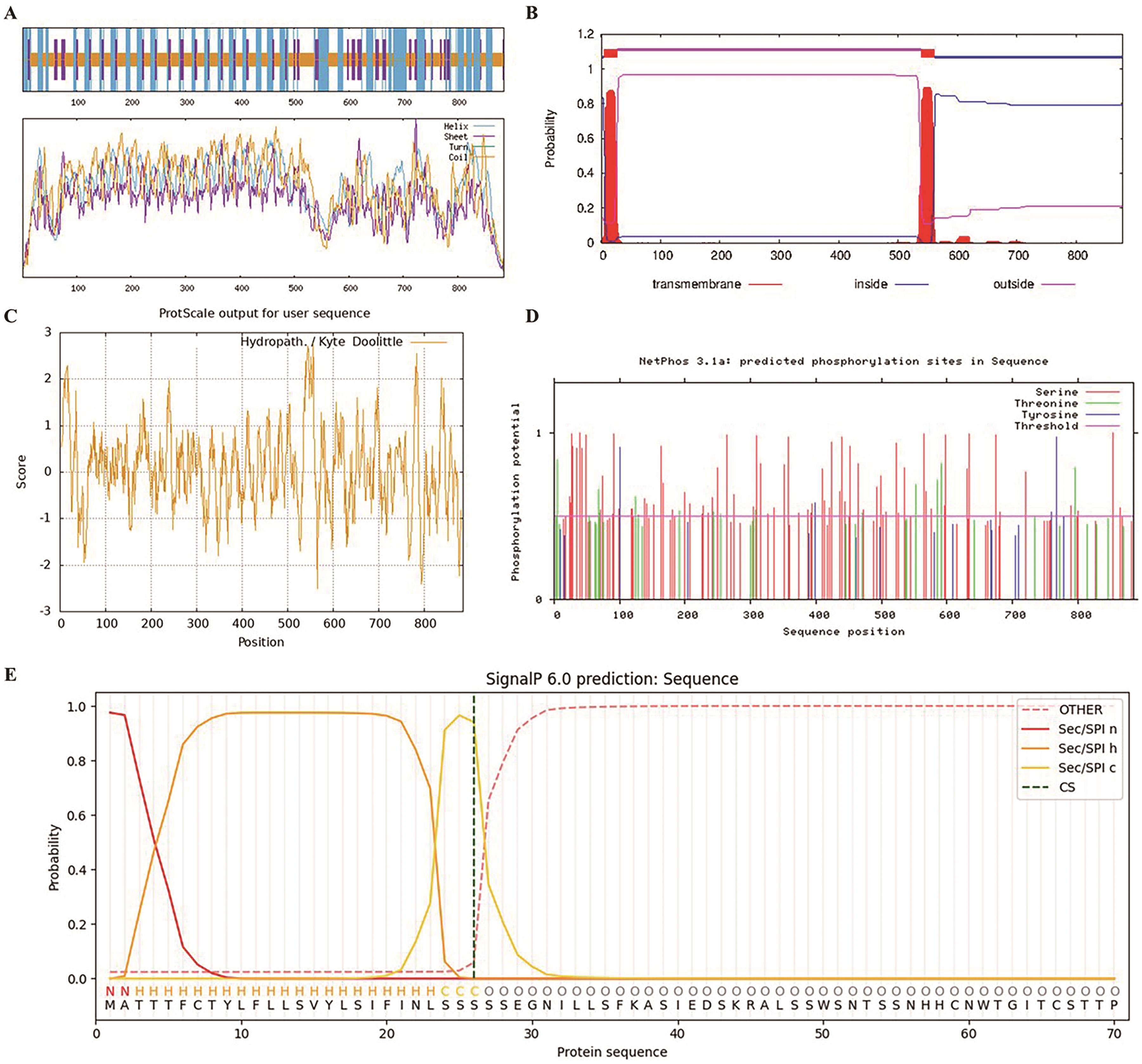
图1 GmRLK19的生物信息学分析A:二级结构预测;B:跨膜结构预测;C:亲/疏水性预测;D:磷酸化位点预测;E:信号肽预测
Fig. 1 Bioinformatics analysis of GmRLK19A: Secondary structure prediction; B: transmembrane structure prediction; C: hydrophilic/hydrophobic prediction; D: phosphorylation site prediction; E: signal peptide prediction
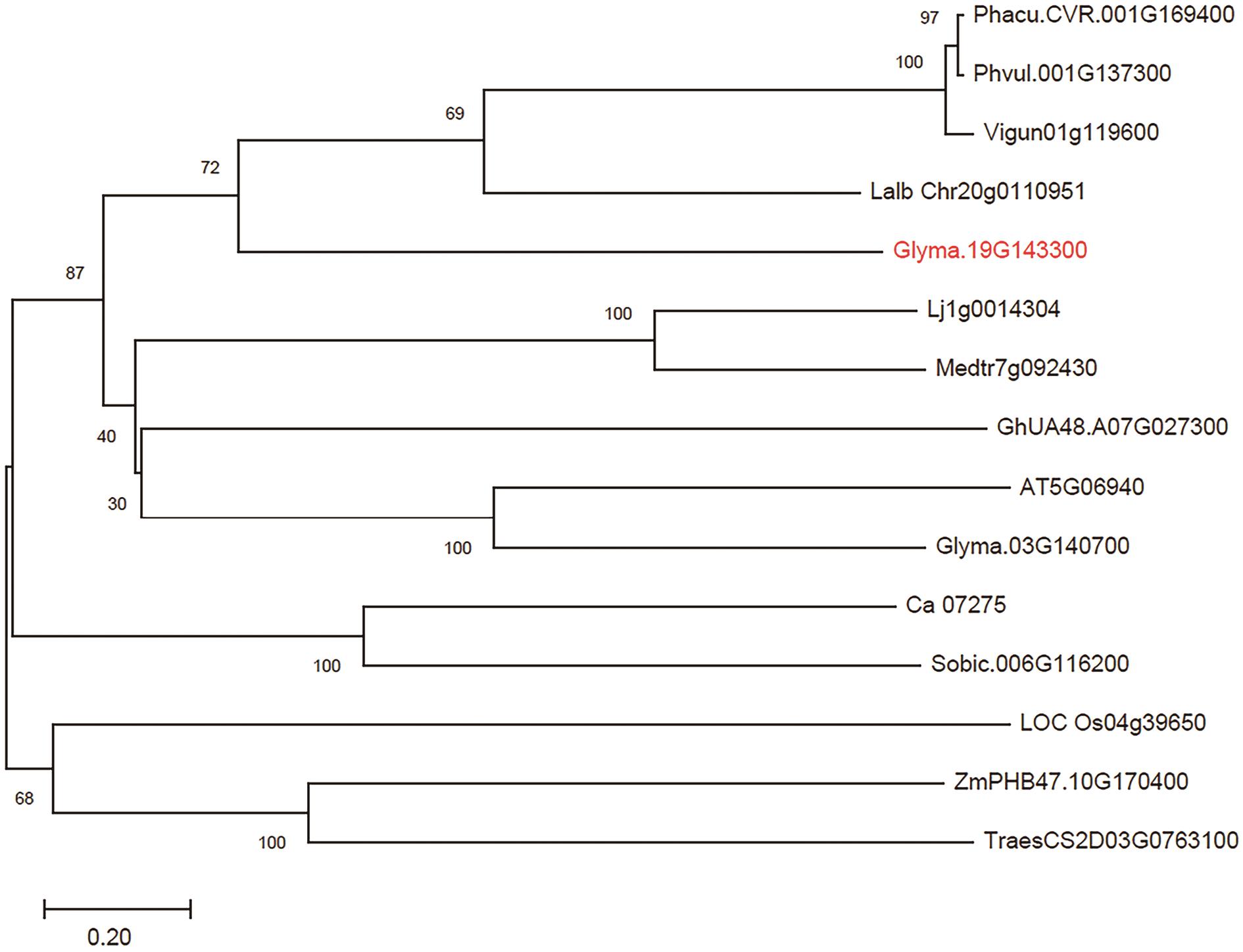
图2 GmRLK19及其同源蛋白的系统进化树分析GmRLK19及其同源蛋白的进化树分析,节点上的数字表示1 000次Bootstrap值。使用与GmRLK19同源的14个蛋白质的氨基酸全长序列绘制,包括,Phacu.CVR.001G169400,尖叶菜豆;Phvul.001G137300,菜豆/四季豆;Vigun01g119600,豇豆;Lalb Chr20g0110951,白羽扇豆;Glyma.19G143300,大豆;Lj1g0014304,百脉根;Medtr7g092430,蒺藜苜蓿;GhUA48.A07G027300,棉花;AT5G06940,拟南芥;Glyma.03G140700,大豆;Ca_07275,鹰嘴豆;Sobic.006G116200,高粱;LOC Os04g39650,水稻;ZmPHB47.10G170400,玉米;TraesCS2D03G0763100,小麦
Fig. 2 Phylogenetic tree analysis of GmRLK19 and its homologous proteinPhylogenetic tree analysis of GmRLK19 and homologous proteins. The numbers in the nodes indicate 1 000-time Bootstrap. It is depicted using the full-length amino acid sequences of the 14 proteins that are homologous with GmRLK19, including Phacu.CVR.001G169400 (Phaseolus vulgaris); Phvul.001G137300 (Phaseolus acutifolius); Vigun01g119600 (Vigna unguiculata); Lalb Chr20g0110951 (Lupinus albus); Glyma.19G143300 (Glycine max); Lj1g0014304 (Lotus japonicus); Medtr7g092430 (Medicago truncatula); GhUA48.A07G027300 (Gossypium hirsutum); AT5G06940 (Arabidopsis thaliana); Glyma.03G140700 (Glycine max); Ca_07275 (Cicer arietinum); Sobic.006G116200 (Sorghum bicolor); LOC Os04g39650 (Oryza sativa); ZmPHB47.10G170400 (Zea mays); TraesCS2D03G0763100 (Triticum aestivum)
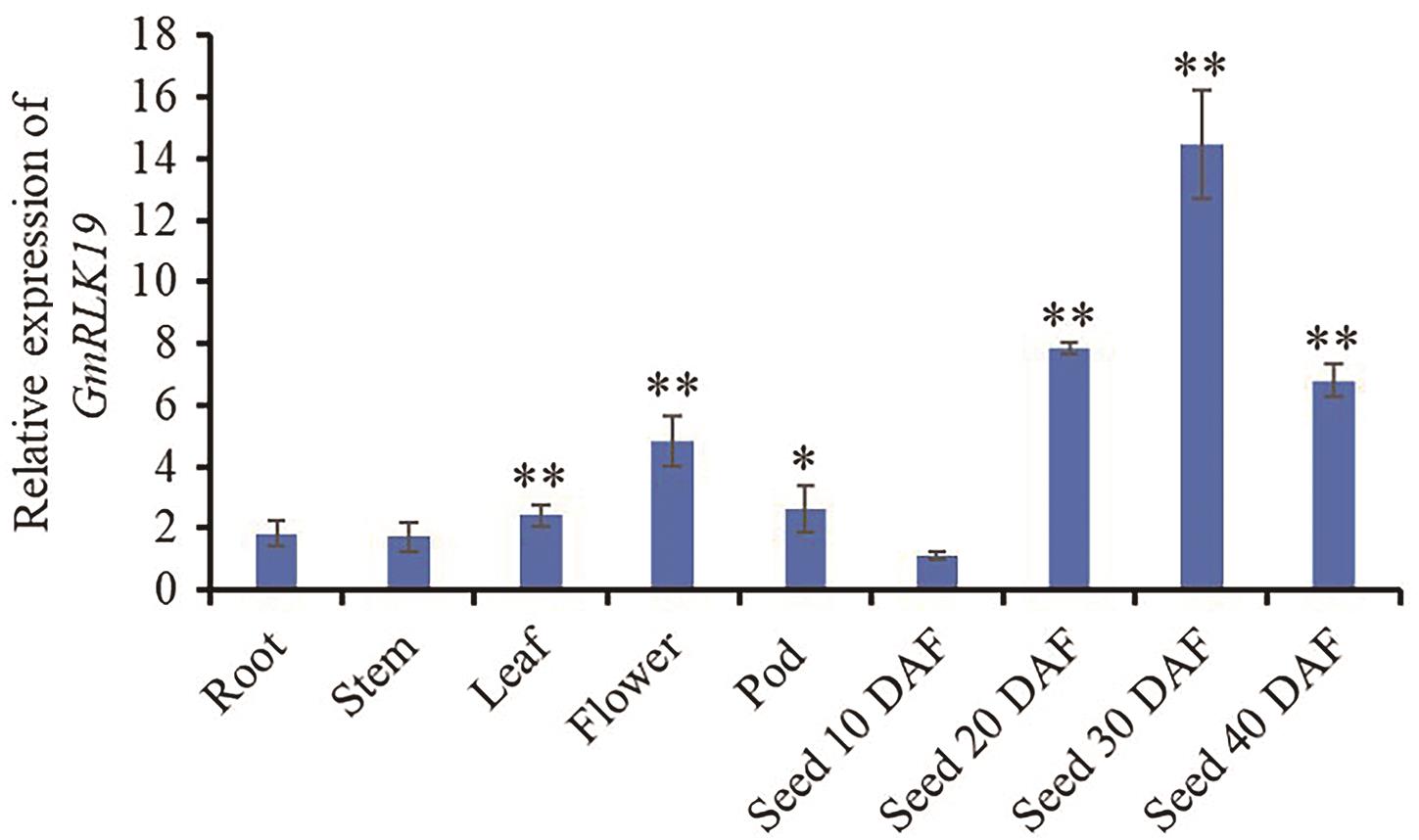
图3 GmRLK19在大豆不同组织中的相对表达量各组织样品分别选自PI595843品种14 d苗龄的根、茎、叶、2 cm的荚以及开花后10-40 d发育的籽粒。不同组织和不同发育时期籽粒的基因相对表达量均分别以开花后10 d的籽粒中的表达量进行标准化,即作为对照(相对表达量 = 1)。误差线代表标准误差(n = 3)。Root:根;Stem:茎;Leaf:叶片;Flower:花;Pod:荚;Seed 10 DAF - Seed 40 DAF分别表示开花后10、20、30和40 d的籽粒。*和**分别表示各个组织与10 DAF籽粒在0.05和0.01水平上的差异显著性(Student’s t-test, two-tail)
Fig. 3 Relative expressions of GmRLK19 gene in different soybean tissuesThe tissue samples were selected from the root, stem, leaf and 2 cm pod of the PI595843 of 14-day old seedlings, and the seeds at different development stages at 10-40 d after flowering were also collected. The relative expressions were normalized in the seed at 10DAF from PI595843, as the control (relative expression = 1). The error bars indicate the standard error (n = 3). Seed 10 DAF-Seed 40 DAF: the seed of the 10, 20, 30, and 40 d after flowering, respectively. * and ** indicate significant difference between tissue samples and seed at 10 DAF at 0.01 and 0.05 level (Student’s t-test, two-tail), respectively
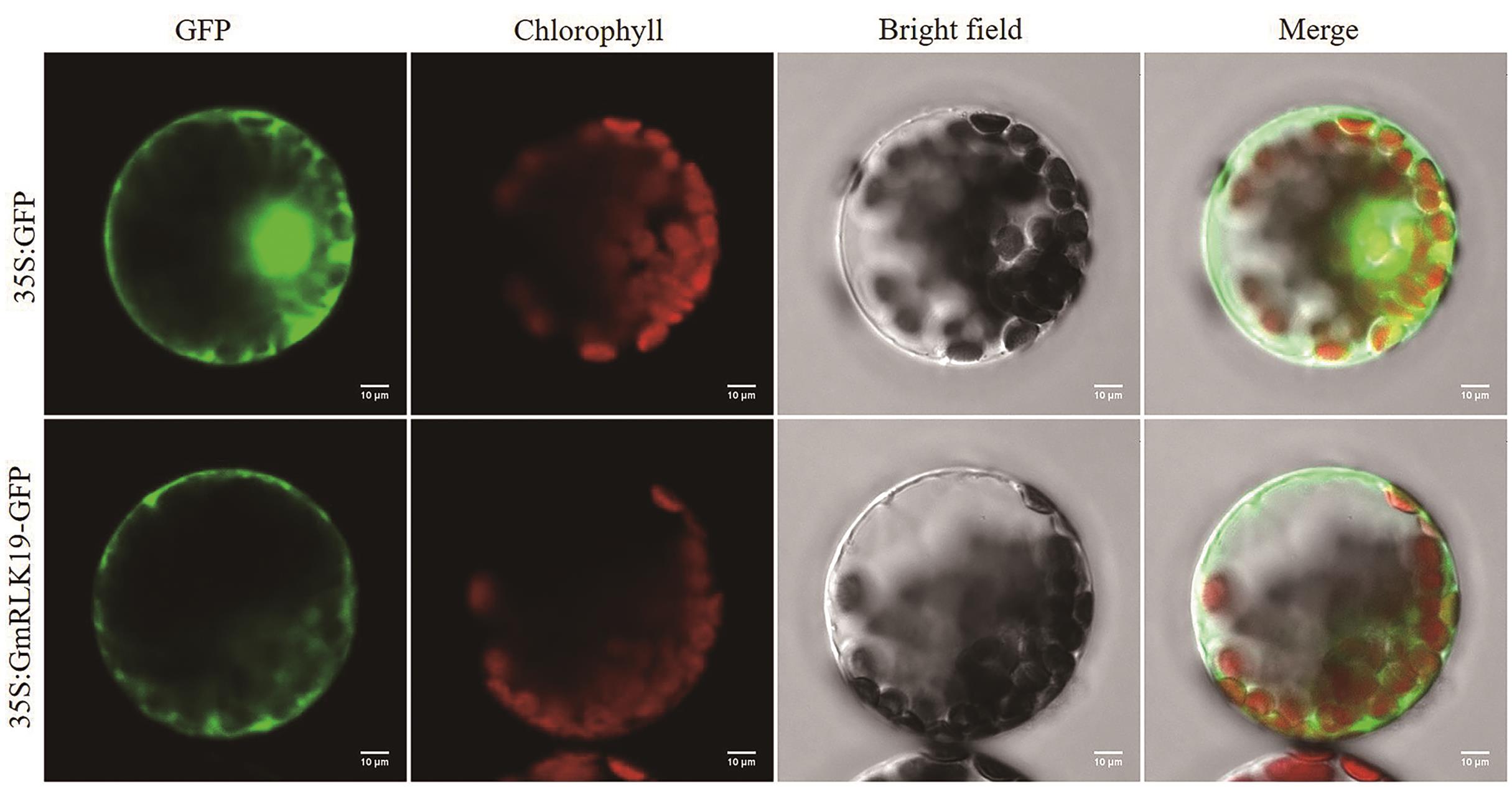
图4 GmRLK19蛋白的亚细胞定位在CaMV 35S启动子控制下,GFP蛋白及GmRLK19-GFP融合蛋白在拟南芥原生质体中的瞬时表达。GFP,绿色荧光蛋白;Chlorophyll,叶绿体自发荧光;Bright field,明场;Merge,叠加。比例尺为10 μm
Fig. 4 Subcellular localization of GmRLK19 proteinTransient expression of GFP protein or GmRLK19-GFP fusion protein under the control of CaMV 35S promoter in tobacco cells. GFP, green fluorescence protein. Chlorophyll, chloroplast spontaneous fluorescence. Bright field, brighten field. Merged, merged images of the GFP, chlorophyll, and bright field images. Scale bar = 10 μm
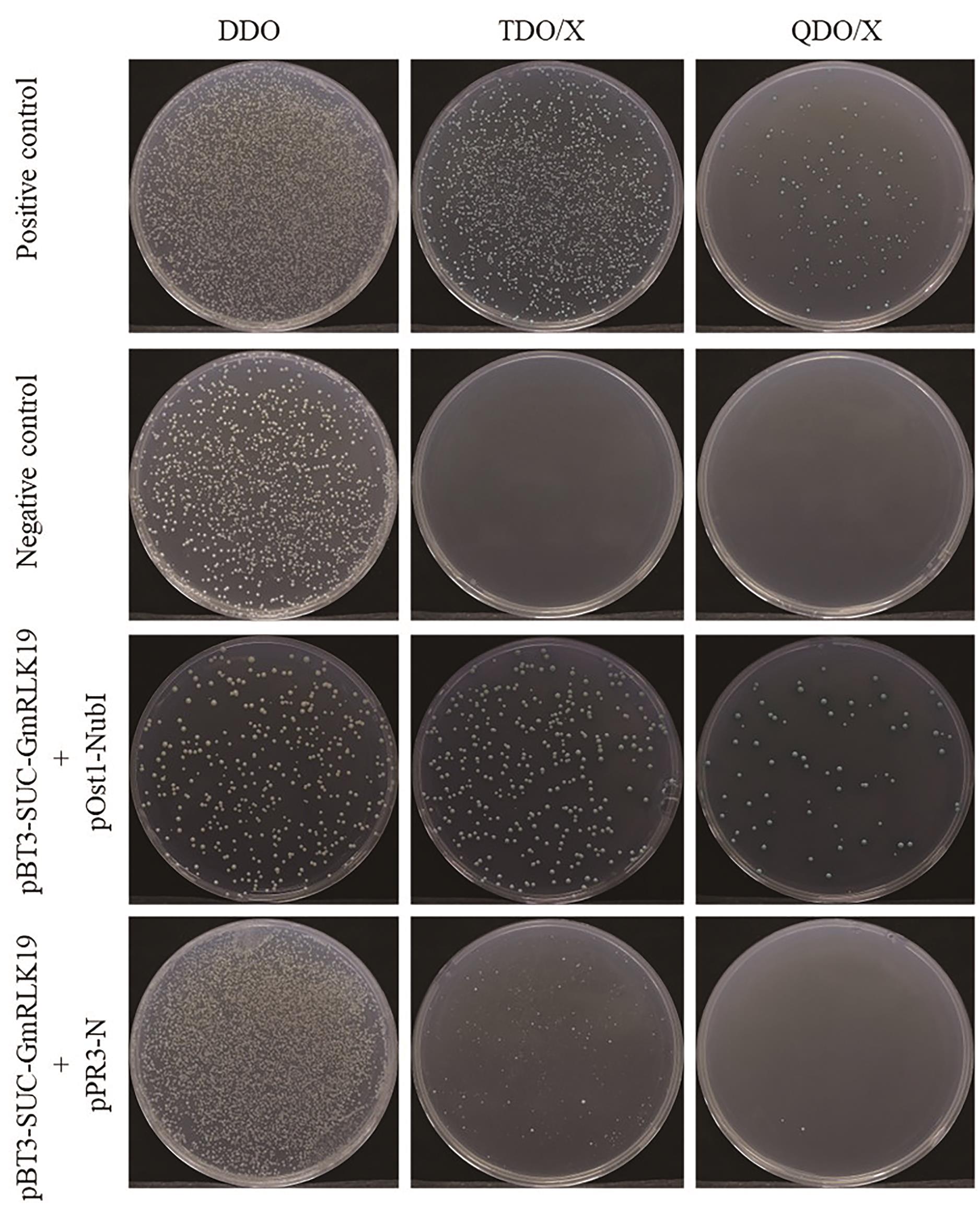

图5 自激活检测结果从上到下依次为阳性对照、阴性对照、功能验证组及自激活组;从左到右依次为DDO、TDO/X和QDO/X平板
Fig. 5 Results of self-activation detectionFrom top to bottom, the groups are: positive control, negative control, functional validation group, and self-activation group. From left to right, the plates are: DDO, TDO/X, and QDO/X plate
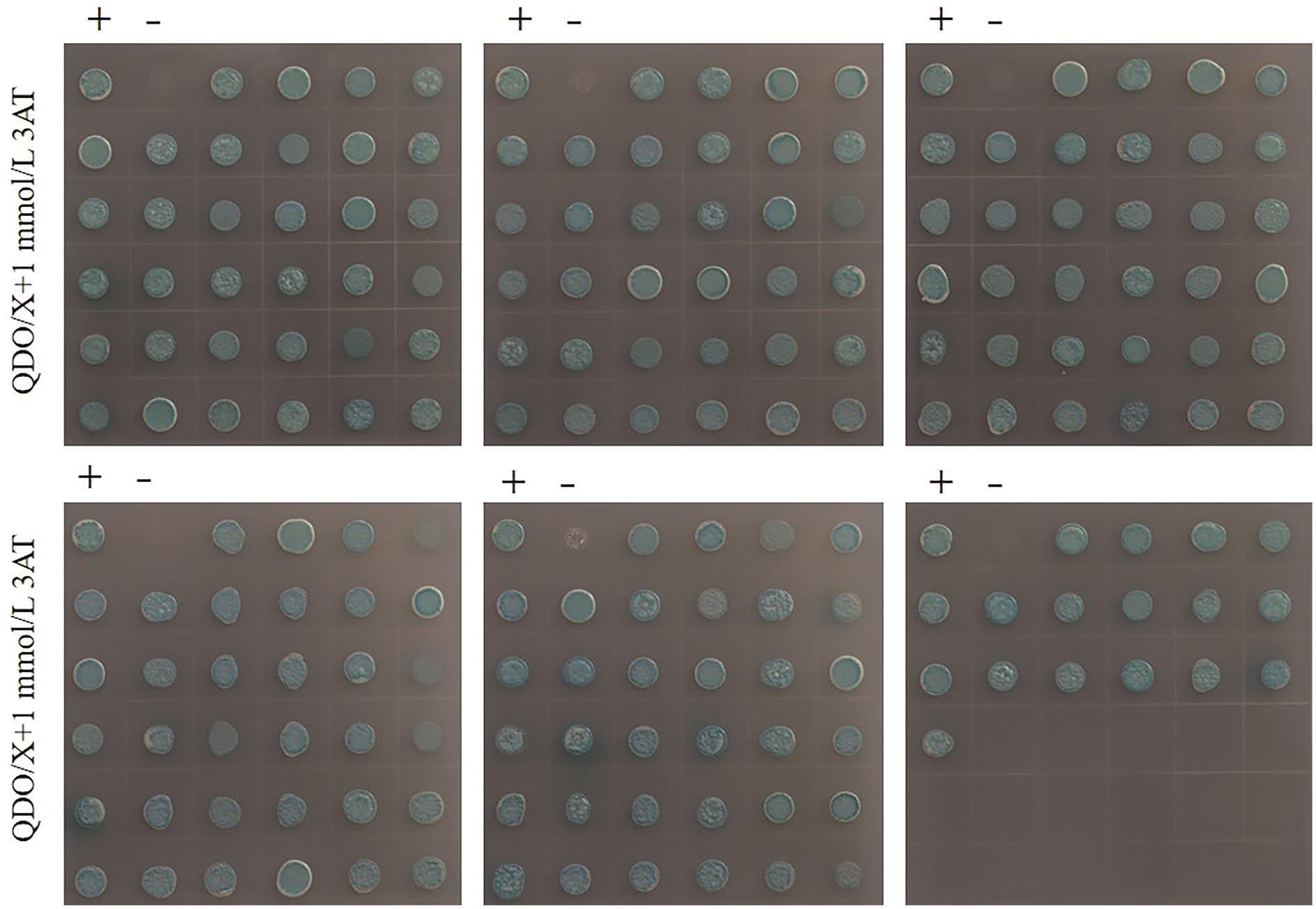

图6 双杂交文库筛选获得的187个阳性克隆“+”代表阳性对照,pTSU2-APP+NubG-Fe65;“-”代表阴性对照,pTSU2-APP+pPR3-N
Fig. 6 The 187 positive clones obtained double hybridization library screening“+” indicates positive control, pTSU2-APP+NubG-Fe65; “-” indicates negative control, pTSU2-APP+pPR3-N
| [1] | Kato S, Sayama T, Fujii K, et al. A major and stable QTL associated with seed weight in soybean across multiple environments and genetic backgrounds [J]. Theor Appl Genet, 2014, 127(6): 1365-1374. |
| [2] | Gao CX. Genome engineering for crop improvement and future agriculture [J]. Cell, 2021, 184(6): 1621-1635. |
| [3] | Li J, Zhang YH, Ma RR, et al. Identification of ST1 reveals a selection involving hitchhiking of seed morphology and oil content during soybean domestication [J]. Plant Biotechnol J, 2022, 20(6): 1110-1121. |
| [4] | Duan ZB, Zhang M, Zhang ZF, et al. Natural allelic variation of GmST05 controlling seed size and quality in soybean [J]. Plant Biotechnol J, 2022, 20(9): 1807-1818. |
| [5] | Liang S, Duan ZB, He XM, et al. Natural variation in GmSW17 controls seed size in soybean [J]. Nat Commun, 2024, 15: 7417. |
| [6] | Goettel W, Zhang HY, Li Y, et al. POWR1 is a domestication gene pleiotropically regulating seed quality and yield in soybean [J]. Nat Commun, 2022, 13: 3051. |
| [7] | Di Q, Dong LD, Jiang L, et al. Genome-wide association study and RNA-seq identifies GmWRI1-like transcription factor related to the seed weight in soybean [J]. Front Plant Sci, 2023, 14: 1268511. |
| [8] | Wang SD, Yokosho K, Guo RZ, et al. The soybean sugar transporter GmSWEET15 mediates sucrose export from endosperm to early embryo [J]. Plant Physiol, 2019, 180(4): 2133-2141. |
| [9] | Wang SD, Liu SL, Wang J, et al. Simultaneous changes in seed size, oil content and protein content driven by selection of SWEET homologues during soybean domestication [J]. Natl Sci Rev, 2020, 7(11): 1776-1786. |
| [10] | Hu Y, Liu Y, Tao JJ, et al. GmJAZ3 interacts with GmRR18a and GmMYC2a to regulate seed traits in soybean [J]. J Integr Plant Biol, 2023, 65(8): 1983-2000. |
| [11] | Hu Y, Liu Y, Lu L, et al. Global analysis of seed transcriptomes reveals a novel PLATZ regulator for seed size and weight control in soybean [J]. New Phytol, 2023, 240(6): 2436-2454. |
| [12] | Soltabayeva A, Dauletova N, Serik S, et al. Receptor-like kinases (LRR-RLKs) in response of plants to biotic and abiotic stresses [J]. Plants, 2022, 11(19): 2660. |
| [13] | Morinaka Y, Sakamoto T, Inukai Y, et al. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice [J]. Plant Physiol, 2006, 141(3): 924-931. |
| [14] | Jiang YH, Bao L, Jeong SY, et al. XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice [J]. Plant J, 2012, 70(3): 398-408. |
| [15] | Garcia D, Saingery V, Chambrier P, et al. Arabidopsis haiku mutants reveal new controls of seed size by endosperm [J]. Plant Physiol, 2003, 131(4): 1661-1670. |
| [16] | Luo M, Dennis ES, Berger F, et al. MINISEED3 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR)KINASE gene, are regulators of seed size in Arabidopsis [J]. Proc Natl Acad Sci USA, 2005, 102(48): 17531-17536. |
| [17] | He CM, Wang J, Dong R, et al. Overexpression of an antisense RNA of maize receptor-like kinase gene ZmRLK7 enlarges the organ and seed size of transgenic Arabidopsis plants [J]. Front Plant Sci, 2020, 11: 579120. |
| [18] | Zhou FL, Guo Y, Qiu LJ. Genome-wide identification and evolutionary analysis of leucine-rich repeat receptor-like protein kinase genes in soybean [J]. BMC Plant Biol, 2016, 16(1): 58. |
| [19] | Kim S, Kim SJ, Shin YJ, et al. An atypical soybean leucine-rich repeat receptor-like kinase, GmLRK1, may be involved in the regulation of cell elongation [J]. Planta, 2009, 229(4): 811-821. |
| [20] | Li XP, Gan R, Li PL, et al. Identification and functional characterization of a leucine-rich repeat receptor-like kinase gene that is involved in regulation of soybean leaf senescence [J]. Plant Mol Biol, 2006, 61(6): 829-844. |
| [21] | Yang L, Wu KC, Gao P, et al. GsLRPK, a novel cold-activated leucine-rich repeat receptor-like protein kinase from Glycine soja, is a positive regulator to cold stress tolerance [J]. Plant Sci, 2014, 215/216: 19-28. |
| [22] | Xu MG, Kong KK, Miao L, et al. Identification of major quantitative trait loci and candidate genes for seed weight in soybean [J]. Theor Appl Genet, 2023, 136(1): 22. |
| [23] | Larkin MA, Blackshields G, Brown NP, et al. Clustal W and clustal X version 2.0 [J]. Bioinformatics, 2007, 23(21): 2947-2948. |
| [24] | Tamura K, Stecher G, Peterson D, et al. MEGA6: molecular evolutionary genetics analysis version 6.0 [J]. Mol Biol Evol, 2013, 30(12): 2725-2729. |
| [25] | Yoo SD, Cho YH, Sheen J. Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis [J]. Nat Protoc, 2007, 2(7): 1565-1572. |
| [26] | Dievart A, Gottin C, Périn C, et al. Origin and diversity of plant receptor-like kinases [J]. Annu Rev Plant Biol, 2020, 71: 131-156. |
| [27] | Gampala SS, Kim TW, He JX, et al. An essential role for 14-3-3 proteins in brassinosteroid signal transduction in Arabidopsis [J]. Dev Cell, 2007, 13(2): 177-189. |
| [28] | Parry LIC G. Complex regulation of the TIR1/AFB family of auxin receptors [J]. Proc Natl Acad Sci U S A, 2009, 106(52): 22540-22545. |
| [29] | Qu J, Kang SG, Hah C, et al. Molecular and cellular characterization of GA-Stimulated Transcripts GASA4 and GASA6 in Arabidopsis thaliana [J]. Plant Sci, 2016, 246: 1-10. |
| [30] | Li N, Xu R, Li YH. Molecular networks of seed size control in plants [J]. Annu Rev Plant Biol, 2019, 70: 435-463. |
| [31] | Lu X, Xiong Q, Cheng T, et al. A PP2C-1 allele underlying a quantitative trait locus enhances soybean 100-seed weight [J]. Mol Plant, 2017, 10(5): 670-684. |
| [32] | Jia ML, Li YN, Wang ZY, et al. TaIAA21 represses TaARF25-mediated expression of TaERFs required for grain size and weight development in wheat [J]. Plant J, 2021, 108(6): 1754-1767. |
| [33] | Wang L, Yang YM, Yang ZY, et al. GmFtsH25 overexpression increases soybean seed yield by enhancing photosynthesis and photosynthates [J]. J Integr Plant Biol, 2023, 65(4): 1026-1040. |
| [34] | Hu DZ, Li X, Yang ZY, et al. Downregulation of a gibberellin 3β- hydroxylase enhances photosynthesis and increases seed yield in soybean [J]. New Phytol, 2022, 235(2): 502-517. |
| [1] | 苏燕竹, 李达, 张爱爱, 刘永光, 张秀荣, 薛其勤. 大豆CAD基因家族的鉴定及表达分析[J]. 生物技术通报, 2026, 42(4): 101-113. |
| [2] | 刘青媛, 吴洪启, 陈秀娥, 陈剑, 姜远泽, 何燕子, 喻奇伟, 刘仁祥. 转录因子NtMYB96a调控烟草耐旱性的功能研究[J]. 生物技术通报, 2026, 42(4): 239-250. |
| [3] | 陈登科, 兰刚, 夏芝, 侯保国, 杨六六, 曹彩荣, 李朋波, 吴翠翠. 花生ZF-HD基因家族的鉴定和非生物胁迫响应分析[J]. 生物技术通报, 2026, 42(4): 114-128. |
| [4] | 江昕桦, 方天宇, 张晶晶, 李相媛, 张邦跃, 廖晓珊, 荣朵艳. 地钱MpPP2A-C基因的鉴定及功能分析[J]. 生物技术通报, 2026, 42(4): 216-226. |
| [5] | 刘林娅, 刘欢艳, 梁鑫钰, 宋姝熠, 何斌, 王绪英, 黄亚成. ‘红阳’猕猴桃BGAL基因家族的全基因组鉴定与表达分析[J]. 生物技术通报, 2026, 42(3): 312-323. |
| [6] | 张高翔, 吴玉碧, 郭亚静, 纪薇, 杨忠义. 葡萄WD40基因家族鉴定及表达量分析[J]. 生物技术通报, 2026, 42(3): 324-337. |
| [7] | 李天源, 亓新亮, 刘珊, 张建成, 王鹏飞, 张帅, 贾璐婷, 穆霄鹏. 欧李SPL基因家族的鉴定及在果实发育过程中的表达分析[J]. 生物技术通报, 2026, 42(3): 362-373. |
| [8] | 杨子涵, 李葵秀, 李俊良, 吕文慧, 何温婷, 刘旭燕, 刘冠泽. 三七RALF基因家族鉴定及表达分析[J]. 生物技术通报, 2026, 42(2): 278-292. |
| [9] | 任云儿, 伍国强, 成斌, 魏明. 甜菜BvATGs基因家族全基因组鉴定及盐胁迫下表达模式分析[J]. 生物技术通报, 2026, 42(1): 184-197. |
| [10] | 杨跃琴, 邢英, 仲子荷, 田维军, 杨雪清, 王建旭. 甲基汞胁迫下水稻OsMATE34的表达及功能分析[J]. 生物技术通报, 2026, 42(1): 86-94. |
| [11] | 张月, 戴月华, 张莹莹, 李奥辉, 李楚慧, 薛金爱, 秦慧彬, 陈妍, 聂萌恩, 张海平. 大豆烯酰辅酶A还原酶ECR14基因的克隆与功能分析[J]. 生物技术通报, 2026, 42(1): 95-104. |
| [12] | 陈静欢, 房国楠, 朱文豪, 叶广继, 苏旺, 贺苗苗, 杨生龙, 周云. 马铃薯种质资源淀粉表征及相关基因表达分析[J]. 生物技术通报, 2026, 42(1): 170-183. |
| [13] | 李亚涛, 张志鹏, 赵梦瑶, 吕镇, 甘恬, 魏浩, 吴书凤, 马玉超. 根瘤菌Bd1的全基因组分析及TetR3对细胞生长和结瘤的负调控功能[J]. 生物技术通报, 2025, 41(9): 289-301. |
| [14] | 李珊, 马登辉, 马红义, 姚文孔, 尹晓. 葡萄SKP1基因家族鉴定与表达分析[J]. 生物技术通报, 2025, 41(9): 147-158. |
| [15] | 黄国栋, 邓宇星, 程宏伟, 但焱南, 周会汶, 吴兰花. 大豆ZIP基因家族鉴定及响应铝胁迫的表达分析[J]. 生物技术通报, 2025, 41(9): 71-81. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||