








Biotechnology Bulletin ›› 2026, Vol. 42 ›› Issue (3): 283-293.doi: 10.13560/j.cnki.biotech.bull.1985.2025-0704
Previous Articles Next Articles
LUO Wei1,2( ), GONG Ao1, ZHONG Yang3, HU Di1, ZHOU Hong-yuan1, ZHANG Hong-xin1, AI Ju1, LUO You-wei4, GAO Dong-li1(
), GONG Ao1, ZHONG Yang3, HU Di1, ZHOU Hong-yuan1, ZHANG Hong-xin1, AI Ju1, LUO You-wei4, GAO Dong-li1( )
)
Received:2025-07-01
Online:2026-03-26
Published:2026-04-23
Contact:
GAO Dong-li
E-mail:1136689701@qq.com;gdongli@126.com
LUO Wei, GONG Ao, ZHONG Yang, HU Di, ZHOU Hong-yuan, ZHANG Hong-xin, AI Ju, LUO You-wei, GAO Dong-li. Pleiotropic Effects of SEPALLATA2 Knock-out on Fruit and Wart Development in Cucumber[J]. Biotechnology Bulletin, 2026, 42(3): 283-293.
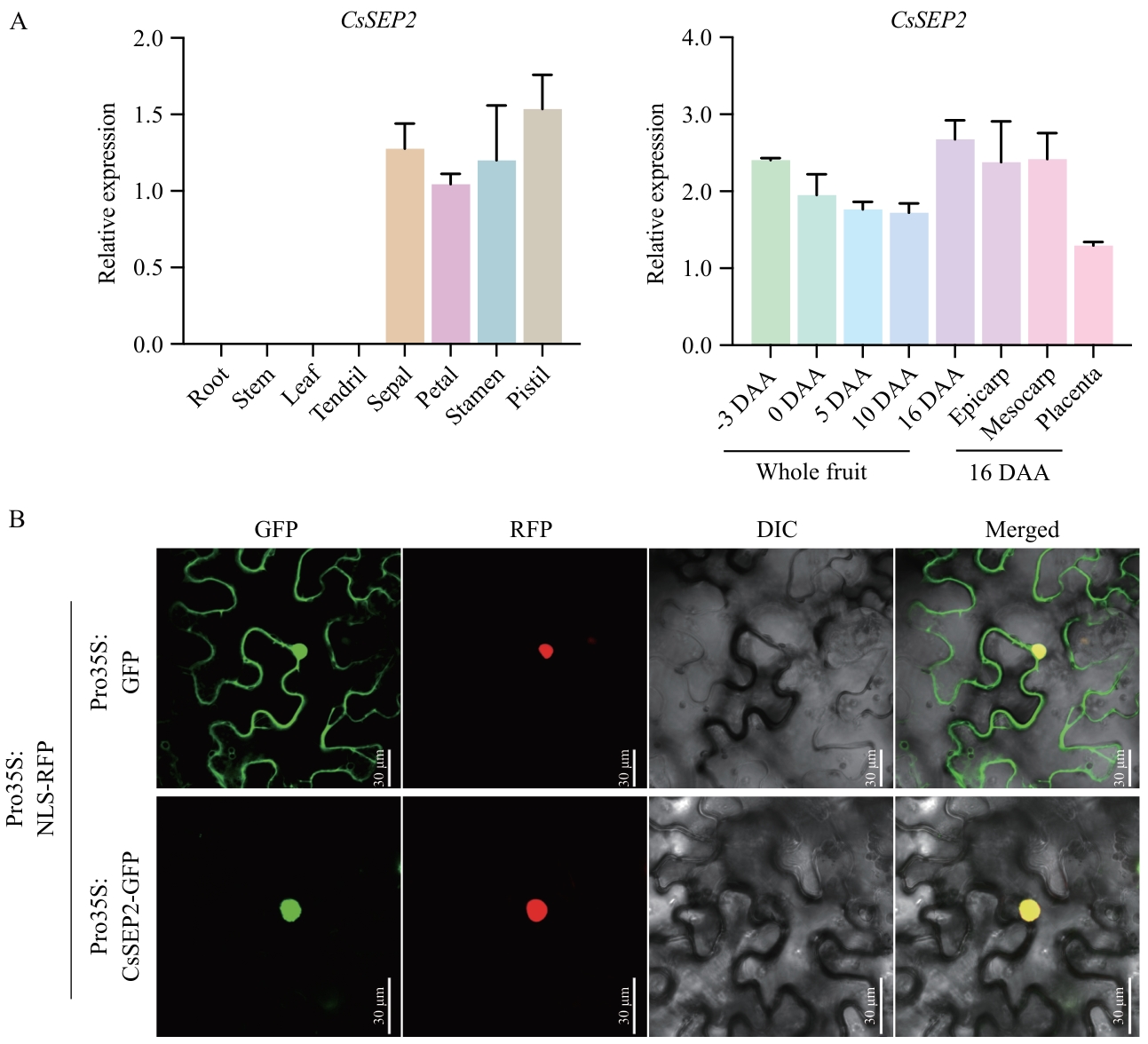
Fig. 1 Expression analyses (A) and subcellular localization (B) of gene CsSEP2In Fig. A, the expressions of CsSEP2 in thefruits of -3, 0, 5, 10, and 16 DAA and epicarp, mesocarp and placenta from the central parts of fruits at 16 DAA were examined. Data are means ± SD, n = 3. DAA refers to days after anthesis, the same below
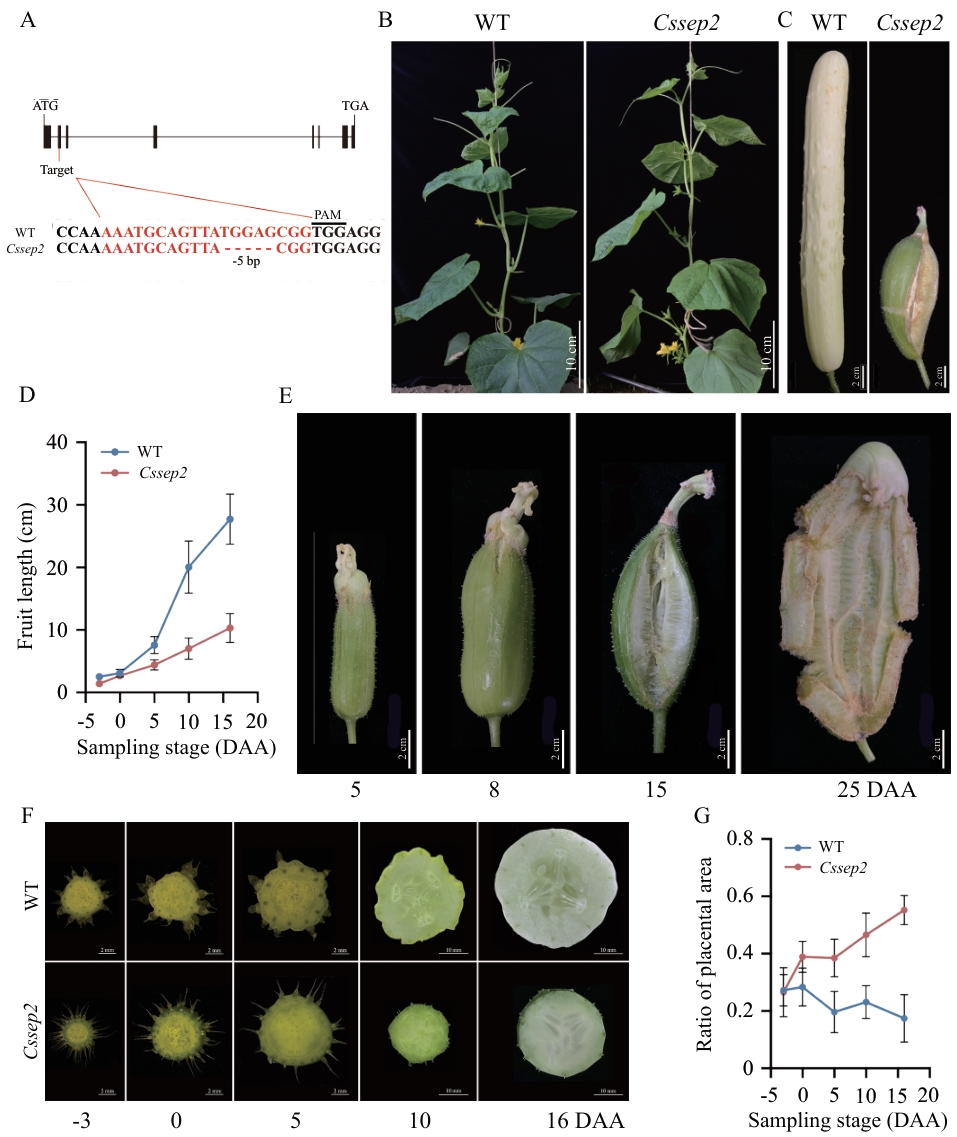
Fig. 2 Knock-out of gene CsSEP2 resulted in abnormal fruit developmentA: Structure of the CsSEP2 gene and the mutated loci in the Cssep2 mutants. Exons and introns are denoted by black boxes and lines, respectively. The red nucleotide sequences are the editing target site of the gene, and the three underlined bases indicate the protospacer adjacent motif (PAM). B, C: Representative plants (B) and fruits (C) of WT and Cssep2 mutants. D: Measurement of the fruit lengths of WT and Cssep2 mutants. Data are means ± SD, n = 5. E: Macroscopic observation of fruit dehiscence of the Cssep2 mutants. F: Comparative observation of the fruit placentas of WT and Cssep2 mutants. G: The ratio of placenta areas to fruit cross areas of WT and Cssep2 mutants. Values are means ± SD, n ≥ 5
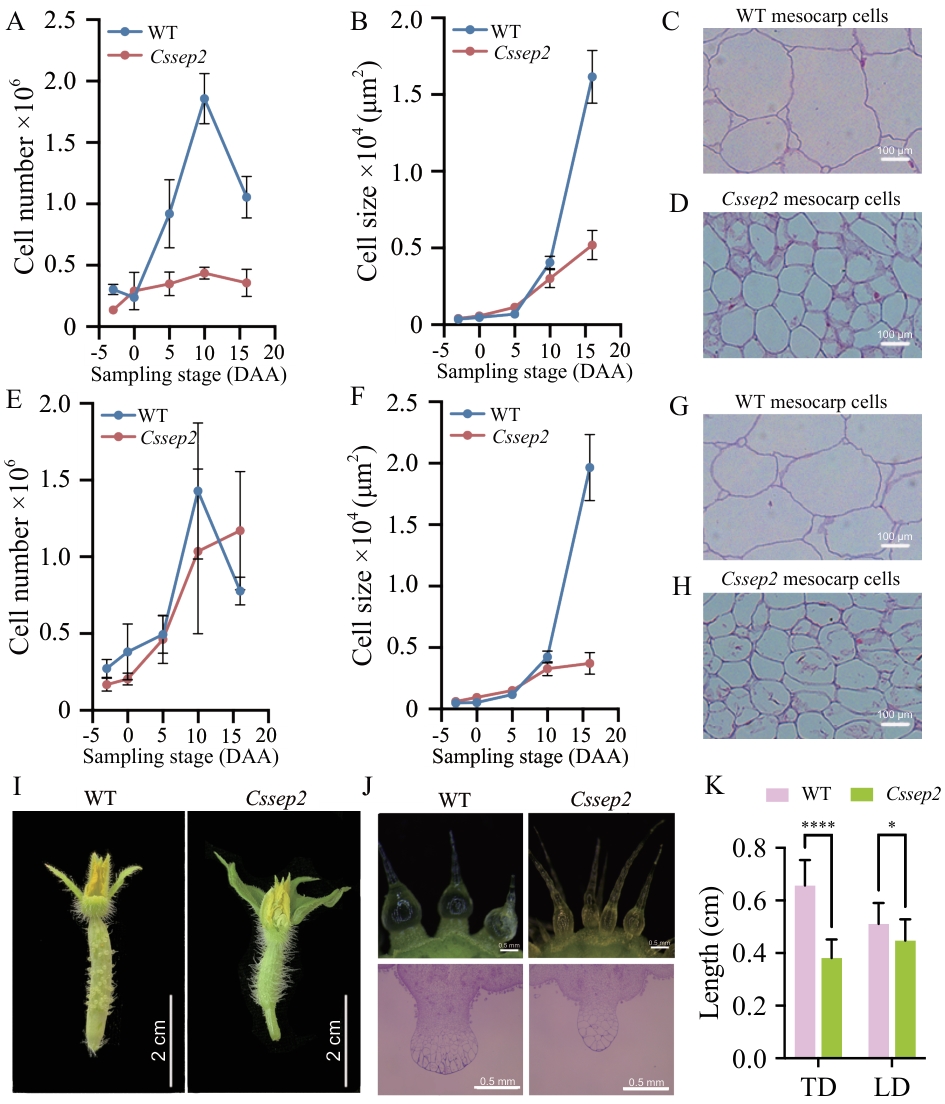
Fig. 3 Cellular dissection of the fruit and wart phenotype of the Cssep2 mutantsA, B: Cell number and size of fruit mesocarp along the longitudinal axis. Values are means ± SD, n = 540. C, D: Longitudinal sections of the fruit mesocarp at 16 DAA. E, F: Cell number and size of fruit mesocarp along the transversal axis. Values are means ± SD, n = 540. G, H: Transversal sections of the fruit mesocarp at 16 DAA. I: Fruit spine of WT and Cssep2 mutants at -3 DAA. J: Macroscopic observation of the tubercule at -3 DAA. K: Measurement of the spine base of WT and Cssep2 mutants at -3 DAA. TD: Traverse diameter, LD: longitudinal diameter; * P < 0.05; **** P < 0.000 1. Values are means ± SD, n = 18. The same below
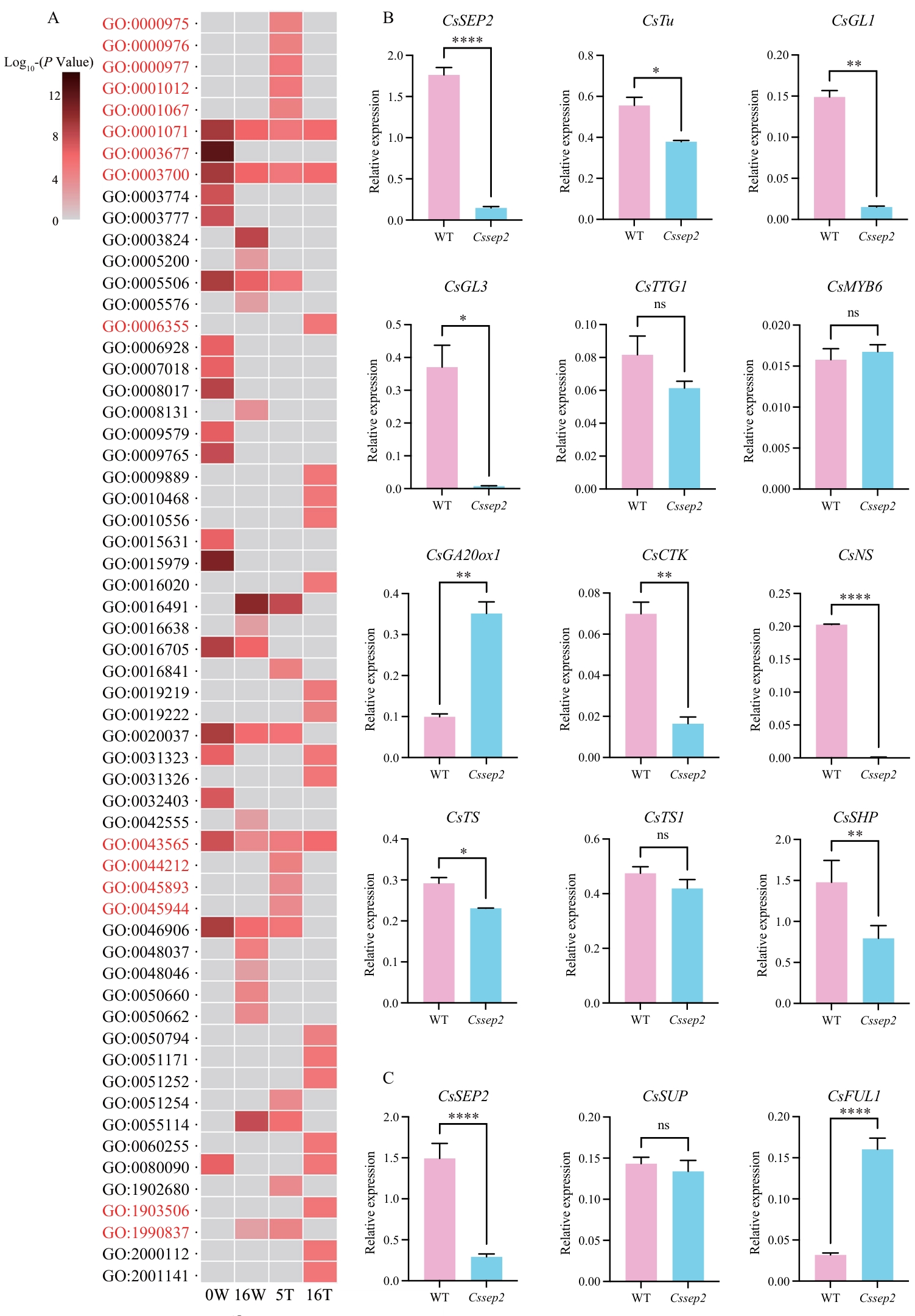
Fig. 4 GO enrichment and RT-qPCR analysis of differentially expressed genesA: GO enrichment analysis of DEGs at 5T (placenta at 5 DAA), 16T (placenta at 16 DAA), 0W (epicarp at 0 DAA) and 16W (epicarp at 16 DAA). GO terms related with DNA binding and regulation of transcription are highlighted with red. B: Expression analysis of genes in the epicarp tissues of WT and Cssep2 mutants. C: Expression analysis of genes in the placenta tissues of WT and Cssep2 mutants; **** P < 0.000 1; ** P < 0.01; * P < 0.05; ns: not significant difference
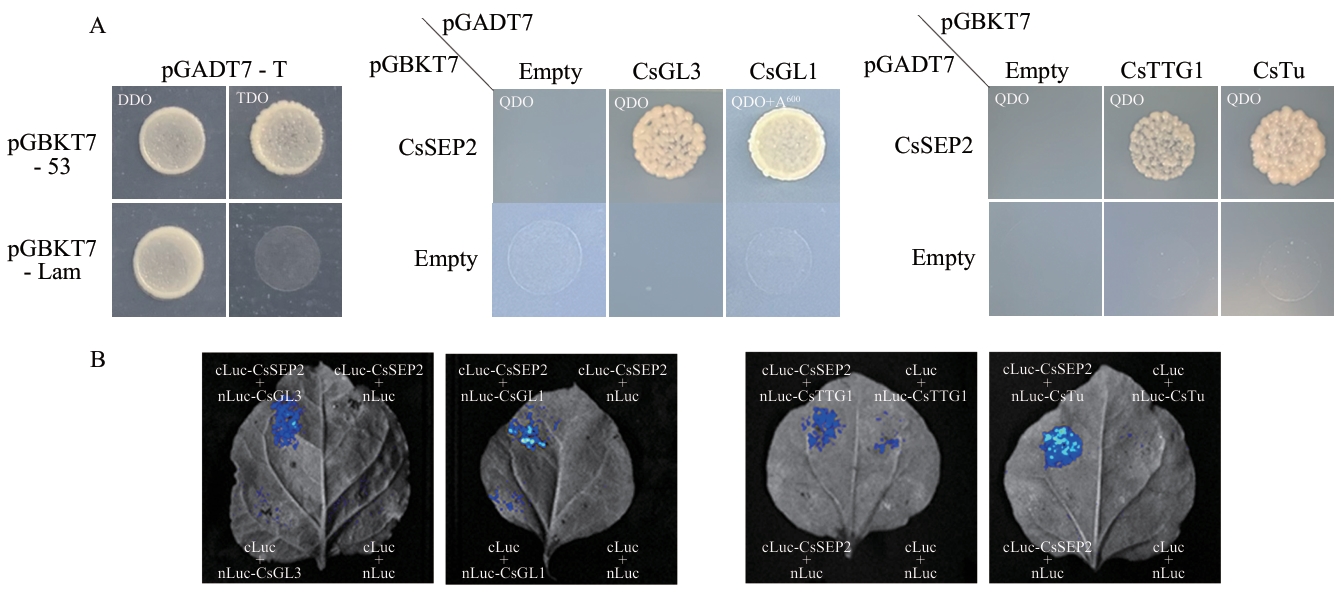
Fig. 5 Identification of proteins interacting with CsSEP2A: Yeast two-hybrid assay results. QDO: Quadruple dropout, SD/-Leu-Trp-His-Ade. A600: Addition of Aureobasidin A with a final concentration of 600 ng/mL. B: Firefly luciferase complementation imaging analysis
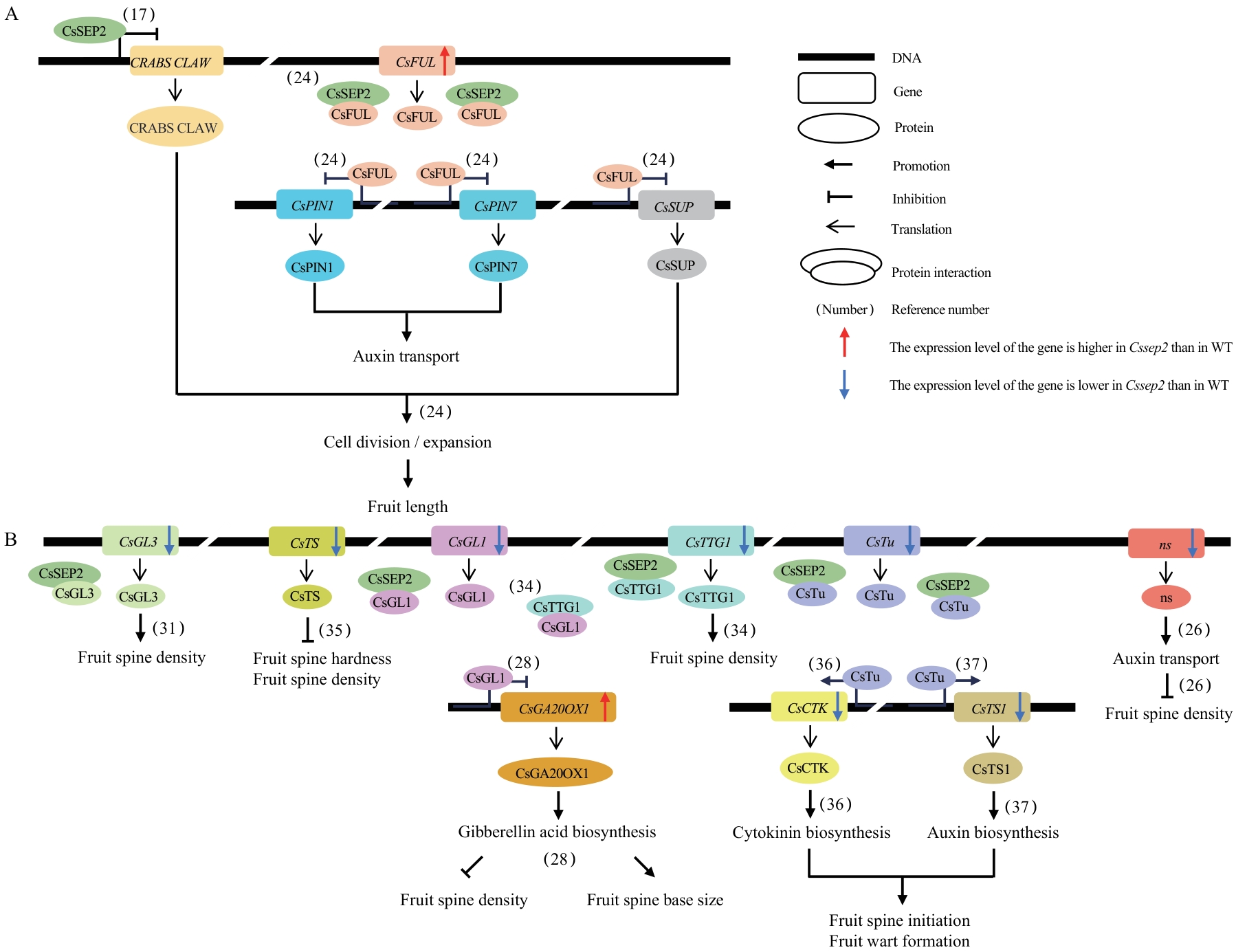
Fig. 6 Molecular network of fruit-related traits regulated by CsSEP2A, B: Schematic diagram of the CsSEP2-invovled molecular network of fruit development (A) and warty traits (B). The captions of the figure are in the upper right corner. CsSEP2 inhibits the lengths of cucumber fruits by suppressing the expression of CRABS CLAW[17]. CsFUL regulates fruit length by inhibiting the expressions of CsPIN1, CsPIN7 and CsSUP, and can also form protein complex with CsSEP2 to exert its functions[21]. Notably, the expressions of both CsFUL and CsSUP changed significantly in Cssep2. Compared to WT, the expressions of CsGL3[22], CsTS[23], CsGL1[24], CsTTG1[25], CsTu[26-27], and ns[28] were lower in Cssep2 mutants. These genes have been confirmed to be involved in the regulation of the formation and development of fruit spines and warts. Among them, ns encodes an auxin transporter and inhibits the density of fruit spines[28]. CsGL1 can not only inhibit the expression of CsGA20OX1, affecting the synthesis of GA and altering the density of fruit spines and the size of the fruit spine base[24], but also interacts with CsTTG1 to form a complex to regulate the density of fruit spines[25]. CsTu promotes the expressions of CsCTK[26] and CsTS1[27], altering the synthesis of cytokinin and auxin and facilitating the initiation of fruit spines and the formation of fruit warts. In Cssep2 mutants, the expressions of CsGA20OX1, CsCTK, and CsTS1 also changed to varying degrees. In addition, this study further revealed that CsSEP2 interacts with CsGL3, CsGL1, CsTTG,1 and CsTu
| [1] | Bowman JL, Moyroud E. Reflections on the ABC model of flower development [J]. Plant Cell, 2024, 36(5): 1334-1357. |
| [2] | Lin ZY, Zhu GF, Lu CQ, et al. Functional conservation and divergence of SEPALLATA-like genes in floral development in Cymbidium sinense [J]. Front Plant Sci, 2023, 14: 1209834. |
| [3] | Ditta G, Pinyopich A, Robles P, et al. The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity [J]. Curr Biol, 2004, 14(21): 1935-1940. |
| [4] | Jack T. Relearning our ABCs: new twists on an old model [J]. Trends Plant Sci, 2001, 6(7): 310-316. |
| [5] | Tan XM, Li YR, Song MR, et al. The molecular mechanism of interaction between SEPALLATA3 and APETALA1 in Arabidopsis thaliana [J]. Plant Direct, 2025, 9(4): e70052. |
| [6] | Hugouvieux V, Blanc-Mathieu R, Janeau A, et al. SEPALLATA-driven MADS transcription factor tetramerization is required for inner whorl floral organ development [J]. Plant Cell, 2024, 36(9): 3435-3450. |
| [7] | Gan ZC, Wu XX, Biahomba SAM, et al. Genome-wide identification, evolution, and expression characterization of the pepper (Capsicum spp.) MADS-box gene family [J]. Genes, 2022, 13(11): 2047. |
| [8] | Cui ZB, Wang XW, Dai YD, et al. Transcription factor OsNF-YC1 regulates grain size by coordinating the transcriptional activation of OsMADS1 in Oryza sativa L [J]. Plant J, 2024, 119(3): 1465-1480. |
| [9] | Ampomah-Dwamena C, Morris BA, Sutherland P, et al. Down-regulation of TM29, a tomato SEPALLATA homolog, causes parthenocarpic fruit development and floral reversion [J]. Plant Physiol, 2002, 130(2): 605-617. |
| [10] | Vallarino JG, Merchante C, Sánchez-Sevilla JF, et al. Characterizing the involvement of FaMADS9 in the regulation of strawberry fruit receptacle development [J]. Plant Biotechnol J, 2020, 18(4): 929-943. |
| [11] | Ireland HS, Yao JL, Tomes S, et al. Apple SEPALLATA1/2-like genes control fruit flesh development and ripening [J]. Plant J, 2013, 73(6): 1044-1056. |
| [12] | Upadhyay RK, Tucker ML, Mattoo AK. Ethylene and RIPENING INHIBITOR modulate expression of SlHSP17.7A, B class I small heat shock protein genes during tomato fruit ripening [J]. Front Plant Sci, 2020, 11: 975. |
| [13] | Ito Y, Kitagawa M, Ihashi N, et al. DNA-binding specificity, transcriptional activation potential, and the rin mutation effect for the tomato fruit-ripening regulator RIN [J]. Plant J, 2008, 55(2): 212-223. |
| [14] | Zhou Y, Hu LF, Song JB, et al. Isolation and characterization of a MADS-box gene in cucumber (Cucumis sativus L.) that affects flowering time and leaf morphology in transgenic Arabidopsis [J]. Biotechnol Biotechnol Equip, 2019, 33(1): 54-63. |
| [15] | Cheng YW, He P, Jiang LW, et al. Identification and characterization of a SEPALLATA-like MADS-box gene from cucumber (Cucumis sativus L.) [J]. Not Bot Horti Agrobo, 2019, 47(4): 1168-1177. |
| [16] | Wang X, Gao DL, Sun JJ, et al. An exon skipping in a SEPALLATA-Like gene is associated with perturbed floral and fruits development in cucumber [J]. J Integr Plant Biol, 2016, 58(9): 766-771. |
| [17] | Song WY, Xie Y, Liu B, et al. Single nucleotide polymorphisms in SEPALLATA 2 underlie fruit length variation in cucurbits [J]. Plant Cell, 2024, 36(10): 4607-4621. |
| [18] | Xin TX, Tian HJ, Ma YL, et al. Targeted creating new mutants with compact plant architecture using CRISPR/Cas9 genome editing by an optimized genetic transformation procedure in cucurbit plants [J]. Hortic Res, 2022, 9: uhab086. |
| [19] | Kanehisa M, Furumichi M, Sato Y, et al. KEGG: biological systems database as a model of the real world [J]. Nucleic Acids Res, 2025, 53(D1): D672-D677. |
| [20] | Wang LP, Yu G, Macho AP, et al. Split-luciferase complementation imaging assay to study protein-protein interactions in Nicotiana benthamiana [J]. Bio Protoc, 2021, 11(23): e4237. |
| [21] | Zhao JY, Jiang L, Che G, et al. A functional allele of CsFUL1 regulates fruit length through repressing CsSUP and inhibiting auxin transport in cucumber [J]. Plant Cell, 2019, 31(6): 1289-1307. |
| [22] | Pan YP, Bo KL, Cheng ZH, et al. The loss-of-function GLABROUS 3 mutation in cucumber is due to LTR-retrotransposon insertion in a class IV HD-ZIP transcription factor gene CsGL3 that is epistatic over CsGL1 [J]. BMC Plant Biol, 2015, 15: 302. |
| [23] | Guo CL, Yang XQ, Wang YL, et al. Identification and mapping of ts (tender spines), a gene involved in soft spine development in Cucumis sativus [J]. Theor Appl Genet, 2018, 131(1): 1-12. |
| [24] | Qiang Li CC. The identification of Cucumis sativus Glabrous 1 (CsGL1) required for the formation of trichomes uncovers a novel function for the homeodomain-leucine zipper I gene [J]. J Exp Bot, 2015, 66(9): 2515-2526. |
| [25] | Chunhua Chen SY. The WD-repeat protein CsTTG1 regulates fruit wart formation through interaction with the homeodomain-leucine zipper I protein mict [J]. Plant Physiol, 2016, 171(2): 1156-1168. |
| [26] | Yang XQ, Zhang WW, He HL, et al. Tuberculate fruit gene Tu encodes a C2H2 zinc finger protein that is required for the warty fruit phenotype in cucumber (Cucumis sativus L.) [J]. Plant J, 2014, 78(6): 1034-1046. |
| [27] | Yang S, Wen CL, Liu B, et al. A CsTu-TS1 regulatory module promotes fruit tubercule formation in cucumber [J]. Plant Biotechnol J, 2019, 17(1): 289-301. |
| [28] | Xie Q, Liu PN, Shi LX, et al. Combined fine mapping, genetic diversity, and transcriptome profiling reveals that the auxin transporter gene ns plays an important role in cucumber fruit spine development [J]. Theor Appl Genet, 2018, 131(6): 1239-1252. |
| [29] | Zhang ZH, Zou WH, Lin PX, et al. Evolution and function of MADS-box transcription factors in plants [J]. Int J Mol Sci, 2024, 25(24): 13278. |
| [30] | Lai XL, Stigliani A, Lucas J, et al. Genome-wide binding of SEPALLATA3 and AGAMOUS complexes determined by sequential DNA-affinity purification sequencing [J]. Nucleic Acids Res, 2020, 48(17): 9637-9648. |
| [31] | Lai XL, Vega-Léon R, Hugouvieux V, et al. The intervening domain is required for DNA-binding and functional identity of plant MADS transcription factors [J]. Nat Commun, 2021, 12(1): 4760. |
| [32] | Yang S, Cai YL, Liu XW, et al. A CsMYB6-CsTRY module regulates fruit trichome initiation in cucumber [J]. J Exp Bot, 2018, 69(8): 1887-1902. |
| [33] | Chen CH, Liu ML, Jiang L, et al. Transcriptome profiling reveals roles of meristem regulators and polarity genes during fruit trichome development in cucumber (Cucumis sativus L.) [J]. J Exp Bot, 2014, 65(17): 4943-4958. |
| [34] | Zhao JL, Pan JS, Guan Y, et al. Micro-trichome as a class I homeodomain-leucine zipper gene regulates multicellular trichome development in Cucumis sativus [J]. J Integr Plant Biol, 2015, 57(11): 925-935. |
| [35] | Cui JY, Miao H, Ding LH, et al. A new glabrous gene (csgl3) identified in trichome development in cucumber (Cucumis sativus L.) [J]. PLoS One, 2016, 11(2): e0148422. |
| [36] | Wang YL, Nie JT, Chen HM, et al. Identification and mapping of Tril, a homeodomain-leucine zipper gene involved in multicellular trichome initiation in Cucumis sativus [J]. Theor Appl Genet, 2016, 129(2): 305-316. |
| [37] | Zhang HY, Wang LN, Zheng SS, et al. A fragment substitution in the promoter of CsHDZIV11/CsGL3 is responsible for fruit spine density in cucumber (Cucumis sativus L.) [J]. Theor Appl Genet, 2016, 129(7): 1289-1301. |
| [38] | Ferrándiz C, Fourquin C. Role of the FUL-SHP network in the evolution of fruit morphology and function [J]. J Exp Bot, 2014, 65(16): 4505-4513. |
| [39] | Ferrándiz C, Liljegren SJ, Yanofsky MF. Negative regulation of the SHATTERPROOF genes by FRUITFULL during Arabidopsis fruit development [J]. Science, 2000, 289(5478): 436-438. |
| [40] | Cheng ZH, Zhuo SB, Liu XF, et al. The MADS-box gene CsSHP participates in fruit maturation and floral organ development in cucumber [J]. Front Plant Sci, 2019, 10: 1781. |
| [41] | Wang LN, Cao CX, Zheng SS, et al. Transcriptomic analysis of short-fruit 1 (sf1) reveals new insights into the variation of fruit-related traits in Cucumis sativus [J]. Sci Rep, 2017, 7(1): 2950. |
| [42] | Wang J, Tian PP, Sun JJ, et al. CsMYC2 is involved in the regulation of phenylpropanoid biosynthesis induced by trypsin in cucumber (Cucumis sativus) during storage [J]. Plant Physiol Biochem, 2023, 196: 65-74. |
| [1] | YIN Yue, QIN Xiao-ya, MI Jia, AN Wei, HE Jun, ZHANG Feng-feng. Identification of FBN Gene Family and Its Relationship with Carotenoids Metabolism in Lyciumbarbarum [J]. Biotechnology Bulletin, 2026, 42(3): 338-348. |
| [2] | LI Tian-yuan, QI Xin-liang, LIU Shan, ZHANG Jian-cheng, WANG Peng-fei, ZHANG Shuai, JIA Lu-ting, MU Xiao-peng. Identification of Cerasus humilisSPL Gene Family and Expression Analysis during Fruit Development [J]. Biotechnology Bulletin, 2026, 42(3): 362-373. |
| [3] | LIU Na, ZENG Bao-zhen, JIA Zhao-xing, ZHU Ying-fang. Advances in Epigenetic Regulation of Tomato Fruit Development and Ripening [J]. Biotechnology Bulletin, 2026, 42(3): 37-47. |
| [4] | HU Qiu-ling, CHEN Ling, HUANG Jia-yi, ZHAO Zi-qiao, PAN Lu-yi, LIU Hui-li, LIU Tai-bo. Advances in the Regulation of Fruit Development by Polyamines [J]. Biotechnology Bulletin, 2026, 42(3): 203-212. |
| [5] | MA Shi-jie, LI Zheng, LI Wei, GUO Yang-dong, ZHANG Na. Research Progress in Light Signaling Regulation of Fruit Development in Horticultural Crops [J]. Biotechnology Bulletin, 2026, 42(3): 5-18. |
| [6] | ZHANG Gao-xiang, WU Yu-bi, GUO Ya-jing, JI Wei, YANG Zhong-yi. Identification and Expression Analysis of WD40 Gene Family in Grape [J]. Biotechnology Bulletin, 2026, 42(3): 324-337. |
| [7] | LI Cheng-quan, SHI Qing-hua, YANG Xiao-yu. microRNA-based Regulatory Network for Fruit Development of Horticultural Crops: From Molecular Mechanism to Germplasm Innovation [J]. Biotechnology Bulletin, 2026, 42(3): 19-36. |
| [8] | LIU Jia-li, SONG Jing-rong, ZHAO Wen-yu, ZHANG Xin-yuan, ZHAO Zi-yang, CAO Yi-bo, ZHANG Ling-yun. Identification of the R2R3-MYB Gene and Expression Analysis of Flavonoid Regulatory Genes in Blueberry [J]. Biotechnology Bulletin, 2025, 41(9): 124-138. |
| [9] | WANG Bin, LI Jian-rong, ZHAN Zhao-xia, YUAN Xiao. Cloning of CsGR-RBP3 and Its Functional Roles in Cold Tolerance of Harvested Cucumber [J]. Biotechnology Bulletin, 2025, 41(6): 155-166. |
| [10] | LUO Si-fang, ZHANG Zu-ming, XIE Li-fang, GUO Zi-jing, CHEN Zhao-xing, YANG Yue-hua, YAN Xiang, ZHANG Hong-ming. Genome-wide Identification of GATA Gene Family of Jindou Kumquat (Fortunella hindsii) and Their Expression Analysis in Fruit Development [J]. Biotechnology Bulletin, 2025, 41(5): 218-230. |
| [11] | FAN Zong-qiang, FENG Jing-han, ZHENG Li-xue, WANG Shuo, PENG Xiang-qian, CHEN Fang. Study on the Control and Induced Resistance in Cucumber with Bacillus subtilis B579 against Cucumber Fusarium Wilt [J]. Biotechnology Bulletin, 2024, 40(7): 226-234. |
| [12] | XIAO Ya-ru, JIA Ting-ting, LUO Dan, WU Zhe, LI Li-xia. Cloning and Expression Analysis of CsERF025L Transcription Factor in Cucumber [J]. Biotechnology Bulletin, 2024, 40(4): 159-166. |
| [13] | YANG Chong, CHENG Sha-sha, AI Chang-feng, ZHAO Xuan, LIU Meng-jun. Identification of ABF/AREB Gene Family and Their Expression Analysis in Jujube Fruit [J]. Biotechnology Bulletin, 2024, 40(11): 184-191. |
| [14] | CHU Rui, LI Zhao-xuan, ZHANG Xue-qing, YANG Dong-ya, CAO Hang-hang, ZHANG Xue-yan. Screening and Identification of Antagonistic Bacillus spp. Against Cucumber Fusarium wilt and Its Biocontrol Effect [J]. Biotechnology Bulletin, 2023, 39(8): 262-271. |
| [15] | XIE Yang, XING Yu-meng, ZHOU Guo-yan, LIU Mei-yan, YIN Shan-shan, YAN Li-ying. Transcriptome Analysis of Diploid and Autotetraploid in Cucumber Fruit [J]. Biotechnology Bulletin, 2023, 39(3): 152-162. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||