








生物技术通报 ›› 2024, Vol. 40 ›› Issue (7): 273-284.doi: 10.13560/j.cnki.biotech.bull.1985.2024-0068
收稿日期:2024-01-16
出版日期:2024-07-26
发布日期:2024-07-30
通讯作者:
彭涛,男,博士,副教授,研究方向:苔藓植物学、湿地生态学及生物多样性保护;E-mail: pengtao@gznu.edu.cn;作者简介:黄丹,女,硕士,研究方向:生态学;E-mail: 1093617061@qq.com
基金资助:
HUANG Dan1( ), JIANG Shan2(
), JIANG Shan2( ), PENG Tao1(
), PENG Tao1( )
)
Received:2024-01-16
Published:2024-07-26
Online:2024-07-30
摘要:
【目的】 细胞色素P450单氧化酶98(Cytochrome P450 monooxygenase 98, CYP98)是苯丙烷途径中的关键限速酶,拟探究其在角苔植物中是否参与苯丙烷途径中生物合成及抗病功能,为今后研究早期陆生植物的进化和适应逆境胁迫的生理机制提供参考。【方法】 利用RACE技术从褐角苔中克隆 FfCYP98 cDNA全长序列,对其进行生物信息学和亚细胞定位分析,并通过构建过表达载体转化拟南芥突变体 FfCYP98-OE1 进行功能验证。【结果】 FfCYP98 cDNA序列开放阅读框1 305 bp,与苔藓植物芽孢角苔和小立碗藓CYP98在进化关系上最近,亚细胞定位结果显示FfCYP98定位于细胞核和细胞质。过表达FfCYP98基因后发现FfCYP98-OE1植株总酚、总黄酮和苯丙烷合成途径中C4H、4CL和C3H的表达量均显著高于拟南芥WT植株。另外,用灰霉菌侵染褐角苔和拟南芥FfCYP98-OE1植株后发现FfCYP98表达量显著上调,FfCYP98-OE1植株枯死的速度明显比WT植株慢,且拟南芥FfCYP98-OE1植株中总酚、总黄酮和苯丙烷合成途径中C4H、HCT、C3H和CAD四个基因的表达量均显著高于WT植株。【结论】 褐角苔FfCYP98可能通过调控苯丙烷途径合成相关基因的表达,参与了苯丙烷途径中酚类和黄酮类化合物的生物合成,并与植物的抗病性有关。
黄丹, 姜山, 彭涛. 褐角苔FfCYP98基因克隆及其功能分析[J]. 生物技术通报, 2024, 40(7): 273-284.
HUANG Dan, JIANG Shan, PENG Tao. Cloning of FfCYP98 Gene and Its Functional Analysis in Folioceros fuciformis[J]. Biotechnology Bulletin, 2024, 40(7): 273-284.
| 引物名称Primer name | 引物序列Primer sequence(5'-3') |
|---|---|
| JB-F | ATGAGGAGTTTGCGAAGCA |
| JB-R | CCTTCCCCTCGTCGTCCAC |
| 3'RACE 特异性引物F1 | AGGGGCCGCAGTCTCATTTCGT |
| 3'RACE 特异性引物F2 | CTCCTGGGGCTGCAGAAGCAGT |
| 5'RACE 特异性引物R1 | AACGCGAGGCGCGTAATGTTGT |
| 5'RACE 特异性引物R2 | GCCGAGAGGTACGTCTTGGGGTT |
| FfCYP98-F | TCTAGAATGGGAGCAACTCTGAACGT |
| FfCYP98-R | GGATCCGACGGTTTTGGCATACAAGG |
| HACtin-F | ACGCCTGAACATCTCCTGAA |
| HACtin-R | CGTGCAGAACAAGAACTCGT |
| qpcr-F | CGCAGTCTCATTTCGTGGAC |
| qpcr-R | TCCTTCTGCACCTTCCTCTG |
| 跨载体-F | ccaaccacgtcttcaaagca |
| 跨载体-R | atccagactgaatgcccaca |
| ACtion2-F | GGAAGGATCTGTACGGTAAC |
| ACtion2-R | GGACCTGCCTCATCTTACT |
| Action-F | TTCAATGTCCCTGCCATGTATG |
| Action-R | AATACCGGTTGTACGACCAC |
| PAL-F | TCTGCGAAAAGGATTTACTCAAAGTT |
| PAL-R | CTACAAGGATCATCCGCGTATGT |
| C4H-F | CCTCCAGGACAGTCTAAAGTGGATACTA |
| C4H-R | CGATTATGGAGTGGTTAAGGATGTG |
| HCT-F | TCTTTATGGGACCTGGTGGAATT |
| HCT-R | GATAAGCTGCCATCATTAGTAGGACTTG |
| 4CL-F | TCAACCCGGTGAGATTTGTA |
| 4CL-R | TCGTCATCGATCAATCCAAT |
| C3H-F | AAGTCTAGTGGAGCGAAACAGCATT |
| C3H-R | TCCTCACTAAGATCATACTGATCCTTCA |
| CAD-F | TGCGATTGGATCCGATGGAA |
| CAD-R | GCTTCCCTGCCTCAGTCATA |
表1 本研究所用引物序列
Table 1 Primer sequences used in the study
| 引物名称Primer name | 引物序列Primer sequence(5'-3') |
|---|---|
| JB-F | ATGAGGAGTTTGCGAAGCA |
| JB-R | CCTTCCCCTCGTCGTCCAC |
| 3'RACE 特异性引物F1 | AGGGGCCGCAGTCTCATTTCGT |
| 3'RACE 特异性引物F2 | CTCCTGGGGCTGCAGAAGCAGT |
| 5'RACE 特异性引物R1 | AACGCGAGGCGCGTAATGTTGT |
| 5'RACE 特异性引物R2 | GCCGAGAGGTACGTCTTGGGGTT |
| FfCYP98-F | TCTAGAATGGGAGCAACTCTGAACGT |
| FfCYP98-R | GGATCCGACGGTTTTGGCATACAAGG |
| HACtin-F | ACGCCTGAACATCTCCTGAA |
| HACtin-R | CGTGCAGAACAAGAACTCGT |
| qpcr-F | CGCAGTCTCATTTCGTGGAC |
| qpcr-R | TCCTTCTGCACCTTCCTCTG |
| 跨载体-F | ccaaccacgtcttcaaagca |
| 跨载体-R | atccagactgaatgcccaca |
| ACtion2-F | GGAAGGATCTGTACGGTAAC |
| ACtion2-R | GGACCTGCCTCATCTTACT |
| Action-F | TTCAATGTCCCTGCCATGTATG |
| Action-R | AATACCGGTTGTACGACCAC |
| PAL-F | TCTGCGAAAAGGATTTACTCAAAGTT |
| PAL-R | CTACAAGGATCATCCGCGTATGT |
| C4H-F | CCTCCAGGACAGTCTAAAGTGGATACTA |
| C4H-R | CGATTATGGAGTGGTTAAGGATGTG |
| HCT-F | TCTTTATGGGACCTGGTGGAATT |
| HCT-R | GATAAGCTGCCATCATTAGTAGGACTTG |
| 4CL-F | TCAACCCGGTGAGATTTGTA |
| 4CL-R | TCGTCATCGATCAATCCAAT |
| C3H-F | AAGTCTAGTGGAGCGAAACAGCATT |
| C3H-R | TCCTCACTAAGATCATACTGATCCTTCA |
| CAD-F | TGCGATTGGATCCGATGGAA |
| CAD-R | GCTTCCCTGCCTCAGTCATA |

图2 FfCYP98的生物信息学分析 A:FfCYP98理化性质分析和二级结构预测;B:FfCYP98的三维结构预测;C:FfCYP98氨基酸序亲水性与疏水性分析;D:FfCYP98蛋白质跨膜结构域分析;E:FfCYP98 的系统发生树
Fig. 2 Bioinformatics analysis of the FfCYP98 A: Physicochemical characterization and secondary structure prediction of FfCYP98. B: Tertiary structure prediction of FfCYP98. C: Analysis of hydrophilicity and hydrophobicity of FfCYP98 amino acid sequences. D: Analysis of the transmembrane structural domains of FfCYP98. E: Phylogenetic tree of FfCYP98

图3 FfCYP98 蛋白的亚细胞定位 星号表示细胞核,箭头表示细胞质
Fig. 3 Subcellular localization of the protein FfCYP98 The asterisk indicates the nucleus, and the arrow indicates the cytoplasm

图4 转基因拟南芥鉴定和转基因拟南芥中总酚、总黄酮和苯丙烷合成途径基因的相对表达量 A:抗性筛选拟南芥;B:分子鉴定转基因拟南芥,M:Marker = 15 000 bp,1:空白对照,2:WT;C:FfCYP98基因在转基因拟南芥中的表达量(***P< 0.001,****P< 0.0001);D:转基因拟南芥总酚和总黄酮检测;E:转基因拟南芥苯丙烷合成途径基因的表达分析
Fig. 4 Identification of transgenic A. thaliana and relative expressions of total phenols, total flavonoids and phenylpropane synthesis pathway genes in transgenic A. thaliana A: Resistance screening for Arabidopsis thaliana. B: Identification of transgenic A. thaliana, M: Marker = 15 000 bp, 1: Control, 2: WT. C: Expression of the FfCYP98 gene intransgenic A. thaliana(***P< 0.001,****P< 0.0001). D: Detection of total phenols and total flavonoids in transgenic A. thaliana. E: Expression analysis of transgenic A. thaliana phenylpropane synthesis pathway genes
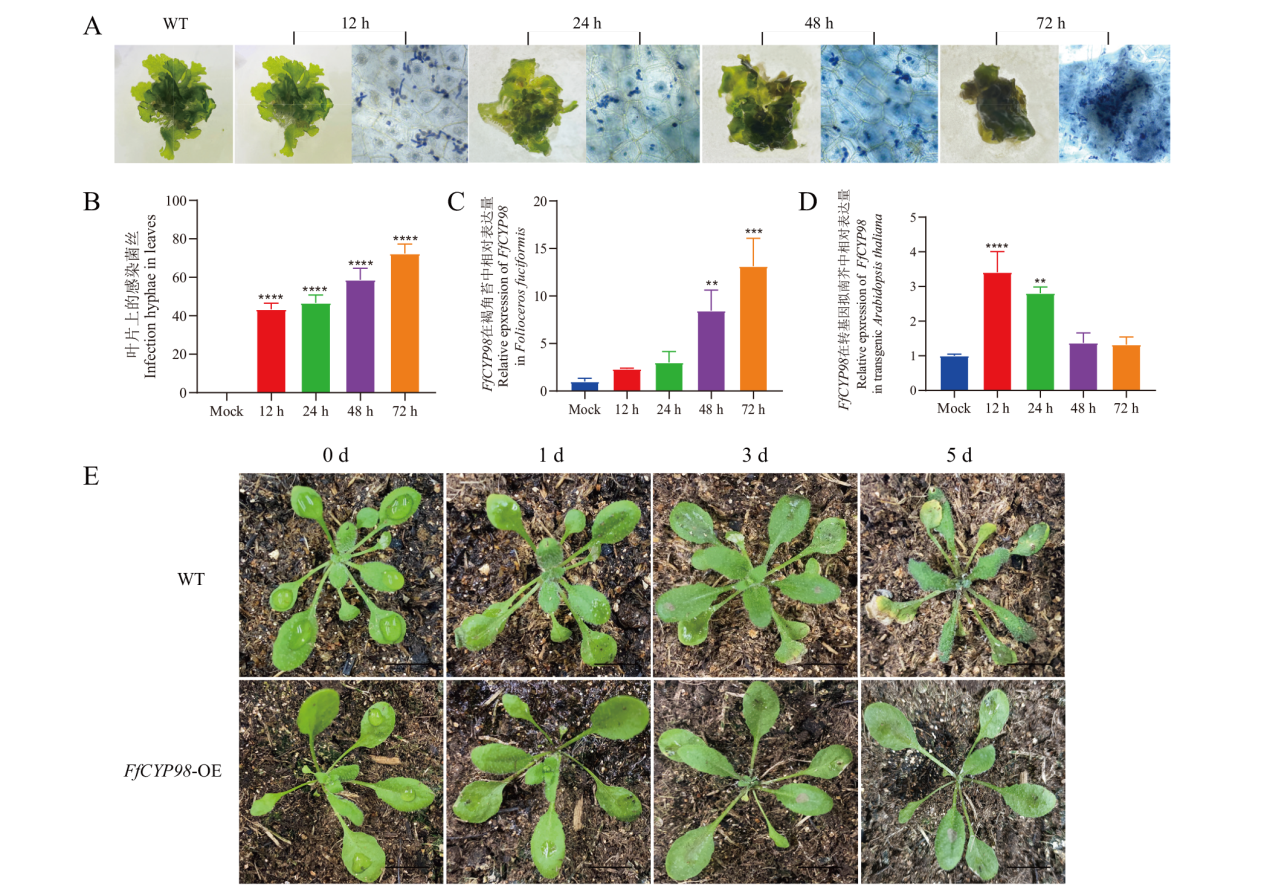
图5 灰霉菌在褐角苔中的感染行为和灰霉菌侵染下FfCYP98在转基因拟南芥中的表达特性 A:灰霉菌侵染褐角苔,Bar=100 μm; B:灰霉菌侵入菌丝率的比较;C: 不同时间灰霉菌侵染褐角苔后FfCYP98表达水平;D:灰霉菌感染转基因拟南芥形态观察,bar= 1 cm;E:不同时间灰霉菌侵染转基因拟南芥后FfCYP98表达水平。**P< 0.01,***P< 0.001,****P< 0.0001
Fig. 5 Infected behavior of B. cinerea on F. fuciformis and characterization of FfCYP98 expression in transgenic A. thaliana under B. cinerea infestation A: B. cinerea infests Folioceros fuciformis, bar=100 μm. B: Comparison of the invasion rate of hyphae in F. fuciformis. C: Expression of FfCYP98 at different times of F. fuciformis under the stress of B. cinerea. D: Observation of transgenic A. thaliana infection with B. cinerea, bar= 1 cm. E: Expressions of FfCYP98 at different times of transgenic A. thaliana under the stress of B. cinerea, **P< 0.01, ***P< 0.001, ****P< 0.0001
| [1] | Kenrick P, Wellman CH, Schneider H, et al. A timeline for terrestrialization: consequences for the carbon cycle in the Palaeozoic[J]. Philos Trans R Soc Lond B Biol Sci, 2012, 367(1588): 519-536. |
| [2] |
Yin RH, Ulm R. How plants cope with UV-B: from perception to response[J]. Curr Opin Plant Biol, 2017, 37: 42-48.
doi: S1369-5266(16)30228-X pmid: 28411583 |
| [3] | Vanholme R, Storme V, Vanholme B, et al. A systems biology view of responses to lignin biosynthesis perturbations in Arabidopsis[J]. Plant Cell, 2012, 24(9): 3506-3529. |
| [4] |
Davies KM, Jibran R, Zhou YF, et al. The evolution of flavonoid biosynthesis: a bryophyte perspective[J]. Front Plant Sci, 2020, 11: 7.
doi: 10.3389/fpls.2020.00007 pmid: 32117358 |
| [5] | Alber AV, Renault H, Basilio-Lopes A, et al. Evolution of coumaroyl conjugate 3-hydroxylases in land plants: lignin biosynthesis and defense[J]. Plant J, 2019, 99(5): 924-936. |
| [6] | Kono Y, Kashine S, Yoneyama T, et al. Iron chelation by chlorogenic acid as a natural antioxidant[J]. Biosci Biotechnol Biochem, 1998, 62(1): 22-27. |
| [7] | Clé C, Hill LM, Niggeweg R, et al. Modulation of chlorogenic acid biosynthesis in Solanum lycopersicum; consequences for phenolic accumulation and UV-tolerance[J]. Phytochemistry, 2008, 69(11): 2149-2156. |
| [8] |
Barbehenn R, Dukatz C, Holt C, et al. Feeding on poplar leaves by caterpillars potentiates foliar peroxidase action in their guts and increases plant resistance[J]. Oecologia, 2010, 164(4): 993-1004.
doi: 10.1007/s00442-010-1733-y pmid: 20680646 |
| [9] |
Escamilla-Treviño LL, Shen H, Hernandez T, et al. Early lignin pathway enzymes and routes to chlorogenic acid in switchgrass(Panicum virgatum L.)[J]. Plant Mol Biol, 2014, 84(4-5): 565-576.
doi: 10.1007/s11103-013-0152-y pmid: 24190737 |
| [10] |
Vanholme R, Cesarino I, Rataj K, et al. Caffeoyl shikimate esterase(CSE)is an enzyme in the lignin biosynthetic pathway in Arabidopsis[J]. Science, 2013, 341(6150): 1103-1106.
doi: 10.1126/science.1241602 pmid: 23950498 |
| [11] | 王雪霞, 薛永常, 赵文超. 木质素生物合成中C3H/HCT的研究进展[J]. 生命的化学, 2008, 28(5): 650-653. |
| Wang XX, Xue YC, Zhao WC. The research progress of C3H/HCT in lignin biosynthesis[J]. Chem Life, 2008, 28(5): 650-653. | |
| [12] |
Renault H, De Marothy M, Jonasson G, et al. Gene duplication leads to altered membrane topology of a cytochrome P450 enzyme in seed plants[J]. Mol Biol Evol, 2017, 34(8): 2041-2056.
doi: 10.1093/molbev/msx160 pmid: 28505373 |
| [13] | 胡人亮. 漫话苔藓[J]. 植物杂志, 1978(4): 41-43. |
| Hu RL. Manga Moss[J]. Plants, 1978(4): 41-43. | |
| [14] |
Basile A, Giordano S, López-Sáez JA, et al. Antibacterial activity of pure flavonoids isolated from mosses[J]. Phytochemistry, 1999, 52(8): 1479-1482.
pmid: 10647220 |
| [15] | Umezawa T. Diversity in lignan biosynthesis[J]. Phytochem Rev, 2003, 2(3): 371-390. |
| [16] | Xu ZY, Zhang DD, Hu J, et al. Comparative genome analysis of lignin biosynthesis gene families across the plant kingdom[J]. BMC Bioinformatics, 2009, 10(Suppl 11): S3. |
| [17] | Jiang S, Tian X, Huang XL, et al. Physcomitrium patens CAD1 has distinct roles in growth and resistance to biotic stress[J]. BMC Plant Biol, 2022, 22(1): 518. |
| [18] | Garcia C, Sérgio C, Villarreal JC, et al. The HornwortsDendrocerosNees andMegacerosCampb. in São tomé e Príncipe(africa, gulf of Guinea)with the description of Dendroceros paivaesp. nov[J]. Cryptogam Bryol, 2012, 33(1): 3-21. |
| [19] | Renzaglia KS, Villarreal JC, Duff RJ, et al. New insights into morphology, anatomy, and systematics of hornworts[M]//Shaw AJ, ed. Bryophyte Biology. Cambridge: Cambridge University Press, 2010: 139-172. |
| [20] | Qiu YL, Li LB, Wang B, et al. The deepest divergences in land plants inferred from phylogenomic evidence[J]. Proc Natl Acad Sci U S A, 2006, 103(42): 15511-15516. |
| [21] | Ernst L, Wohl J, Bauerbach E, et al. Hydroxycinnamoyltransferase and CYP98 in phenolic metabolism in the rosmarinic acid-producing hornwort Anthoceros agrestis[J]. Planta, 2022, 255(4): 75. |
| [22] |
Zhang J, Fu XX, Li RQ, et al. The hornwort genome and early land plant evolution[J]. Nat Plants, 2020, 6(2): 107-118.
doi: 10.1038/s41477-019-0588-4 pmid: 32042158 |
| [23] |
刘阳星月, 张迪, 姚亚亚, 等. 大豆过敏原11S球蛋白G2中A2链结合表位的预测[J]. 食品科学, 2018, 39(18): 152-158.
doi: 10.7506/spkx1002-6630-201818024 |
| Liu YXY, Zhang D, Yao YY, et al. Mapping of ig G-binding epitopes on the A2 chain of the major soybean allergen 11S globulin G2[J]. Food Sci, 2018, 39(18): 152-158. | |
| [24] | 何姗. 农杆菌介导CmWRKY15-1基因对菊花的遗传转化[D]. 沈阳: 沈阳农业大学, 2020. |
| He S. Agrobacterium-mediated genetic transformation of CmWRKY15-1 gene to Chrysanthemum[D]. Shenyang: Shenyang Agricultural University, 2020. | |
| [25] | 魏加练. 基于转录组的粗山羊草响应镉胁迫关键基因挖掘及AetHIPP28的功能验证[D]. 贵阳: 贵州师范大学, 2022. |
| Wei JL. Identification and the function of Aegilops tauschii AetHIPP28 to cadmium stress based on transcriptome sequencing[D]. Guiyang: Guizhou Normal University, 2022. | |
| [26] | 李保华, 赵美琦. 梨叶抗黑星病作用机制研究[J]. 莱阳农学院学报, 2003, 20(3): 187-188. |
| Li BH, Zhao MQ. Resistant mechanism of pear leaves against Venturia nashicola[J]. J Laiyang Agric Coll, 2003, 20(3): 187-188. | |
| [27] |
Clough SJ, Bent AF. Floral dip: a simplified method for Agrobacte-rium-mediated transformation of Arabidopsis thaliana[J]. Plant J, 1998, 16(6): 735-743.
doi: 10.1046/j.1365-313x.1998.00343.x pmid: 10069079 |
| [28] |
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2ΔΔCT Method[J]. Methods, 2001, 25(4): 402-408.
doi: 10.1006/meth.2001.1262 pmid: 11846609 |
| [29] | 王殿, 庄亚妹, 徐华, 等. 甘蓝型油菜CCCH基因BnaA07g26050D的克隆及表达分析[J]. 山东师范大学学报: 自然科学版, 2017, 32(4): 137-143. |
| Wang D, Zhuang YM, Xu H, et al. Cloning and expression of the ccch gene bnaa07g26050d in brassica napus L[J]. J Shandong Norm Univ Nat Sci, 2017, 32(4): 137-143. | |
| [30] |
张蕊杰, 顾茁, 薛辰旭, 等. 青稞萌芽富集酚类物质品种筛选及条件优化[J]. 食品与发酵工业, 2022, 48(15): 193-200.
doi: 10.13995/j.cnki.11-1802/ts.028750 |
| Zhang RJ, Gu Z, Xue CX, et al. Screening of highland barley varieties using phenolics accumulation as indicator and the effect of germination conditions on phenolics[J]. Food Ferment Ind, 2022, 48(15): 193-200. | |
| [31] | Dong NQ, Lin HX. Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions[J]. J Integr Plant Biol, 2021, 63(1): 180-209. |
| [32] | Mizutani M, Ward E, DiMaio J, et al. Molecular cloning and sequencing of a cDNA encoding mung bean cytochrome P450(P450C4H)possessing cinnamate 4-hydroxylase activity[J]. Biochem Biophys Res Commun, 1993, 190(3): 875-880. |
| [33] |
Teutsch HG, Hasenfratz MP, Lesot A, et al. Isolation and sequence of a cDNA encoding the Jerusalem artichoke cinnamate 4-hydroxylase, a major plant cytochrome P450 involved in the general phenylpropanoid pathway[J]. Proc Natl Acad Sci USA, 1993, 90(9): 4102-4106.
doi: 10.1073/pnas.90.9.4102 pmid: 8097885 |
| [34] |
Lavhale SG, Kalunke RM, Giri AP. Structural, functional and evolutionary diversity of 4-coumarate-CoA ligase in plants[J]. Planta, 2018, 248(5): 1063-1078.
doi: 10.1007/s00425-018-2965-z pmid: 30078075 |
| [35] | Chen XH, Wang HT, Li XY, et al. Molecular cloning and functional analysis of 4-Coumarate: CoA ligase 4(4CL-like 1)from Fraxinus mandshurica and its role in abiotic stress tolerance and cell wall synthesis[J]. BMC Plant Biol, 2019, 19(1): 231. |
| [36] | Costa MA, Collins RE, Anterola AM, et al. An in silico assessment of gene function and organization of the phenylpropanoid pathway metabolic networks in Arabidopsis thaliana and limitations thereof[J]. Phytochemistry, 2003, 64(6): 1097-1112. |
| [37] | 潘琛, 任百光, 盖颖. 对香豆酰辅酶A酯生物合成及纯化方法[J]. 北京林业大学学报, 2016, 38(3): 120-124. |
| Pan C, Ren BG, Gai Y. Method of enzymatic synthesis and purification of p-coumaroyl-CoA[J]. J Beijing For Univ, 2016, 38(3): 120-124. | |
| [38] | 李洋, 唐雪冰, 李晓峰, 等. NtC3H基因对烟草类黄酮及绿原酸合成的影响[J]. 中国烟草科学, 2016, 37(1): 8-13. |
| Li Y, Tang XB, Li XF, et al. The influence of NtC3H on the synthesis of flavonoids and chlorogenic acid in tobacco[J]. Chin Tob Sci, 2016, 37(1): 8-13. | |
| [39] | Xin JK, Che TM, Huang XL, et al. A comprehensive view of metabolic responses to CYP98 perturbation in ancestral plants[J]. Plant Physiol Biochem, 2023, 201: 107793. |
| [40] | 高梅. 小立碗藓4- 香豆酰莽草酸/喹啉3'-羟化酶基因功能的初步研究[D]. 贵阳: 贵州师范大学, 2020. |
| Gao M. A preliminary study on the gene function of 4-coumaruylshikimate/quinoline 3'-hydroxylase[D]. Guiyang: Guizhou Normal University, 2020. | |
| [41] |
Boerjan W, Ralph J, Baucher M. Lignin biosynthesis[J]. Annu Rev Plant Biol, 2003, 54: 519-546.
pmid: 14503002 |
| [42] | 何利, 唐翠明, 戴凡炜, 等. 防御酶与植物抗青枯病关系研究进展[J]. 广东农业科学, 2013, 40(12): 101-103. |
| He L, Tang CM, Dai FW, et al. Relationship between the defense enzyme and the disease resistance of plants to bacterial wilt[J]. Guangdong Agric Sci, 2013, 40(12): 101-103. | |
| [43] | Li C, Zhang MM, Wei S, et al. Study of Walnut brown rot caused by Botryosphaeria dothidea in the Guizhou Province of China[J]. Crop Prot, 2023, 163: 106118. |
| [44] | Wang D, Xu H, Huang JY, et al. The Arabidopsis CCCH protein C3H14 contributes to basal defense against Botrytis cinerea mainly through the WRKY33-dependent pathway[J]. Plant Cell Environ, 2020, 43(7): 1792-1806. |
| [1] | 庞梦真, 徐汉琴, 刘海燕, 宋娟, 王佳涵, 孙丽娜, 姬佩梅, 尹泽芝, 胡又川, 赵晓萌, 梁闪闪, 张泗举, 栾维江. 水稻黄化早抽穗突变体 hz1 的基因鉴定及功能分析[J]. 生物技术通报, 2024, 40(7): 125-136. |
| [2] | 范宗强, 冯靖涵, 郑丽雪, 王硕, 彭向前, 陈芳. 枯草芽孢杆菌B579对黄瓜枯萎病的防治及其诱导抗性研究[J]. 生物技术通报, 2024, 40(7): 226-234. |
| [3] | 沈真辉, 曹瑶, 杨林雷, 罗祥英, 子灵山, 陆青青, 李荣春. 金耳和毛韧革菌麦角硫因生物合成基因的克隆及生物信息学分析[J]. 生物技术通报, 2024, 40(7): 259-272. |
| [4] | 王玉书, 赵琳琳, 赵爽, 胡琦, 白慧霞, 王欢, 曹业萍, 范震宇. 大白菜BrCYP83B1基因的克隆及表达分析[J]. 生物技术通报, 2024, 40(6): 152-160. |
| [5] | 郝思怡, 张君珂, 王斌, 曲朋燕, 李瑞得, 程春振. 香蕉ELF3的克隆与表达分析[J]. 生物技术通报, 2024, 40(5): 131-140. |
| [6] | 杜泽光, 任少文, 张凤勤, 李梅兰, 李改珍, 齐仙惠. 大白菜BrMLP328的克隆、表达及功能验证[J]. 生物技术通报, 2024, 40(4): 122-129. |
| [7] | 刘换换, 杨立春, 李火根. 北美鹅掌楸LtMYB305基因的克隆及功能分析[J]. 生物技术通报, 2024, 40(4): 179-188. |
| [8] | 钟匀, 林春, 刘正杰, 董陈文华, 毛自朝, 李兴玉. 芦笋皂苷合成相关糖基转移酶基因克隆及原核表达分析[J]. 生物技术通报, 2024, 40(4): 255-263. |
| [9] | 彭羽佳, 李文萃, 刘勇波. 昆虫对杀虫剂和转Bt基因植物的抗性进化机制研究进展[J]. 生物技术通报, 2024, 40(4): 40-51. |
| [10] | 方天一, 岳艳玲. 根系分泌物介导的植物对土传病害的抗性机制[J]. 生物技术通报, 2024, 40(3): 52-61. |
| [11] | 杨艳, 胡洋, 刘霓如, 殷璐, 杨锐, 王鹏飞, 穆霄鹏, 张帅, 程春振, 张建成. ‘红满堂’苹果MbbZIP43基因的克隆与功能研究[J]. 生物技术通报, 2024, 40(2): 146-159. |
| [12] | 任延靖, 张鲁刚, 赵孟良, 李江, 邵登魁. 白菜种子cDNA酵母文库的构建及BrTTG1互作蛋白的筛选及分析[J]. 生物技术通报, 2024, 40(2): 223-232. |
| [13] | 邹修为, 岳佳妮, 李志宇, 戴良英, 李魏. 水稻热激转录因子HsfA2b调控非生物胁迫抗性的功能分析[J]. 生物技术通报, 2024, 40(2): 90-98. |
| [14] | 王俊芳, 黄秋斌, 张飘丹, 张彭湃. Surfactin的结构、生物合成及其在生物防治中的作用[J]. 生物技术通报, 2024, 40(1): 100-112. |
| [15] | 朱毅, 柳唐镜, 宫国义, 张洁, 王晋芳, 张海英. 西瓜ClPP2C3克隆及表达分析[J]. 生物技术通报, 2024, 40(1): 243-249. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||